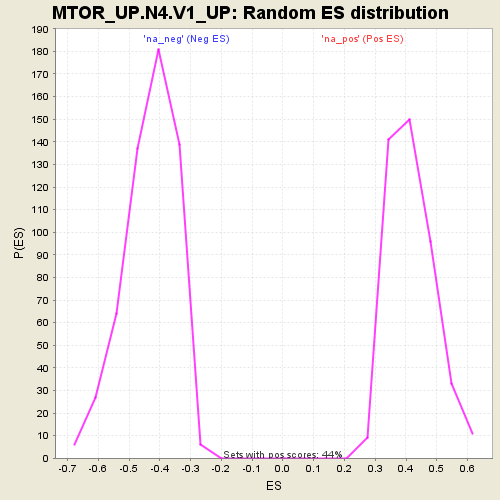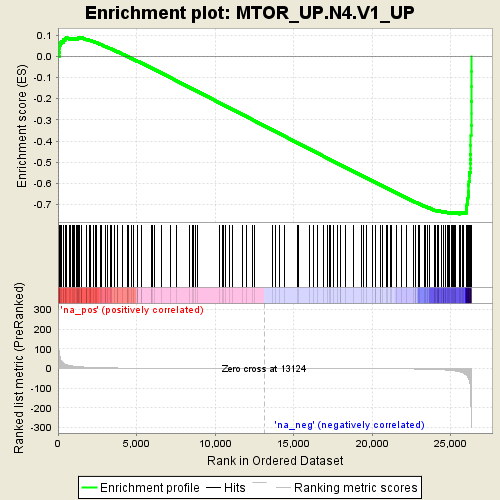
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | MTOR_UP.N4.V1_UP |
| Enrichment Score (ES) | -0.74464524 |
| Normalized Enrichment Score (NES) | -1.7292444 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.08763764 |
| FWER p-Value | 0.72 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LDLR | 63 | 89.302 | 0.0191 | No | ||
| 2 | MSMO1 | 74 | 78.192 | 0.0375 | No | ||
| 3 | TUBB2A | 105 | 65.807 | 0.0521 | No | ||
| 4 | MAP1A | 132 | 57.350 | 0.0649 | No | ||
| 5 | PHACTR2 | 202 | 40.583 | 0.0721 | No | ||
| 6 | DHCR7 | 331 | 27.123 | 0.0737 | No | ||
| 7 | SDF2L1 | 350 | 26.338 | 0.0793 | No | ||
| 8 | LARS2 | 442 | 22.350 | 0.0812 | No | ||
| 9 | IDH1 | 464 | 21.758 | 0.0856 | No | ||
| 10 | DNMT3B | 506 | 20.433 | 0.0890 | No | ||
| 11 | RRP9 | 724 | 15.790 | 0.0845 | No | ||
| 12 | SLC6A6 | 778 | 15.032 | 0.0861 | No | ||
| 13 | FKBP11 | 938 | 12.867 | 0.0831 | No | ||
| 14 | NQO2 | 988 | 12.307 | 0.0842 | No | ||
| 15 | ARG2 | 1068 | 11.448 | 0.0839 | No | ||
| 16 | SEC24D | 1182 | 10.500 | 0.0821 | No | ||
| 17 | VARS | 1220 | 10.228 | 0.0831 | No | ||
| 18 | KLHL7 | 1246 | 9.949 | 0.0846 | No | ||
| 19 | DBN1 | 1280 | 9.747 | 0.0856 | No | ||
| 20 | TBRG4 | 1282 | 9.742 | 0.0880 | No | ||
| 21 | FAS | 1372 | 9.028 | 0.0867 | No | ||
| 22 | HYAL3 | 1479 | 8.385 | 0.0847 | No | ||
| 23 | GRM2 | 1492 | 8.346 | 0.0862 | No | ||
| 24 | PIR | 1518 | 8.211 | 0.0872 | No | ||
| 25 | RIT1 | 1786 | 6.889 | 0.0787 | No | ||
| 26 | GAS2L1 | 1811 | 6.769 | 0.0794 | No | ||
| 27 | ELOVL6 | 1839 | 6.687 | 0.0800 | No | ||
| 28 | ARL3 | 1981 | 6.116 | 0.0760 | No | ||
| 29 | CCDC86 | 2048 | 5.893 | 0.0749 | No | ||
| 30 | SLC39A4 | 2088 | 5.750 | 0.0748 | No | ||
| 31 | PTS | 2256 | 5.233 | 0.0697 | No | ||
| 32 | CAMKV | 2386 | 4.878 | 0.0659 | No | ||
| 33 | NANS | 2419 | 4.792 | 0.0659 | No | ||
| 34 | ETNK1 | 2708 | 4.158 | 0.0558 | No | ||
| 35 | ISOC2 | 2754 | 4.070 | 0.0551 | No | ||
| 36 | SLCO4A1 | 2989 | 3.641 | 0.0470 | No | ||
| 37 | ITPR1 | 2995 | 3.635 | 0.0477 | No | ||
| 38 | LPCAT4 | 3146 | 3.360 | 0.0428 | No | ||
| 39 | ADARB1 | 3336 | 3.081 | 0.0363 | No | ||
| 40 | ACACA | 3408 | 2.975 | 0.0343 | No | ||
| 41 | ABHD6 | 3577 | 2.771 | 0.0285 | No | ||
| 42 | CHUK | 3810 | 2.518 | 0.0203 | No | ||
| 43 | MGAT4A | 3815 | 2.513 | 0.0207 | No | ||
| 44 | GK | 4091 | 2.229 | 0.0107 | No | ||
| 45 | VPS54 | 4418 | 1.916 | -0.0013 | No | ||
| 46 | ASPHD1 | 4435 | 1.895 | -0.0014 | No | ||
| 47 | UGCG | 4469 | 1.868 | -0.0023 | No | ||
| 48 | EBNA1BP2 | 4700 | 1.692 | -0.0107 | No | ||
| 49 | RBM28 | 4771 | 1.642 | -0.0129 | No | ||
| 50 | PAK1IP1 | 5028 | 1.454 | -0.0224 | No | ||
| 51 | RRP12 | 5036 | 1.445 | -0.0223 | No | ||
| 52 | SAR1B | 5060 | 1.433 | -0.0228 | No | ||
| 53 | MAST2 | 5317 | 1.283 | -0.0323 | No | ||
| 54 | CDK16 | 5341 | 1.270 | -0.0329 | No | ||
| 55 | TMEM135 | 5918 | 0.964 | -0.0547 | No | ||
| 56 | CHORDC1 | 5954 | 0.948 | -0.0558 | No | ||
| 57 | OPN3 | 6021 | 0.922 | -0.0581 | No | ||
| 58 | GBE1 | 6131 | 0.886 | -0.0621 | No | ||
| 59 | DLEU1 | 6583 | 0.732 | -0.0791 | No | ||
| 60 | COQ2 | 7181 | 0.554 | -0.1019 | No | ||
| 61 | TFPT | 7536 | 0.478 | -0.1153 | No | ||
| 62 | BNIP3 | 8337 | 0.349 | -0.1458 | No | ||
| 63 | TSNAXIP1 | 8535 | 0.318 | -0.1533 | No | ||
| 64 | IFI30 | 8636 | 0.305 | -0.1570 | No | ||
| 65 | NME7 | 8763 | 0.289 | -0.1618 | No | ||
| 66 | CALML4 | 8889 | 0.273 | -0.1665 | No | ||
| 67 | LHX2 | 10291 | 0.137 | -0.2200 | No | ||
| 68 | RABEPK | 10491 | 0.125 | -0.2276 | No | ||
| 69 | IDH3A | 10539 | 0.123 | -0.2294 | No | ||
| 70 | LGMN | 10636 | 0.114 | -0.2330 | No | ||
| 71 | COX10 | 10938 | 0.096 | -0.2445 | No | ||
| 72 | ITK | 11077 | 0.088 | -0.2498 | No | ||
| 73 | MRPL22 | 11734 | 0.053 | -0.2749 | No | ||
| 74 | VPS33B | 11979 | 0.045 | -0.2842 | No | ||
| 75 | GH1 | 12388 | 0.027 | -0.2998 | No | ||
| 76 | OPRL1 | 12530 | 0.021 | -0.3052 | No | ||
| 77 | KIF21B | 13676 | -0.019 | -0.3490 | No | ||
| 78 | PCCA | 13862 | -0.026 | -0.3561 | No | ||
| 79 | MIPEP | 14123 | -0.037 | -0.3660 | No | ||
| 80 | DOK2 | 14447 | -0.049 | -0.3783 | No | ||
| 81 | NUP155 | 15252 | -0.087 | -0.4091 | No | ||
| 82 | EIF2B3 | 15330 | -0.092 | -0.4120 | No | ||
| 83 | RPAP1 | 15992 | -0.133 | -0.4373 | No | ||
| 84 | CYTL1 | 16278 | -0.152 | -0.4481 | No | ||
| 85 | USH1C | 16533 | -0.173 | -0.4578 | No | ||
| 86 | STX4 | 16906 | -0.208 | -0.4720 | No | ||
| 87 | PFKM | 17177 | -0.236 | -0.4822 | No | ||
| 88 | PDXK | 17300 | -0.249 | -0.4869 | No | ||
| 89 | COASY | 17376 | -0.259 | -0.4897 | No | ||
| 90 | SREBF1 | 17541 | -0.279 | -0.4959 | No | ||
| 91 | PANK3 | 17811 | -0.310 | -0.5061 | No | ||
| 92 | GFPT1 | 17970 | -0.331 | -0.5120 | No | ||
| 93 | IL21R | 18015 | -0.337 | -0.5137 | No | ||
| 94 | CASR | 18318 | -0.384 | -0.5251 | No | ||
| 95 | PDHX | 18835 | -0.480 | -0.5447 | No | ||
| 96 | CNIH4 | 19338 | -0.588 | -0.5638 | No | ||
| 97 | YRDC | 19473 | -0.624 | -0.5688 | No | ||
| 98 | CD79B | 19615 | -0.663 | -0.5740 | No | ||
| 99 | CLPB | 20040 | -0.794 | -0.5900 | No | ||
| 100 | MCPH1 | 20189 | -0.844 | -0.5955 | No | ||
| 101 | GALK2 | 20225 | -0.859 | -0.5966 | No | ||
| 102 | PJA1 | 20524 | -0.970 | -0.6078 | No | ||
| 103 | DDN | 20555 | -0.983 | -0.6087 | No | ||
| 104 | GTF2E2 | 20629 | -1.011 | -0.6113 | No | ||
| 105 | SLC25A32 | 20909 | -1.144 | -0.6217 | No | ||
| 106 | GALE | 20953 | -1.168 | -0.6230 | No | ||
| 107 | FGF1 | 21184 | -1.300 | -0.6315 | No | ||
| 108 | PLS1 | 21218 | -1.318 | -0.6325 | No | ||
| 109 | PTPRK | 21535 | -1.506 | -0.6442 | No | ||
| 110 | GPRC5C | 21896 | -1.762 | -0.6575 | No | ||
| 111 | E2F6 | 22173 | -2.013 | -0.6676 | No | ||
| 112 | TRPM6 | 22608 | -2.456 | -0.6836 | No | ||
| 113 | BSPRY | 22743 | -2.612 | -0.6881 | No | ||
| 114 | CUTC | 22792 | -2.680 | -0.6893 | No | ||
| 115 | SLC31A1 | 22957 | -2.889 | -0.6949 | No | ||
| 116 | ASCC1 | 23033 | -2.995 | -0.6970 | No | ||
| 117 | SLC4A1AP | 23309 | -3.437 | -0.7067 | No | ||
| 118 | OPCML | 23382 | -3.578 | -0.7086 | No | ||
| 119 | SYNPO | 23506 | -3.787 | -0.7124 | No | ||
| 120 | SLC7A5 | 23621 | -4.053 | -0.7158 | No | ||
| 121 | ARL4A | 23626 | -4.062 | -0.7150 | No | ||
| 122 | RRP15 | 23988 | -4.979 | -0.7276 | No | ||
| 123 | NFIL3 | 24002 | -5.017 | -0.7269 | No | ||
| 124 | GOT1 | 24030 | -5.126 | -0.7267 | No | ||
| 125 | BHLHE40 | 24138 | -5.407 | -0.7295 | No | ||
| 126 | POLR3G | 24148 | -5.433 | -0.7285 | No | ||
| 127 | PHGDH | 24203 | -5.598 | -0.7292 | No | ||
| 128 | PLCB1 | 24226 | -5.673 | -0.7287 | No | ||
| 129 | CLIC4 | 24240 | -5.709 | -0.7278 | No | ||
| 130 | SSTR2 | 24422 | -6.466 | -0.7332 | No | ||
| 131 | EAF2 | 24525 | -6.869 | -0.7355 | No | ||
| 132 | GLRX2 | 24554 | -7.024 | -0.7348 | No | ||
| 133 | EIF4EBP1 | 24571 | -7.108 | -0.7337 | No | ||
| 134 | MFAP4 | 24652 | -7.457 | -0.7350 | No | ||
| 135 | FAH | 24691 | -7.612 | -0.7346 | No | ||
| 136 | GCH1 | 24774 | -8.103 | -0.7358 | No | ||
| 137 | MARS | 24894 | -8.832 | -0.7383 | No | ||
| 138 | APOL6 | 24931 | -9.126 | -0.7374 | No | ||
| 139 | SLC3A2 | 25078 | -10.370 | -0.7405 | No | ||
| 140 | ASS1 | 25113 | -10.709 | -0.7393 | No | ||
| 141 | E2F5 | 25198 | -11.551 | -0.7397 | No | ||
| 142 | MRPL39 | 25226 | -11.847 | -0.7379 | No | ||
| 143 | SPRED2 | 25290 | -12.547 | -0.7373 | No | ||
| 144 | TM6SF1 | 25337 | -13.101 | -0.7359 | No | ||
| 145 | TRAF3IP2 | 25567 | -16.643 | -0.7406 | Yes | ||
| 146 | ADAM9 | 25606 | -17.916 | -0.7378 | Yes | ||
| 147 | PYCR1 | 25769 | -21.972 | -0.7387 | Yes | ||
| 148 | CARS | 25844 | -24.758 | -0.7356 | Yes | ||
| 149 | FAM129A | 25984 | -32.982 | -0.7330 | Yes | ||
| 150 | SLC6A9 | 25989 | -33.245 | -0.7251 | Yes | ||
| 151 | BCHE | 26012 | -34.895 | -0.7176 | Yes | ||
| 152 | ANXA3 | 26030 | -37.029 | -0.7093 | Yes | ||
| 153 | SLC39A14 | 26039 | -37.712 | -0.7006 | Yes | ||
| 154 | TIMM44 | 26063 | -40.273 | -0.6918 | Yes | ||
| 155 | SLC1A5 | 26086 | -42.020 | -0.6825 | Yes | ||
| 156 | LIMA1 | 26094 | -42.892 | -0.6725 | Yes | ||
| 157 | BCAT1 | 26106 | -44.511 | -0.6622 | Yes | ||
| 158 | PSPH | 26121 | -46.794 | -0.6515 | Yes | ||
| 159 | SLC7A11 | 26126 | -47.089 | -0.6403 | Yes | ||
| 160 | RIMS3 | 26155 | -52.334 | -0.6288 | Yes | ||
| 161 | ULBP1 | 26156 | -52.544 | -0.6162 | Yes | ||
| 162 | HERPUD1 | 26159 | -53.015 | -0.6035 | Yes | ||
| 163 | ADM | 26168 | -55.394 | -0.5905 | Yes | ||
| 164 | VLDLR | 26190 | -59.843 | -0.5769 | Yes | ||
| 165 | SARS | 26192 | -60.122 | -0.5625 | Yes | ||
| 166 | CEBPG | 26210 | -66.827 | -0.5471 | Yes | ||
| 167 | ASNS | 26241 | -83.978 | -0.5280 | Yes | ||
| 168 | CEBPB | 26244 | -86.643 | -0.5073 | Yes | ||
| 169 | VEGFA | 26251 | -89.987 | -0.4859 | Yes | ||
| 170 | SLC43A1 | 26263 | -98.209 | -0.4627 | Yes | ||
| 171 | DDIT3 | 26291 | -176.939 | -0.4212 | Yes | ||
| 172 | ATF3 | 26295 | -192.665 | -0.3750 | Yes | ||
| 173 | PCK2 | 26298 | -198.705 | -0.3273 | Yes | ||
| 174 | IFRD1 | 26303 | -230.785 | -0.2720 | Yes | ||
| 175 | TRIB3 | 26304 | -235.323 | -0.2154 | Yes | ||
| 176 | DDIT4 | 26311 | -299.207 | -0.1437 | Yes | ||
| 177 | CHAC1 | 26314 | -299.207 | -0.0719 | Yes | ||
| 178 | TSC22D3 | 26317 | -299.207 | 0.0000 | Yes |