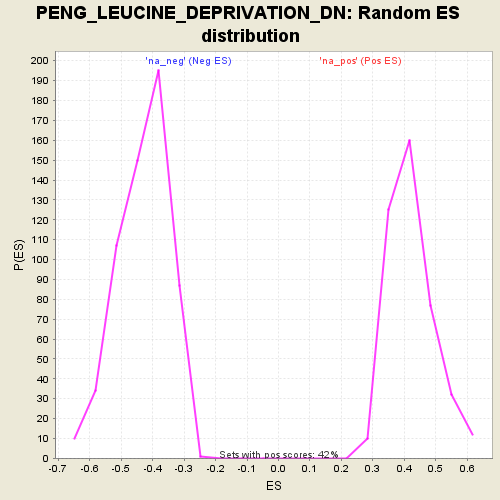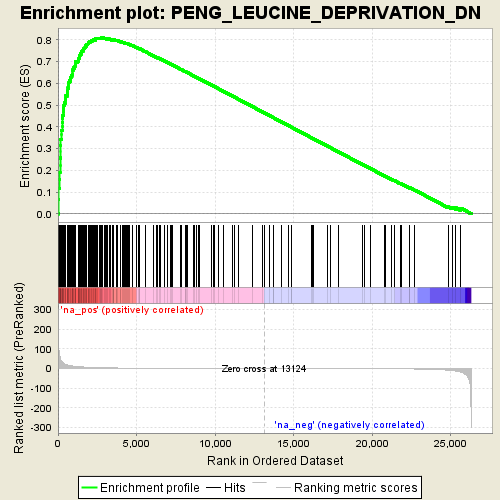
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PENG_LEUCINE_DEPRIVATION_DN |
| Enrichment Score (ES) | 0.81081855 |
| Normalized Enrichment Score (NES) | 1.9273204 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IFITM1 | 30 | 118.511 | 0.0661 | Yes | ||
| 2 | HSPA8 | 56 | 93.481 | 0.1182 | Yes | ||
| 3 | IDI1 | 88 | 70.501 | 0.1571 | Yes | ||
| 4 | SQLE | 112 | 62.535 | 0.1917 | Yes | ||
| 5 | MLEC | 128 | 58.306 | 0.2242 | Yes | ||
| 6 | TUBB4B | 131 | 57.538 | 0.2568 | Yes | ||
| 7 | HSPA5 | 151 | 52.337 | 0.2858 | Yes | ||
| 8 | AMD1 | 152 | 52.233 | 0.3154 | Yes | ||
| 9 | KPNA2 | 160 | 49.616 | 0.3433 | Yes | ||
| 10 | DNAJA1 | 219 | 38.203 | 0.3628 | Yes | ||
| 11 | PDIA4 | 222 | 37.994 | 0.3842 | Yes | ||
| 12 | EI24 | 260 | 34.262 | 0.4023 | Yes | ||
| 13 | CALR | 266 | 32.957 | 0.4208 | Yes | ||
| 14 | DHX9 | 284 | 31.114 | 0.4378 | Yes | ||
| 15 | PNP | 301 | 29.501 | 0.4539 | Yes | ||
| 16 | UBAP2L | 327 | 27.414 | 0.4685 | Yes | ||
| 17 | EIF2S1 | 344 | 26.480 | 0.4830 | Yes | ||
| 18 | UBE2N | 368 | 25.440 | 0.4965 | Yes | ||
| 19 | CCNB1 | 387 | 24.655 | 0.5098 | Yes | ||
| 20 | MANF | 448 | 22.191 | 0.5201 | Yes | ||
| 21 | TRIP13 | 457 | 21.942 | 0.5323 | Yes | ||
| 22 | YWHAH | 496 | 20.725 | 0.5426 | Yes | ||
| 23 | RASSF2 | 572 | 18.817 | 0.5504 | Yes | ||
| 24 | MLLT11 | 588 | 18.479 | 0.5603 | Yes | ||
| 25 | EIF4G1 | 621 | 17.869 | 0.5692 | Yes | ||
| 26 | PSME4 | 628 | 17.753 | 0.5791 | Yes | ||
| 27 | GLO1 | 669 | 16.943 | 0.5872 | Yes | ||
| 28 | SF3A1 | 683 | 16.727 | 0.5962 | Yes | ||
| 29 | FAM98A | 687 | 16.541 | 0.6054 | Yes | ||
| 30 | NOP16 | 706 | 16.201 | 0.6139 | Yes | ||
| 31 | SF3B4 | 806 | 14.791 | 0.6185 | Yes | ||
| 32 | RBM14 | 808 | 14.771 | 0.6269 | Yes | ||
| 33 | HSPE1 | 829 | 14.316 | 0.6343 | Yes | ||
| 34 | MRPL3 | 888 | 13.527 | 0.6397 | Yes | ||
| 35 | MRPS12 | 910 | 13.251 | 0.6464 | Yes | ||
| 36 | DHX30 | 911 | 13.236 | 0.6539 | Yes | ||
| 37 | TGFB1 | 913 | 13.218 | 0.6614 | Yes | ||
| 38 | POLRMT | 957 | 12.584 | 0.6669 | Yes | ||
| 39 | MAP2K3 | 1002 | 12.112 | 0.6721 | Yes | ||
| 40 | PSMD1 | 1070 | 11.432 | 0.6760 | Yes | ||
| 41 | PPIB | 1084 | 11.233 | 0.6819 | Yes | ||
| 42 | EMD | 1090 | 11.185 | 0.6881 | Yes | ||
| 43 | FASN | 1094 | 11.155 | 0.6943 | Yes | ||
| 44 | SCD | 1102 | 11.098 | 0.7003 | Yes | ||
| 45 | CDK2AP2 | 1268 | 9.840 | 0.6996 | Yes | ||
| 46 | XPO6 | 1289 | 9.708 | 0.7043 | Yes | ||
| 47 | DYNLL1 | 1319 | 9.478 | 0.7086 | Yes | ||
| 48 | PTBP1 | 1323 | 9.452 | 0.7138 | Yes | ||
| 49 | HSP90AA1 | 1345 | 9.282 | 0.7183 | Yes | ||
| 50 | NUTF2 | 1359 | 9.145 | 0.7230 | Yes | ||
| 51 | CCNA2 | 1369 | 9.062 | 0.7278 | Yes | ||
| 52 | UNG | 1403 | 8.838 | 0.7316 | Yes | ||
| 53 | RCC1 | 1411 | 8.790 | 0.7363 | Yes | ||
| 54 | EBP | 1467 | 8.488 | 0.7390 | Yes | ||
| 55 | MCM4 | 1485 | 8.358 | 0.7431 | Yes | ||
| 56 | NMT1 | 1500 | 8.281 | 0.7473 | Yes | ||
| 57 | TDG | 1554 | 8.034 | 0.7498 | Yes | ||
| 58 | SLC29A1 | 1591 | 7.776 | 0.7528 | Yes | ||
| 59 | SEC61B | 1634 | 7.567 | 0.7555 | Yes | ||
| 60 | PRMT1 | 1642 | 7.527 | 0.7595 | Yes | ||
| 61 | TMEM109 | 1688 | 7.341 | 0.7620 | Yes | ||
| 62 | PSMD11 | 1710 | 7.211 | 0.7652 | Yes | ||
| 63 | GGA2 | 1727 | 7.147 | 0.7687 | Yes | ||
| 64 | FDPS | 1746 | 7.063 | 0.7720 | Yes | ||
| 65 | PSMF1 | 1785 | 6.897 | 0.7745 | Yes | ||
| 66 | NUP98 | 1819 | 6.752 | 0.7770 | Yes | ||
| 67 | ACAT2 | 1837 | 6.703 | 0.7802 | Yes | ||
| 68 | HSP90AB1 | 1905 | 6.375 | 0.7813 | Yes | ||
| 69 | PLCG1 | 1918 | 6.321 | 0.7844 | Yes | ||
| 70 | AHSA1 | 1955 | 6.206 | 0.7865 | Yes | ||
| 71 | PDIA6 | 1980 | 6.116 | 0.7891 | Yes | ||
| 72 | SSB | 1998 | 6.067 | 0.7919 | Yes | ||
| 73 | CTSC | 2055 | 5.878 | 0.7931 | Yes | ||
| 74 | HPRT1 | 2135 | 5.621 | 0.7932 | Yes | ||
| 75 | TUBB4A | 2146 | 5.591 | 0.7960 | Yes | ||
| 76 | CYCS | 2189 | 5.436 | 0.7975 | Yes | ||
| 77 | ALG8 | 2241 | 5.280 | 0.7986 | Yes | ||
| 78 | VDAC1 | 2272 | 5.196 | 0.8004 | Yes | ||
| 79 | FCHO1 | 2328 | 5.023 | 0.8011 | Yes | ||
| 80 | FKBP2 | 2390 | 4.870 | 0.8015 | Yes | ||
| 81 | PTGES3 | 2391 | 4.867 | 0.8043 | Yes | ||
| 82 | WDR1 | 2395 | 4.857 | 0.8069 | Yes | ||
| 83 | NCL | 2432 | 4.764 | 0.8083 | Yes | ||
| 84 | DHCR24 | 2520 | 4.587 | 0.8075 | Yes | ||
| 85 | XRCC5 | 2613 | 4.389 | 0.8065 | Yes | ||
| 86 | TIMM17A | 2674 | 4.256 | 0.8066 | Yes | ||
| 87 | PTPN6 | 2728 | 4.129 | 0.8069 | Yes | ||
| 88 | OGFR | 2747 | 4.090 | 0.8086 | Yes | ||
| 89 | CSE1L | 2773 | 4.034 | 0.8099 | Yes | ||
| 90 | MLF2 | 2809 | 3.960 | 0.8108 | Yes | ||
| 91 | NXF1 | 2953 | 3.692 | 0.8074 | No | ||
| 92 | TCP1 | 3035 | 3.553 | 0.8064 | No | ||
| 93 | NOP56 | 3113 | 3.435 | 0.8054 | No | ||
| 94 | HSP90B1 | 3151 | 3.351 | 0.8059 | No | ||
| 95 | PSMA3 | 3248 | 3.209 | 0.8040 | No | ||
| 96 | LDHA | 3339 | 3.076 | 0.8023 | No | ||
| 97 | PSMD14 | 3434 | 2.947 | 0.8004 | No | ||
| 98 | GSPT1 | 3503 | 2.866 | 0.7994 | No | ||
| 99 | TBC1D2B | 3514 | 2.839 | 0.8006 | No | ||
| 100 | HSPA4 | 3529 | 2.822 | 0.8017 | No | ||
| 101 | PSMA5 | 3694 | 2.650 | 0.7969 | No | ||
| 102 | RRP1B | 3696 | 2.648 | 0.7984 | No | ||
| 103 | STIP1 | 3782 | 2.547 | 0.7966 | No | ||
| 104 | PSMB7 | 3958 | 2.358 | 0.7912 | No | ||
| 105 | ERH | 3990 | 2.325 | 0.7914 | No | ||
| 106 | ILF3 | 4120 | 2.204 | 0.7877 | No | ||
| 107 | PSMD2 | 4164 | 2.139 | 0.7872 | No | ||
| 108 | COPB2 | 4245 | 2.065 | 0.7854 | No | ||
| 109 | PRPF40A | 4277 | 2.035 | 0.7853 | No | ||
| 110 | PSMD3 | 4351 | 1.970 | 0.7837 | No | ||
| 111 | LDHB | 4389 | 1.937 | 0.7833 | No | ||
| 112 | PIK3R3 | 4491 | 1.856 | 0.7805 | No | ||
| 113 | UMPS | 4570 | 1.795 | 0.7786 | No | ||
| 114 | KDM1A | 4707 | 1.686 | 0.7743 | No | ||
| 115 | ALDOA | 4962 | 1.503 | 0.7655 | No | ||
| 116 | FARSA | 5151 | 1.379 | 0.7590 | No | ||
| 117 | PGAM1 | 5155 | 1.378 | 0.7597 | No | ||
| 118 | SPINT2 | 5209 | 1.349 | 0.7584 | No | ||
| 119 | LPCAT1 | 5545 | 1.150 | 0.7463 | No | ||
| 120 | PAPOLA | 6073 | 0.906 | 0.7266 | No | ||
| 121 | ZC3H3 | 6255 | 0.837 | 0.7202 | No | ||
| 122 | ACOT7 | 6361 | 0.804 | 0.7166 | No | ||
| 123 | AURKA | 6451 | 0.778 | 0.7137 | No | ||
| 124 | FKBP4 | 6553 | 0.747 | 0.7102 | No | ||
| 125 | EMP3 | 6798 | 0.660 | 0.7013 | No | ||
| 126 | GART | 6957 | 0.616 | 0.6956 | No | ||
| 127 | MTHFD1 | 7190 | 0.551 | 0.6870 | No | ||
| 128 | MPL | 7250 | 0.538 | 0.6851 | No | ||
| 129 | API5 | 7264 | 0.535 | 0.6849 | No | ||
| 130 | DNAJC11 | 7824 | 0.424 | 0.6637 | No | ||
| 131 | SRM | 7877 | 0.416 | 0.6620 | No | ||
| 132 | CYB561 | 8112 | 0.378 | 0.6532 | No | ||
| 133 | FAM89B | 8130 | 0.376 | 0.6528 | No | ||
| 134 | BYSL | 8164 | 0.371 | 0.6517 | No | ||
| 135 | ADRM1 | 8251 | 0.361 | 0.6486 | No | ||
| 136 | CCT5 | 8648 | 0.304 | 0.6337 | No | ||
| 137 | CCT2 | 8673 | 0.302 | 0.6329 | No | ||
| 138 | UBC | 8798 | 0.286 | 0.6283 | No | ||
| 139 | NBL1 | 8914 | 0.269 | 0.6241 | No | ||
| 140 | DNAJB6 | 9034 | 0.256 | 0.6197 | No | ||
| 141 | IFFO1 | 9743 | 0.183 | 0.5927 | No | ||
| 142 | PGK1 | 9909 | 0.170 | 0.5865 | No | ||
| 143 | EIF4G2 | 9946 | 0.167 | 0.5852 | No | ||
| 144 | PSMA7 | 10198 | 0.144 | 0.5757 | No | ||
| 145 | GCDH | 10525 | 0.123 | 0.5633 | No | ||
| 146 | BRD2 | 11121 | 0.086 | 0.5406 | No | ||
| 147 | SNRPA1 | 11250 | 0.079 | 0.5357 | No | ||
| 148 | LIMK2 | 11516 | 0.064 | 0.5256 | No | ||
| 149 | WDR43 | 12366 | 0.028 | 0.4931 | No | ||
| 150 | LRRC32 | 13049 | 0.003 | 0.4671 | No | ||
| 151 | CAD | 13116 | 0.000 | 0.4645 | No | ||
| 152 | VRK1 | 13468 | -0.011 | 0.4511 | No | ||
| 153 | CCT4 | 13728 | -0.021 | 0.4412 | No | ||
| 154 | CCT6A | 13733 | -0.021 | 0.4411 | No | ||
| 155 | UBE2S | 14235 | -0.042 | 0.4219 | No | ||
| 156 | MRPL28 | 14646 | -0.056 | 0.4063 | No | ||
| 157 | TOMM40 | 14887 | -0.066 | 0.3971 | No | ||
| 158 | PSMC3IP | 14896 | -0.067 | 0.3969 | No | ||
| 159 | ME2 | 16123 | -0.142 | 0.3500 | No | ||
| 160 | TK1 | 16232 | -0.150 | 0.3460 | No | ||
| 161 | MFSD11 | 16237 | -0.150 | 0.3459 | No | ||
| 162 | PFKM | 17177 | -0.236 | 0.3101 | No | ||
| 163 | DDX21 | 17375 | -0.259 | 0.3027 | No | ||
| 164 | MMP15 | 17876 | -0.319 | 0.2838 | No | ||
| 165 | EIF2B4 | 19360 | -0.594 | 0.2274 | No | ||
| 166 | POLR2J | 19513 | -0.634 | 0.2219 | No | ||
| 167 | MRPL12 | 19897 | -0.749 | 0.2077 | No | ||
| 168 | DDX1 | 20762 | -1.072 | 0.1753 | No | ||
| 169 | FANCI | 20836 | -1.109 | 0.1731 | No | ||
| 170 | COPS2 | 21203 | -1.310 | 0.1598 | No | ||
| 171 | TUBA4A | 21428 | -1.440 | 0.1521 | No | ||
| 172 | BCL2L1 | 21802 | -1.687 | 0.1388 | No | ||
| 173 | CEBPZ | 21879 | -1.749 | 0.1369 | No | ||
| 174 | ETF1 | 22379 | -2.197 | 0.1190 | No | ||
| 175 | DYRK1A | 22401 | -2.220 | 0.1195 | No | ||
| 176 | RNF126 | 22726 | -2.589 | 0.1085 | No | ||
| 177 | EIF5 | 24866 | -8.648 | 0.0316 | No | ||
| 178 | TNFRSF1A | 25105 | -10.602 | 0.0285 | No | ||
| 179 | BST2 | 25313 | -12.832 | 0.0279 | No | ||
| 180 | TFRC | 25640 | -18.457 | 0.0259 | No |