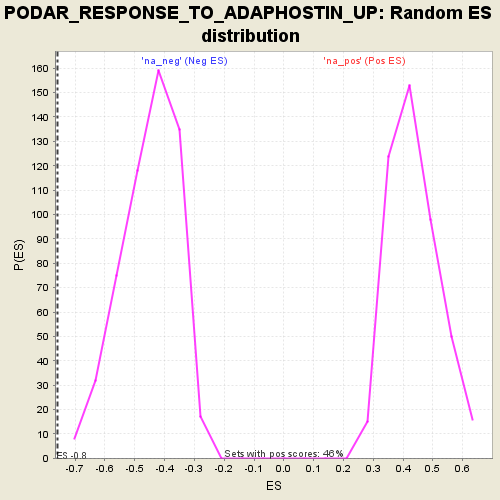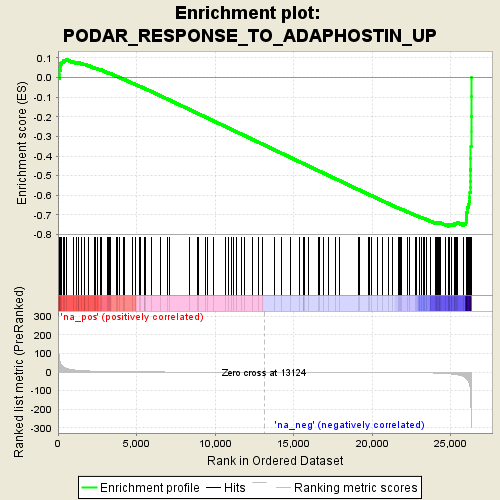
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | PODAR_RESPONSE_TO_ADAPHOSTIN_UP |
| Enrichment Score (ES) | -0.7582266 |
| Normalized Enrichment Score (NES) | -1.6868664 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.20575094 |
| FWER p-Value | 0.991 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TUBB2A | 105 | 65.807 | 0.0177 | No | ||
| 2 | TNFSF9 | 108 | 63.761 | 0.0387 | No | ||
| 3 | BBC3 | 149 | 52.488 | 0.0545 | No | ||
| 4 | DNAJB1 | 161 | 49.447 | 0.0704 | No | ||
| 5 | RAP2B | 246 | 35.107 | 0.0787 | No | ||
| 6 | HEXIM1 | 324 | 27.700 | 0.0849 | No | ||
| 7 | NLRP1 | 419 | 23.262 | 0.0890 | No | ||
| 8 | TUBA1A | 529 | 19.799 | 0.0914 | No | ||
| 9 | KLHL21 | 955 | 12.622 | 0.0793 | No | ||
| 10 | SEC24D | 1182 | 10.500 | 0.0742 | No | ||
| 11 | CCNG2 | 1274 | 9.814 | 0.0739 | No | ||
| 12 | HDLBP | 1284 | 9.733 | 0.0768 | No | ||
| 13 | PIR | 1518 | 8.211 | 0.0706 | No | ||
| 14 | SOX4 | 1692 | 7.315 | 0.0664 | No | ||
| 15 | SLC38A2 | 1705 | 7.228 | 0.0683 | No | ||
| 16 | MIR22HG | 1959 | 6.191 | 0.0607 | No | ||
| 17 | ADAM17 | 2312 | 5.061 | 0.0489 | No | ||
| 18 | NEAT1 | 2398 | 4.852 | 0.0473 | No | ||
| 19 | P4HA2 | 2497 | 4.639 | 0.0451 | No | ||
| 20 | GTPBP2 | 2692 | 4.205 | 0.0391 | No | ||
| 21 | DNAJB9 | 2733 | 4.122 | 0.0389 | No | ||
| 22 | HMOX1 | 3141 | 3.368 | 0.0245 | No | ||
| 23 | IER5 | 3192 | 3.300 | 0.0236 | No | ||
| 24 | CLU | 3249 | 3.207 | 0.0225 | No | ||
| 25 | ATF4 | 3319 | 3.105 | 0.0209 | No | ||
| 26 | PRDM4 | 3748 | 2.577 | 0.0054 | No | ||
| 27 | ZFAND5 | 3813 | 2.515 | 0.0038 | No | ||
| 28 | PNRC1 | 3888 | 2.423 | 0.0018 | No | ||
| 29 | BSDC1 | 4173 | 2.130 | -0.0083 | No | ||
| 30 | RPL13P5 | 4258 | 2.058 | -0.0109 | No | ||
| 31 | STEAP1 | 4752 | 1.651 | -0.0292 | No | ||
| 32 | GLA | 4957 | 1.508 | -0.0365 | No | ||
| 33 | RPS6KC1 | 5169 | 1.371 | -0.0441 | No | ||
| 34 | CREM | 5243 | 1.328 | -0.0464 | No | ||
| 35 | ANXA1 | 5495 | 1.180 | -0.0556 | No | ||
| 36 | ACTR10 | 5523 | 1.163 | -0.0563 | No | ||
| 37 | MSX1 | 5556 | 1.144 | -0.0571 | No | ||
| 38 | KLF2 | 5920 | 0.963 | -0.0706 | No | ||
| 39 | MGAT2 | 6545 | 0.749 | -0.0942 | No | ||
| 40 | RAB9A | 6970 | 0.613 | -0.1102 | No | ||
| 41 | MAP1LC3B | 7078 | 0.582 | -0.1141 | No | ||
| 42 | EIF2AK3 | 8352 | 0.344 | -0.1626 | No | ||
| 43 | DNAJB2 | 8887 | 0.273 | -0.1829 | No | ||
| 44 | GOLGA2 | 8936 | 0.267 | -0.1847 | No | ||
| 45 | DGKG | 9417 | 0.215 | -0.2029 | No | ||
| 46 | HSPB1 | 9513 | 0.206 | -0.2065 | No | ||
| 47 | ZBTB43 | 9910 | 0.170 | -0.2216 | No | ||
| 48 | MORC3 | 10652 | 0.113 | -0.2498 | No | ||
| 49 | ISG20 | 10851 | 0.100 | -0.2574 | No | ||
| 50 | STX12 | 11037 | 0.090 | -0.2644 | No | ||
| 51 | ZFP36L1 | 11150 | 0.084 | -0.2687 | No | ||
| 52 | ZNF165 | 11389 | 0.071 | -0.2777 | No | ||
| 53 | NCOA2 | 11697 | 0.055 | -0.2894 | No | ||
| 54 | GADD45A | 11891 | 0.047 | -0.2968 | No | ||
| 55 | TRIB1 | 12362 | 0.028 | -0.3147 | No | ||
| 56 | SAT1 | 12782 | 0.011 | -0.3307 | No | ||
| 57 | ARMCX3 | 13026 | 0.005 | -0.3400 | No | ||
| 58 | HBP1 | 13784 | -0.023 | -0.3689 | No | ||
| 59 | MAFG | 14243 | -0.042 | -0.3864 | No | ||
| 60 | IL15 | 14782 | -0.061 | -0.4069 | No | ||
| 61 | NAP1L2 | 15347 | -0.094 | -0.4284 | No | ||
| 62 | RB1CC1 | 15618 | -0.109 | -0.4387 | No | ||
| 63 | HIST1H2BG | 15620 | -0.110 | -0.4387 | No | ||
| 64 | JUND | 15695 | -0.113 | -0.4415 | No | ||
| 65 | GLRX | 15957 | -0.131 | -0.4514 | No | ||
| 66 | CLIC2 | 16590 | -0.179 | -0.4755 | No | ||
| 67 | CDKN1C | 16666 | -0.187 | -0.4783 | No | ||
| 68 | NCF2 | 16924 | -0.210 | -0.4881 | No | ||
| 69 | ZEB1 | 17252 | -0.244 | -0.5005 | No | ||
| 70 | SQSTM1 | 17662 | -0.293 | -0.5160 | No | ||
| 71 | DNAAF1 | 17931 | -0.326 | -0.5261 | No | ||
| 72 | BACH1 | 19119 | -0.539 | -0.5713 | No | ||
| 73 | FOXA1 | 19191 | -0.552 | -0.5738 | No | ||
| 74 | ARID5B | 19772 | -0.706 | -0.5957 | No | ||
| 75 | DNAJB4 | 19862 | -0.738 | -0.5989 | No | ||
| 76 | CCPG1 | 19990 | -0.782 | -0.6035 | No | ||
| 77 | MAFF | 20369 | -0.911 | -0.6176 | No | ||
| 78 | FTH1 | 20690 | -1.038 | -0.6295 | No | ||
| 79 | TMCO3 | 21044 | -1.216 | -0.6426 | No | ||
| 80 | GCLM | 21297 | -1.363 | -0.6518 | No | ||
| 81 | HSPA13 | 21655 | -1.583 | -0.6649 | No | ||
| 82 | CYTIP | 21723 | -1.636 | -0.6669 | No | ||
| 83 | IL23A | 21785 | -1.672 | -0.6687 | No | ||
| 84 | DUSP1 | 21853 | -1.729 | -0.6707 | No | ||
| 85 | GABARAPL1 | 22236 | -2.065 | -0.6846 | No | ||
| 86 | MBNL2 | 22372 | -2.193 | -0.6890 | No | ||
| 87 | CD44 | 22790 | -2.677 | -0.7040 | No | ||
| 88 | RGS2 | 22796 | -2.685 | -0.7034 | No | ||
| 89 | AKR1C2 | 22988 | -2.925 | -0.7097 | No | ||
| 90 | FNDC3A | 23128 | -3.121 | -0.7140 | No | ||
| 91 | TMEM140 | 23292 | -3.401 | -0.7191 | No | ||
| 92 | KLF6 | 23308 | -3.435 | -0.7185 | No | ||
| 93 | ID2 | 23493 | -3.773 | -0.7243 | No | ||
| 94 | CSGALNACT2 | 23739 | -4.328 | -0.7322 | No | ||
| 95 | ZNF267 | 24029 | -5.126 | -0.7416 | No | ||
| 96 | MXI1 | 24115 | -5.336 | -0.7431 | No | ||
| 97 | AKR1C3 | 24117 | -5.347 | -0.7413 | No | ||
| 98 | WDR26 | 24144 | -5.429 | -0.7405 | No | ||
| 99 | NEK1 | 24253 | -5.746 | -0.7428 | No | ||
| 100 | STX3 | 24269 | -5.844 | -0.7414 | No | ||
| 101 | PPP1R15A | 24307 | -6.013 | -0.7408 | No | ||
| 102 | CLK1 | 24334 | -6.145 | -0.7398 | No | ||
| 103 | MFAP4 | 24652 | -7.457 | -0.7494 | No | ||
| 104 | FYN | 24883 | -8.790 | -0.7553 | Yes | ||
| 105 | MARS | 24894 | -8.832 | -0.7528 | Yes | ||
| 106 | GAB2 | 24911 | -8.991 | -0.7504 | Yes | ||
| 107 | IFNGR1 | 24935 | -9.155 | -0.7483 | Yes | ||
| 108 | ASF1A | 25064 | -10.253 | -0.7498 | Yes | ||
| 109 | SLC3A2 | 25078 | -10.370 | -0.7469 | Yes | ||
| 110 | DUSP5 | 25225 | -11.830 | -0.7485 | Yes | ||
| 111 | JUN | 25262 | -12.127 | -0.7459 | Yes | ||
| 112 | TMEM57 | 25325 | -12.934 | -0.7440 | Yes | ||
| 113 | RND3 | 25387 | -13.780 | -0.7418 | Yes | ||
| 114 | ZNF215 | 25437 | -14.475 | -0.7389 | Yes | ||
| 115 | CARS | 25844 | -24.758 | -0.7462 | Yes | ||
| 116 | PALLD | 25978 | -32.108 | -0.7407 | Yes | ||
| 117 | FAM129A | 25984 | -32.982 | -0.7300 | Yes | ||
| 118 | CD55 | 25987 | -33.140 | -0.7192 | Yes | ||
| 119 | NFE2L1 | 26006 | -34.437 | -0.7085 | Yes | ||
| 120 | IGFBP3 | 26019 | -35.792 | -0.6971 | Yes | ||
| 121 | TES | 26033 | -37.078 | -0.6854 | Yes | ||
| 122 | YPEL5 | 26050 | -38.933 | -0.6732 | Yes | ||
| 123 | PMAIP1 | 26053 | -38.997 | -0.6604 | Yes | ||
| 124 | SLC7A11 | 26126 | -47.089 | -0.6476 | Yes | ||
| 125 | PHLDA1 | 26170 | -55.684 | -0.6309 | Yes | ||
| 126 | CEBPG | 26210 | -66.827 | -0.6103 | Yes | ||
| 127 | PSAT1 | 26230 | -76.008 | -0.5859 | Yes | ||
| 128 | ASNS | 26241 | -83.978 | -0.5586 | Yes | ||
| 129 | CEBPB | 26244 | -86.643 | -0.5301 | Yes | ||
| 130 | GARS | 26249 | -89.806 | -0.5006 | Yes | ||
| 131 | VEGFA | 26251 | -89.987 | -0.4710 | Yes | ||
| 132 | DDIT3 | 26291 | -176.939 | -0.4141 | Yes | ||
| 133 | ATF3 | 26295 | -192.665 | -0.3506 | Yes | ||
| 134 | IFRD1 | 26303 | -230.785 | -0.2747 | Yes | ||
| 135 | TRIB3 | 26304 | -235.323 | -0.1970 | Yes | ||
| 136 | DDIT4 | 26311 | -299.207 | -0.0985 | Yes | ||
| 137 | CHAC1 | 26314 | -299.207 | 0.0001 | Yes |