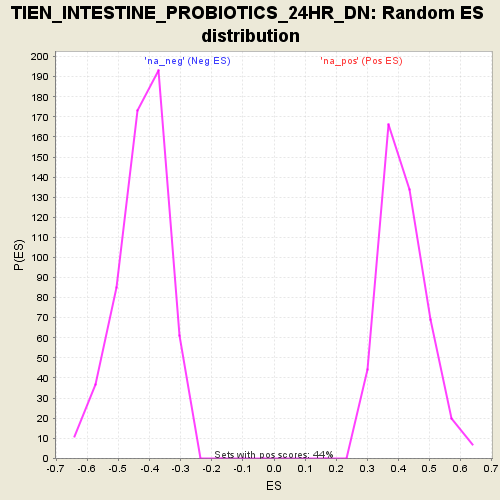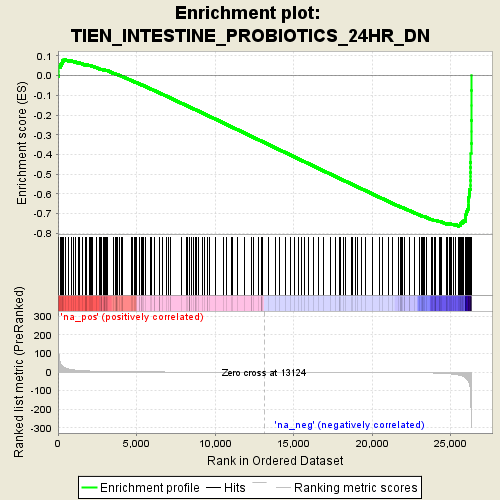
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | TIEN_INTESTINE_PROBIOTICS_24HR_DN |
| Enrichment Score (ES) | -0.7640212 |
| Normalized Enrichment Score (NES) | -1.7978002 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.020581309 |
| FWER p-Value | 0.098 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC6A8 | 55 | 94.569 | 0.0217 | No | ||
| 2 | HSPA8 | 56 | 93.481 | 0.0453 | No | ||
| 3 | HSPA5 | 151 | 52.337 | 0.0549 | No | ||
| 4 | PDIA4 | 222 | 37.994 | 0.0618 | No | ||
| 5 | CALR | 266 | 32.957 | 0.0684 | No | ||
| 6 | GDF15 | 292 | 30.340 | 0.0751 | No | ||
| 7 | SDF2L1 | 350 | 26.338 | 0.0796 | No | ||
| 8 | MANF | 448 | 22.191 | 0.0814 | No | ||
| 9 | MLPH | 677 | 16.829 | 0.0769 | No | ||
| 10 | RNF19B | 823 | 14.447 | 0.0750 | No | ||
| 11 | ACTG1 | 962 | 12.546 | 0.0729 | No | ||
| 12 | PPIB | 1084 | 11.233 | 0.0711 | No | ||
| 13 | CDK2AP2 | 1268 | 9.840 | 0.0666 | No | ||
| 14 | HSPH1 | 1350 | 9.216 | 0.0658 | No | ||
| 15 | EIF4B | 1567 | 7.940 | 0.0595 | No | ||
| 16 | SLC35E1 | 1722 | 7.166 | 0.0554 | No | ||
| 17 | UNC5B | 1769 | 6.981 | 0.0554 | No | ||
| 18 | RPL9 | 1788 | 6.888 | 0.0565 | No | ||
| 19 | HYOU1 | 1806 | 6.782 | 0.0575 | No | ||
| 20 | PDIA6 | 1980 | 6.116 | 0.0525 | No | ||
| 21 | RPS14 | 2047 | 5.896 | 0.0514 | No | ||
| 22 | NUCB2 | 2136 | 5.615 | 0.0495 | No | ||
| 23 | RPS3 | 2200 | 5.404 | 0.0484 | No | ||
| 24 | FAM120A | 2445 | 4.736 | 0.0403 | No | ||
| 25 | RPL23A | 2661 | 4.286 | 0.0331 | No | ||
| 26 | RPS24 | 2715 | 4.152 | 0.0321 | No | ||
| 27 | DNAJB9 | 2733 | 4.122 | 0.0325 | No | ||
| 28 | RPL23 | 2742 | 4.105 | 0.0332 | No | ||
| 29 | EEF2 | 2774 | 4.032 | 0.0331 | No | ||
| 30 | FTL | 2919 | 3.762 | 0.0285 | No | ||
| 31 | PDIA3 | 2945 | 3.710 | 0.0285 | No | ||
| 32 | CDC5L | 2965 | 3.669 | 0.0287 | No | ||
| 33 | RPL34 | 2966 | 3.668 | 0.0296 | No | ||
| 34 | TMED2 | 3029 | 3.568 | 0.0281 | No | ||
| 35 | P4HB | 3059 | 3.515 | 0.0279 | No | ||
| 36 | AFTPH | 3084 | 3.473 | 0.0278 | No | ||
| 37 | HSP90B1 | 3151 | 3.351 | 0.0262 | No | ||
| 38 | EPHB4 | 3507 | 2.853 | 0.0133 | No | ||
| 39 | TRAM1 | 3681 | 2.671 | 0.0073 | No | ||
| 40 | FICD | 3705 | 2.632 | 0.0071 | No | ||
| 41 | EIF5A | 3712 | 2.624 | 0.0075 | No | ||
| 42 | SGSM3 | 3719 | 2.613 | 0.0080 | No | ||
| 43 | RPS27 | 3760 | 2.568 | 0.0071 | No | ||
| 44 | PNRC1 | 3888 | 2.423 | 0.0028 | No | ||
| 45 | RPS23 | 4068 | 2.247 | -0.0035 | No | ||
| 46 | RPL39 | 4087 | 2.231 | -0.0036 | No | ||
| 47 | RPL3 | 4096 | 2.226 | -0.0033 | No | ||
| 48 | EEF1A1 | 4648 | 1.739 | -0.0240 | No | ||
| 49 | RPS18 | 4759 | 1.649 | -0.0278 | No | ||
| 50 | PABPC1 | 4895 | 1.550 | -0.0326 | No | ||
| 51 | TMED7 | 4927 | 1.528 | -0.0334 | No | ||
| 52 | RPL5 | 4999 | 1.476 | -0.0357 | No | ||
| 53 | ST6GALNAC4 | 5203 | 1.352 | -0.0432 | No | ||
| 54 | RPL13A | 5204 | 1.351 | -0.0428 | No | ||
| 55 | RPL37A | 5298 | 1.297 | -0.0460 | No | ||
| 56 | RPS19 | 5342 | 1.269 | -0.0474 | No | ||
| 57 | RPS2 | 5348 | 1.266 | -0.0472 | No | ||
| 58 | RPS6 | 5456 | 1.206 | -0.0510 | No | ||
| 59 | RPL41 | 5567 | 1.139 | -0.0550 | No | ||
| 60 | HKDC1 | 5884 | 0.975 | -0.0668 | No | ||
| 61 | ARHGAP35 | 5969 | 0.942 | -0.0698 | No | ||
| 62 | SEL1L | 6132 | 0.885 | -0.0758 | No | ||
| 63 | RPS7 | 6153 | 0.875 | -0.0763 | No | ||
| 64 | KLHL24 | 6453 | 0.777 | -0.0876 | No | ||
| 65 | RPL13 | 6459 | 0.775 | -0.0876 | No | ||
| 66 | MAN1A2 | 6485 | 0.767 | -0.0883 | No | ||
| 67 | RPS3A | 6650 | 0.709 | -0.0944 | No | ||
| 68 | KLF5 | 6883 | 0.637 | -0.1032 | No | ||
| 69 | RPS11 | 7027 | 0.595 | -0.1085 | No | ||
| 70 | RARRES3 | 7145 | 0.565 | -0.1128 | No | ||
| 71 | ARIH1 | 7152 | 0.562 | -0.1129 | No | ||
| 72 | TXNRD1 | 7886 | 0.415 | -0.1409 | No | ||
| 73 | ARF4 | 8150 | 0.374 | -0.1509 | No | ||
| 74 | HUWE1 | 8212 | 0.366 | -0.1531 | No | ||
| 75 | NDRG1 | 8342 | 0.347 | -0.1580 | No | ||
| 76 | CANX | 8348 | 0.345 | -0.1581 | No | ||
| 77 | LAMP3 | 8359 | 0.344 | -0.1584 | No | ||
| 78 | DNAJC10 | 8399 | 0.338 | -0.1598 | No | ||
| 79 | MTFR1 | 8479 | 0.324 | -0.1627 | No | ||
| 80 | SEL1L3 | 8595 | 0.310 | -0.1671 | No | ||
| 81 | MARCH5 | 8727 | 0.293 | -0.1720 | No | ||
| 82 | ACBD3 | 8774 | 0.287 | -0.1737 | No | ||
| 83 | UBC | 8798 | 0.286 | -0.1745 | No | ||
| 84 | VASP | 8943 | 0.267 | -0.1799 | No | ||
| 85 | RPLP0 | 9178 | 0.242 | -0.1888 | No | ||
| 86 | BTG1 | 9324 | 0.224 | -0.1943 | No | ||
| 87 | NQO1 | 9499 | 0.207 | -0.2010 | No | ||
| 88 | FAF2 | 9525 | 0.205 | -0.2019 | No | ||
| 89 | TPT1 | 9669 | 0.190 | -0.2073 | No | ||
| 90 | PRR15L | 10040 | 0.159 | -0.2214 | No | ||
| 91 | UBXN4 | 10563 | 0.121 | -0.2414 | No | ||
| 92 | CREB1 | 10744 | 0.107 | -0.2482 | No | ||
| 93 | STX5 | 11016 | 0.091 | -0.2586 | No | ||
| 94 | RPL27A | 11079 | 0.088 | -0.2610 | No | ||
| 95 | TBC1D15 | 11445 | 0.068 | -0.2749 | No | ||
| 96 | ELF3 | 11880 | 0.048 | -0.2915 | No | ||
| 97 | RAB2A | 12297 | 0.031 | -0.3075 | No | ||
| 98 | GCNT3 | 12322 | 0.030 | -0.3084 | No | ||
| 99 | KAT6B | 12427 | 0.025 | -0.3123 | No | ||
| 100 | SAT1 | 12782 | 0.011 | -0.3259 | No | ||
| 101 | MAP2K2 | 12922 | 0.007 | -0.3312 | No | ||
| 102 | DTX2P1-UPK3BP1-PMS2P11 | 12973 | 0.005 | -0.3331 | No | ||
| 103 | PJA2 | 12983 | 0.005 | -0.3335 | No | ||
| 104 | ARMCX3 | 13026 | 0.005 | -0.3351 | No | ||
| 105 | RPL12 | 13412 | -0.009 | -0.3498 | No | ||
| 106 | S100P | 13838 | -0.025 | -0.3661 | No | ||
| 107 | CDC6 | 14100 | -0.035 | -0.3761 | No | ||
| 108 | TMED5 | 14484 | -0.051 | -0.3907 | No | ||
| 109 | PTPRF | 14780 | -0.061 | -0.4020 | No | ||
| 110 | NBR1 | 15081 | -0.078 | -0.4135 | No | ||
| 111 | ZMYM4 | 15280 | -0.089 | -0.4211 | No | ||
| 112 | LAMB3 | 15522 | -0.104 | -0.4303 | No | ||
| 113 | ATF2 | 15697 | -0.113 | -0.4369 | No | ||
| 114 | GLRX | 15957 | -0.131 | -0.4468 | No | ||
| 115 | AUH | 16242 | -0.151 | -0.4576 | No | ||
| 116 | RPL4 | 16586 | -0.178 | -0.4707 | No | ||
| 117 | ESRP1 | 16926 | -0.210 | -0.4836 | No | ||
| 118 | RPL10 | 17319 | -0.251 | -0.4986 | No | ||
| 119 | SQSTM1 | 17662 | -0.293 | -0.5116 | No | ||
| 120 | KDELR1 | 17905 | -0.323 | -0.5208 | No | ||
| 121 | NUDT4 | 17928 | -0.325 | -0.5216 | No | ||
| 122 | GFPT1 | 17970 | -0.331 | -0.5230 | No | ||
| 123 | MCTP2 | 18206 | -0.364 | -0.5320 | No | ||
| 124 | LRRFIP1 | 18317 | -0.384 | -0.5361 | No | ||
| 125 | TMEM39A | 18665 | -0.448 | -0.5493 | No | ||
| 126 | MIA3 | 18750 | -0.464 | -0.5524 | No | ||
| 127 | WDR45 | 18971 | -0.509 | -0.5606 | No | ||
| 128 | ZNF236 | 19083 | -0.530 | -0.5648 | No | ||
| 129 | AKR1C1 | 19331 | -0.586 | -0.5741 | No | ||
| 130 | TANK | 19607 | -0.661 | -0.5844 | No | ||
| 131 | TSPAN31 | 20016 | -0.787 | -0.5999 | No | ||
| 132 | ZNF282 | 20464 | -0.947 | -0.6168 | No | ||
| 133 | FTH1 | 20690 | -1.038 | -0.6251 | No | ||
| 134 | TMCO3 | 21044 | -1.216 | -0.6383 | No | ||
| 135 | GCLM | 21297 | -1.363 | -0.6476 | No | ||
| 136 | HSPA13 | 21655 | -1.583 | -0.6609 | No | ||
| 137 | NCOR1 | 21776 | -1.668 | -0.6651 | No | ||
| 138 | TFG | 21822 | -1.705 | -0.6664 | No | ||
| 139 | THOC2 | 21860 | -1.733 | -0.6674 | No | ||
| 140 | TCEA1 | 21870 | -1.744 | -0.6673 | No | ||
| 141 | PDE8A | 21931 | -1.797 | -0.6691 | No | ||
| 142 | MTA1 | 22049 | -1.903 | -0.6731 | No | ||
| 143 | PDCD4 | 22066 | -1.916 | -0.6732 | No | ||
| 144 | BNIP3L | 22352 | -2.182 | -0.6836 | No | ||
| 145 | MBNL2 | 22372 | -2.193 | -0.6838 | No | ||
| 146 | HOOK1 | 22715 | -2.579 | -0.6962 | No | ||
| 147 | AKR1C2 | 22988 | -2.925 | -0.7059 | No | ||
| 148 | FNDC3A | 23128 | -3.121 | -0.7104 | No | ||
| 149 | PRSS3 | 23213 | -3.262 | -0.7128 | No | ||
| 150 | LPIN2 | 23237 | -3.313 | -0.7129 | No | ||
| 151 | SERP1 | 23293 | -3.403 | -0.7141 | No | ||
| 152 | GATAD1 | 23362 | -3.537 | -0.7158 | No | ||
| 153 | ZNF277 | 23469 | -3.733 | -0.7190 | No | ||
| 154 | HSPA9 | 23764 | -4.390 | -0.7291 | No | ||
| 155 | CSNK1A1 | 23817 | -4.510 | -0.7300 | No | ||
| 156 | NFIL3 | 24002 | -5.017 | -0.7358 | No | ||
| 157 | GOT1 | 24030 | -5.126 | -0.7355 | No | ||
| 158 | GOLGA5 | 24046 | -5.172 | -0.7348 | No | ||
| 159 | ADNP2 | 24059 | -5.207 | -0.7339 | No | ||
| 160 | SEC63 | 24263 | -5.803 | -0.7402 | No | ||
| 161 | PPP1R15A | 24307 | -6.013 | -0.7404 | No | ||
| 162 | CLCN3 | 24325 | -6.099 | -0.7395 | No | ||
| 163 | MTDH | 24386 | -6.334 | -0.7402 | No | ||
| 164 | ALDH3B1 | 24749 | -7.958 | -0.7520 | No | ||
| 165 | RARA | 24765 | -8.044 | -0.7506 | No | ||
| 166 | COG5 | 24826 | -8.358 | -0.7508 | No | ||
| 167 | SH3GLB1 | 24930 | -9.123 | -0.7524 | No | ||
| 168 | CEP57 | 24978 | -9.586 | -0.7518 | No | ||
| 169 | SLC3A2 | 25078 | -10.370 | -0.7530 | No | ||
| 170 | RMND5B | 25196 | -11.549 | -0.7546 | No | ||
| 171 | RANBP9 | 25280 | -12.443 | -0.7546 | No | ||
| 172 | PRNP | 25527 | -15.875 | -0.7600 | Yes | ||
| 173 | WIPI1 | 25583 | -17.243 | -0.7578 | Yes | ||
| 174 | TTC3 | 25625 | -18.237 | -0.7548 | Yes | ||
| 175 | LARP6 | 25627 | -18.258 | -0.7502 | Yes | ||
| 176 | JAG1 | 25675 | -19.325 | -0.7471 | Yes | ||
| 177 | EIF1 | 25731 | -20.857 | -0.7440 | Yes | ||
| 178 | TFE3 | 25759 | -21.664 | -0.7395 | Yes | ||
| 179 | UBE2H | 25847 | -25.130 | -0.7365 | Yes | ||
| 180 | CBX4 | 25927 | -29.298 | -0.7322 | Yes | ||
| 181 | GRB10 | 25941 | -30.158 | -0.7251 | Yes | ||
| 182 | TTC17 | 25965 | -31.483 | -0.7180 | Yes | ||
| 183 | ARHGEF2 | 25969 | -31.712 | -0.7102 | Yes | ||
| 184 | C6orf48 | 25975 | -31.926 | -0.7023 | Yes | ||
| 185 | CD55 | 25987 | -33.140 | -0.6944 | Yes | ||
| 186 | TES | 26033 | -37.078 | -0.6868 | Yes | ||
| 187 | SLC1A5 | 26086 | -42.020 | -0.6782 | Yes | ||
| 188 | SLC7A1 | 26112 | -45.465 | -0.6677 | Yes | ||
| 189 | WARS | 26117 | -46.483 | -0.6561 | Yes | ||
| 190 | AARS | 26130 | -48.060 | -0.6444 | Yes | ||
| 191 | SLC1A4 | 26133 | -48.490 | -0.6323 | Yes | ||
| 192 | HERPUD1 | 26159 | -53.015 | -0.6199 | Yes | ||
| 193 | PHLDA1 | 26170 | -55.684 | -0.6063 | Yes | ||
| 194 | SARS | 26192 | -60.122 | -0.5919 | Yes | ||
| 195 | CEBPG | 26210 | -66.827 | -0.5757 | Yes | ||
| 196 | ASNS | 26241 | -83.978 | -0.5557 | Yes | ||
| 197 | CEBPB | 26244 | -86.643 | -0.5340 | Yes | ||
| 198 | GARS | 26249 | -89.806 | -0.5115 | Yes | ||
| 199 | VEGFA | 26251 | -89.987 | -0.4888 | Yes | ||
| 200 | MTHFD2 | 26257 | -95.222 | -0.4650 | Yes | ||
| 201 | SLC43A1 | 26263 | -98.209 | -0.4405 | Yes | ||
| 202 | ATF3 | 26295 | -192.665 | -0.3931 | Yes | ||
| 203 | PCK2 | 26298 | -198.705 | -0.3431 | Yes | ||
| 204 | IFRD1 | 26303 | -230.785 | -0.2851 | Yes | ||
| 205 | TRIB3 | 26304 | -235.323 | -0.2258 | Yes | ||
| 206 | DDIT4 | 26311 | -299.207 | -0.1506 | Yes | ||
| 207 | INHBE | 26312 | -299.207 | -0.0752 | Yes | ||
| 208 | H1F0 | 26316 | -299.207 | 0.0000 | Yes |