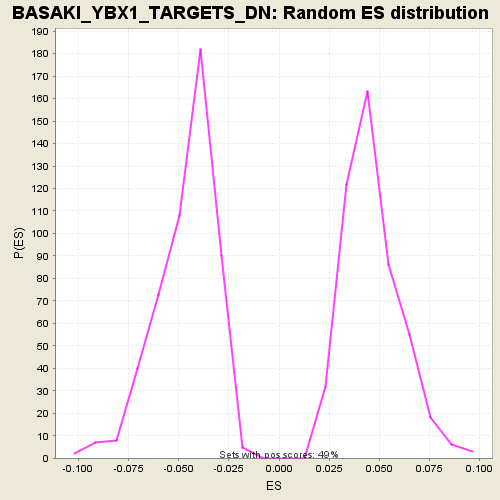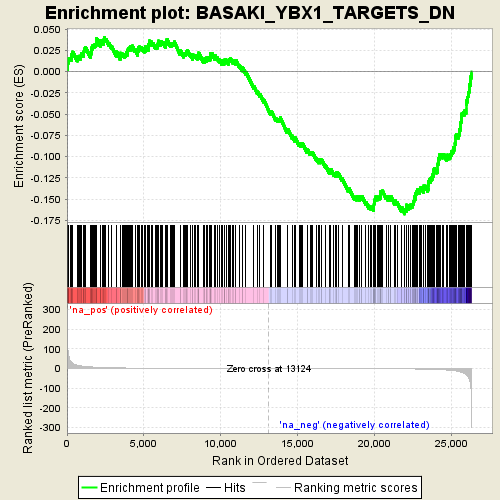
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | BASAKI_YBX1_TARGETS_DN |
| Enrichment Score (ES) | -0.16750826 |
| Normalized Enrichment Score (NES) | -3.610688 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SPP1 | 0 | 299.207 | 0.0028 | No | ||
| 2 | ARRB1 | 20 | 152.207 | 0.0050 | No | ||
| 3 | IFITM1 | 30 | 118.511 | 0.0075 | No | ||
| 4 | STMN3 | 46 | 101.306 | 0.0097 | No | ||
| 5 | LAMB1 | 65 | 88.674 | 0.0119 | No | ||
| 6 | TP53INP1 | 107 | 64.094 | 0.0132 | No | ||
| 7 | HLA-DMA | 118 | 61.525 | 0.0156 | No | ||
| 8 | COBLL1 | 192 | 41.878 | 0.0157 | No | ||
| 9 | NFKBIZ | 270 | 32.545 | 0.0156 | No | ||
| 10 | GDF15 | 292 | 30.340 | 0.0176 | No | ||
| 11 | PNP | 301 | 29.501 | 0.0201 | No | ||
| 12 | ALDOC | 340 | 26.807 | 0.0215 | No | ||
| 13 | H19 | 373 | 25.247 | 0.0231 | No | ||
| 14 | BTN3A2 | 653 | 17.260 | 0.0152 | No | ||
| 15 | MZF1 | 707 | 16.196 | 0.0160 | No | ||
| 16 | STC1 | 740 | 15.599 | 0.0177 | No | ||
| 17 | MMP24 | 796 | 14.910 | 0.0184 | No | ||
| 18 | EGR1 | 901 | 13.382 | 0.0172 | No | ||
| 19 | PFKFB3 | 905 | 13.335 | 0.0200 | No | ||
| 20 | DHRS3 | 933 | 12.896 | 0.0218 | No | ||
| 21 | PTGS1 | 1053 | 11.573 | 0.0200 | No | ||
| 22 | MARCKSL1 | 1056 | 11.539 | 0.0228 | No | ||
| 23 | PDK1 | 1096 | 11.129 | 0.0242 | No | ||
| 24 | SCD | 1102 | 11.098 | 0.0268 | No | ||
| 25 | SLC20A1 | 1200 | 10.407 | 0.0259 | No | ||
| 26 | UCP2 | 1204 | 10.383 | 0.0287 | No | ||
| 27 | IRF2BP2 | 1533 | 8.166 | 0.0189 | No | ||
| 28 | SPRY1 | 1543 | 8.095 | 0.0214 | No | ||
| 29 | TMEM47 | 1564 | 7.957 | 0.0235 | No | ||
| 30 | SSBP4 | 1596 | 7.768 | 0.0251 | No | ||
| 31 | ZNF207 | 1597 | 7.765 | 0.0280 | No | ||
| 32 | TFPI | 1640 | 7.533 | 0.0292 | No | ||
| 33 | PSD3 | 1678 | 7.385 | 0.0306 | No | ||
| 34 | BICD1 | 1745 | 7.067 | 0.0309 | No | ||
| 35 | MEGF9 | 1805 | 6.783 | 0.0315 | No | ||
| 36 | PROS1 | 1866 | 6.553 | 0.0321 | No | ||
| 37 | SF1 | 1892 | 6.431 | 0.0339 | No | ||
| 38 | DCAF16 | 1903 | 6.384 | 0.0364 | No | ||
| 39 | NAV1 | 1906 | 6.374 | 0.0392 | No | ||
| 40 | PTP4A1 | 2168 | 5.519 | 0.0320 | No | ||
| 41 | KDM4B | 2178 | 5.478 | 0.0345 | No | ||
| 42 | RBMXL1 | 2188 | 5.437 | 0.0370 | No | ||
| 43 | NEBL | 2306 | 5.090 | 0.0353 | No | ||
| 44 | WDR1 | 2395 | 4.857 | 0.0348 | No | ||
| 45 | NEAT1 | 2398 | 4.852 | 0.0376 | No | ||
| 46 | SCIN | 2405 | 4.834 | 0.0402 | No | ||
| 47 | MMAA | 2529 | 4.576 | 0.0383 | No | ||
| 48 | NAA38 | 2723 | 4.142 | 0.0337 | No | ||
| 49 | MEIS2 | 2903 | 3.781 | 0.0297 | No | ||
| 50 | NKTR | 3207 | 3.264 | 0.0208 | No | ||
| 51 | TARSL2 | 3214 | 3.255 | 0.0235 | No | ||
| 52 | NADSYN1 | 3448 | 2.934 | 0.0173 | No | ||
| 53 | CARD8 | 3461 | 2.912 | 0.0197 | No | ||
| 54 | CKB | 3479 | 2.888 | 0.0219 | No | ||
| 55 | IBTK | 3581 | 2.764 | 0.0209 | No | ||
| 56 | TNFRSF11A | 3675 | 2.678 | 0.0201 | No | ||
| 57 | MRPL19 | 3765 | 2.564 | 0.0196 | No | ||
| 58 | MGAT4A | 3815 | 2.513 | 0.0205 | No | ||
| 59 | CA12 | 3833 | 2.489 | 0.0227 | No | ||
| 60 | TM2D3 | 3922 | 2.390 | 0.0222 | No | ||
| 61 | BPTF | 3933 | 2.383 | 0.0246 | No | ||
| 62 | JAK3 | 3959 | 2.357 | 0.0265 | No | ||
| 63 | ZNF320 | 4013 | 2.303 | 0.0273 | No | ||
| 64 | ANKRD36 | 4072 | 2.243 | 0.0279 | No | ||
| 65 | RPL31 | 4095 | 2.227 | 0.0299 | No | ||
| 66 | DNAJC3 | 4217 | 2.086 | 0.0281 | No | ||
| 67 | PLOD2 | 4236 | 2.069 | 0.0303 | No | ||
| 68 | GOLGA2P5 | 4423 | 1.912 | 0.0260 | No | ||
| 69 | FAM3C | 4604 | 1.771 | 0.0219 | No | ||
| 70 | CDK12 | 4611 | 1.768 | 0.0245 | No | ||
| 71 | FAM43A | 4643 | 1.745 | 0.0262 | No | ||
| 72 | EEF1A1 | 4648 | 1.739 | 0.0289 | No | ||
| 73 | SEMA4B | 4715 | 1.681 | 0.0292 | No | ||
| 74 | H3F3B | 4816 | 1.613 | 0.0282 | No | ||
| 75 | SF3B1 | 4902 | 1.545 | 0.0277 | No | ||
| 76 | BDH2 | 5029 | 1.452 | 0.0257 | No | ||
| 77 | PDS5B | 5078 | 1.420 | 0.0267 | No | ||
| 78 | SPRY2 | 5085 | 1.415 | 0.0294 | No | ||
| 79 | ST6GALNAC4 | 5203 | 1.352 | 0.0277 | No | ||
| 80 | MIB1 | 5276 | 1.315 | 0.0278 | No | ||
| 81 | RPL37 | 5305 | 1.290 | 0.0296 | No | ||
| 82 | ANKRD50 | 5309 | 1.287 | 0.0323 | No | ||
| 83 | ZNF397 | 5344 | 1.269 | 0.0338 | No | ||
| 84 | VMP1 | 5358 | 1.255 | 0.0362 | No | ||
| 85 | SEPT6 | 5461 | 1.203 | 0.0351 | No | ||
| 86 | MPZL2 | 5579 | 1.131 | 0.0334 | No | ||
| 87 | ABCA5 | 5742 | 1.046 | 0.0300 | No | ||
| 88 | CTBS | 5821 | 1.005 | 0.0299 | No | ||
| 89 | IER5L | 5881 | 0.976 | 0.0305 | No | ||
| 90 | KCTD6 | 5903 | 0.969 | 0.0325 | No | ||
| 91 | CUL4B | 5933 | 0.956 | 0.0342 | No | ||
| 92 | GAL3ST1 | 5959 | 0.946 | 0.0361 | No | ||
| 93 | TM9SF3 | 6102 | 0.894 | 0.0335 | No | ||
| 94 | GBE1 | 6131 | 0.886 | 0.0353 | No | ||
| 95 | TNFSF10 | 6229 | 0.844 | 0.0344 | No | ||
| 96 | IFI27 | 6400 | 0.792 | 0.0307 | No | ||
| 97 | GRB7 | 6411 | 0.789 | 0.0332 | No | ||
| 98 | C11orf58 | 6442 | 0.781 | 0.0349 | No | ||
| 99 | KLHL24 | 6453 | 0.777 | 0.0373 | No | ||
| 100 | SEZ6L2 | 6507 | 0.759 | 0.0381 | No | ||
| 101 | FAM110C | 6746 | 0.672 | 0.0318 | No | ||
| 102 | BTN3A3 | 6801 | 0.659 | 0.0326 | No | ||
| 103 | RAP1GAP | 6862 | 0.641 | 0.0331 | No | ||
| 104 | TFDP2 | 6940 | 0.619 | 0.0330 | No | ||
| 105 | VPS13B | 6961 | 0.615 | 0.0351 | No | ||
| 106 | ARHGAP18 | 7361 | 0.516 | 0.0226 | No | ||
| 107 | GAS2L3 | 7366 | 0.515 | 0.0253 | No | ||
| 108 | NCALD | 7581 | 0.467 | 0.0199 | No | ||
| 109 | GSTM3 | 7613 | 0.460 | 0.0215 | No | ||
| 110 | SBF2 | 7689 | 0.449 | 0.0215 | No | ||
| 111 | SLC25A27 | 7734 | 0.439 | 0.0226 | No | ||
| 112 | VPS13A | 7788 | 0.429 | 0.0234 | No | ||
| 113 | WSB1 | 7827 | 0.423 | 0.0248 | No | ||
| 114 | SLC40A1 | 8024 | 0.392 | 0.0201 | No | ||
| 115 | GPD1L | 8177 | 0.370 | 0.0171 | No | ||
| 116 | PNISR | 8184 | 0.369 | 0.0197 | No | ||
| 117 | ZC3H11A | 8265 | 0.360 | 0.0195 | No | ||
| 118 | NDRG1 | 8342 | 0.347 | 0.0194 | No | ||
| 119 | ZNF253 | 8469 | 0.326 | 0.0174 | No | ||
| 120 | FOS | 8533 | 0.318 | 0.0179 | No | ||
| 121 | HECW2 | 8538 | 0.318 | 0.0205 | No | ||
| 122 | PCSK6 | 8575 | 0.313 | 0.0220 | No | ||
| 123 | KCNK3 | 8877 | 0.274 | 0.0133 | No | ||
| 124 | JUNB | 8956 | 0.266 | 0.0131 | No | ||
| 125 | MEIS1 | 8963 | 0.265 | 0.0157 | No | ||
| 126 | FLRT3 | 9066 | 0.252 | 0.0147 | No | ||
| 127 | CRYM | 9092 | 0.250 | 0.0165 | No | ||
| 128 | SLC45A3 | 9154 | 0.244 | 0.0170 | No | ||
| 129 | MBOAT2 | 9265 | 0.231 | 0.0157 | No | ||
| 130 | NET1 | 9318 | 0.225 | 0.0165 | No | ||
| 131 | BTG1 | 9324 | 0.224 | 0.0192 | No | ||
| 132 | ANKRD29 | 9329 | 0.223 | 0.0218 | No | ||
| 133 | SNORA28 | 9415 | 0.215 | 0.0214 | No | ||
| 134 | TCIRG1 | 9617 | 0.195 | 0.0165 | No | ||
| 135 | SRSF7 | 9632 | 0.194 | 0.0188 | No | ||
| 136 | STXBP3 | 9770 | 0.180 | 0.0164 | No | ||
| 137 | PGK1 | 9909 | 0.170 | 0.0140 | No | ||
| 138 | SLC44A1 | 10066 | 0.157 | 0.0108 | No | ||
| 139 | CLINT1 | 10077 | 0.155 | 0.0133 | No | ||
| 140 | TRIM13 | 10206 | 0.143 | 0.0112 | No | ||
| 141 | HELZ | 10252 | 0.140 | 0.0123 | No | ||
| 142 | XIST | 10267 | 0.139 | 0.0146 | No | ||
| 143 | SCAMP1 | 10366 | 0.132 | 0.0137 | No | ||
| 144 | MED6 | 10498 | 0.124 | 0.0115 | No | ||
| 145 | LCN2 | 10527 | 0.123 | 0.0133 | No | ||
| 146 | UBXN4 | 10563 | 0.121 | 0.0148 | No | ||
| 147 | WIPF2 | 10622 | 0.116 | 0.0154 | No | ||
| 148 | FAM83A | 10762 | 0.106 | 0.0129 | No | ||
| 149 | ISG20 | 10851 | 0.100 | 0.0123 | No | ||
| 150 | GALNT3 | 10948 | 0.095 | 0.0115 | No | ||
| 151 | FAM162A | 10971 | 0.094 | 0.0135 | No | ||
| 152 | PKD2 | 11205 | 0.082 | 0.0074 | No | ||
| 153 | ACER3 | 11380 | 0.071 | 0.0035 | No | ||
| 154 | NT5DC1 | 11415 | 0.069 | 0.0050 | No | ||
| 155 | PLEKHA1 | 11631 | 0.059 | -0.0004 | No | ||
| 156 | C5orf24 | 12142 | 0.038 | -0.0172 | No | ||
| 157 | ANKRD36B | 12367 | 0.028 | -0.0230 | No | ||
| 158 | TTLL3 | 12534 | 0.021 | -0.0265 | No | ||
| 159 | SAT1 | 12782 | 0.011 | -0.0332 | No | ||
| 160 | BCKDHA | 13235 | -0.004 | -0.0477 | No | ||
| 161 | SGK2 | 13274 | -0.005 | -0.0463 | No | ||
| 162 | ATP2C1 | 13578 | -0.015 | -0.0552 | No | ||
| 163 | ZNF626 | 13672 | -0.019 | -0.0559 | No | ||
| 164 | ATRX | 13745 | -0.022 | -0.0558 | No | ||
| 165 | BRWD1 | 13826 | -0.025 | -0.0560 | No | ||
| 166 | C14orf28 | 13851 | -0.026 | -0.0541 | No | ||
| 167 | HDAC4 | 14311 | -0.045 | -0.0689 | No | ||
| 168 | SLC3A1 | 14354 | -0.046 | -0.0677 | No | ||
| 169 | RRAGD | 14635 | -0.056 | -0.0756 | No | ||
| 170 | CD302 | 14800 | -0.062 | -0.0791 | No | ||
| 171 | RFX3 | 14842 | -0.063 | -0.0778 | No | ||
| 172 | BAZ2B | 15086 | -0.078 | -0.0844 | No | ||
| 173 | SECISBP2L | 15195 | -0.084 | -0.0857 | No | ||
| 174 | DPP4 | 15231 | -0.086 | -0.0842 | No | ||
| 175 | ESR1 | 15312 | -0.091 | -0.0844 | No | ||
| 176 | AMN1 | 15602 | -0.109 | -0.0927 | No | ||
| 177 | ZNF614 | 15654 | -0.111 | -0.0918 | No | ||
| 178 | ST3GAL6 | 15813 | -0.121 | -0.0950 | No | ||
| 179 | STOX1 | 15891 | -0.126 | -0.0951 | No | ||
| 180 | GLRX | 15957 | -0.131 | -0.0948 | No | ||
| 181 | ALDH1A2 | 16212 | -0.148 | -0.1017 | No | ||
| 182 | MKLN1 | 16363 | -0.158 | -0.1047 | No | ||
| 183 | DMXL1 | 16400 | -0.161 | -0.1032 | No | ||
| 184 | ACSM3 | 16513 | -0.171 | -0.1047 | No | ||
| 185 | FAM60A | 16563 | -0.175 | -0.1037 | No | ||
| 186 | ATM | 16808 | -0.200 | -0.1102 | No | ||
| 187 | GLCCI1 | 17060 | -0.223 | -0.1171 | No | ||
| 188 | MALAT1 | 17089 | -0.227 | -0.1153 | No | ||
| 189 | PIAS1 | 17153 | -0.234 | -0.1149 | No | ||
| 190 | CDK19 | 17352 | -0.257 | -0.1196 | No | ||
| 191 | KLHDC10 | 17454 | -0.270 | -0.1207 | No | ||
| 192 | ZMYM2 | 17519 | -0.277 | -0.1203 | No | ||
| 193 | FOLR1 | 17544 | -0.279 | -0.1184 | No | ||
| 194 | CPM | 17661 | -0.293 | -0.1200 | No | ||
| 195 | ZNF350 | 17884 | -0.320 | -0.1257 | No | ||
| 196 | PTER | 18282 | -0.377 | -0.1381 | No | ||
| 197 | HEY1 | 18340 | -0.387 | -0.1375 | No | ||
| 198 | PDE9A | 18683 | -0.450 | -0.1478 | No | ||
| 199 | LGALS8 | 18748 | -0.464 | -0.1474 | No | ||
| 200 | TSPAN6 | 18848 | -0.482 | -0.1484 | No | ||
| 201 | CYS1 | 18878 | -0.488 | -0.1466 | No | ||
| 202 | SH3PXD2A | 19005 | -0.513 | -0.1487 | No | ||
| 203 | RUFY3 | 19023 | -0.518 | -0.1465 | No | ||
| 204 | HLA-E | 19140 | -0.542 | -0.1481 | No | ||
| 205 | RICTOR | 19176 | -0.549 | -0.1466 | No | ||
| 206 | IL6ST | 19434 | -0.614 | -0.1536 | No | ||
| 207 | SEC62 | 19620 | -0.664 | -0.1579 | No | ||
| 208 | TSPAN1 | 19723 | -0.691 | -0.1590 | No | ||
| 209 | SPATA20 | 19815 | -0.724 | -0.1596 | No | ||
| 210 | ASPH | 19927 | -0.758 | -0.1611 | No | ||
| 211 | CASZ1 | 19937 | -0.762 | -0.1586 | No | ||
| 212 | FAM13A | 19958 | -0.769 | -0.1565 | No | ||
| 213 | DEPTOR | 19962 | -0.770 | -0.1537 | No | ||
| 214 | TMEM106B | 20009 | -0.786 | -0.1527 | No | ||
| 215 | SNRPN | 20021 | -0.789 | -0.1502 | No | ||
| 216 | GOLGA8A | 20056 | -0.796 | -0.1487 | No | ||
| 217 | C2orf88 | 20085 | -0.804 | -0.1469 | No | ||
| 218 | MALL | 20194 | -0.846 | -0.1482 | No | ||
| 219 | DMXL2 | 20254 | -0.870 | -0.1477 | No | ||
| 220 | ATP7A | 20318 | -0.890 | -0.1472 | No | ||
| 221 | MAFF | 20369 | -0.911 | -0.1463 | No | ||
| 222 | CERS6 | 20374 | -0.913 | -0.1436 | No | ||
| 223 | PAPD5 | 20387 | -0.916 | -0.1412 | No | ||
| 224 | TPD52 | 20450 | -0.940 | -0.1408 | No | ||
| 225 | TMED4 | 20510 | -0.963 | -0.1402 | No | ||
| 226 | ALG13 | 20759 | -1.071 | -0.1469 | No | ||
| 227 | ZNF770 | 20895 | -1.136 | -0.1492 | No | ||
| 228 | DHRS13 | 20910 | -1.144 | -0.1469 | No | ||
| 229 | ZNF615 | 21025 | -1.207 | -0.1485 | No | ||
| 230 | DPY19L4 | 21056 | -1.226 | -0.1468 | No | ||
| 231 | TPCN1 | 21306 | -1.366 | -0.1535 | No | ||
| 232 | SLC25A36 | 21326 | -1.375 | -0.1514 | No | ||
| 233 | DYNC2LI1 | 21469 | -1.467 | -0.1540 | No | ||
| 234 | PRKAB1 | 21750 | -1.651 | -0.1620 | No | ||
| 235 | PAPSS2 | 21768 | -1.660 | -0.1598 | No | ||
| 236 | LMAN1 | 21970 | -1.838 | -0.1647 | Yes | ||
| 237 | SLC27A1 | 21973 | -1.840 | -0.1619 | Yes | ||
| 238 | ZNF780B | 22056 | -1.909 | -0.1622 | Yes | ||
| 239 | PRKD1 | 22063 | -1.913 | -0.1596 | Yes | ||
| 240 | PDCD4 | 22066 | -1.916 | -0.1568 | Yes | ||
| 241 | CGNL1 | 22208 | -2.038 | -0.1594 | Yes | ||
| 242 | GOLGA8B | 22225 | -2.058 | -0.1572 | Yes | ||
| 243 | TRPC6 | 22315 | -2.146 | -0.1577 | Yes | ||
| 244 | FAM189A1 | 22362 | -2.188 | -0.1567 | Yes | ||
| 245 | SMPDL3A | 22438 | -2.278 | -0.1567 | Yes | ||
| 246 | SMCHD1 | 22491 | -2.320 | -0.1558 | Yes | ||
| 247 | TMOD1 | 22504 | -2.334 | -0.1535 | Yes | ||
| 248 | RDH10 | 22550 | -2.377 | -0.1523 | Yes | ||
| 249 | RSF1 | 22564 | -2.398 | -0.1500 | Yes | ||
| 250 | SULT1C2 | 22565 | -2.399 | -0.1471 | Yes | ||
| 251 | YIPF6 | 22646 | -2.504 | -0.1474 | Yes | ||
| 252 | TAF9B | 22674 | -2.544 | -0.1456 | Yes | ||
| 253 | HTRA1 | 22680 | -2.552 | -0.1429 | Yes | ||
| 254 | NT5E | 22714 | -2.579 | -0.1413 | Yes | ||
| 255 | SALL1 | 22773 | -2.660 | -0.1407 | Yes | ||
| 256 | P4HTM | 22785 | -2.670 | -0.1383 | Yes | ||
| 257 | PLCG2 | 22919 | -2.847 | -0.1406 | Yes | ||
| 258 | SLC22A18 | 22941 | -2.876 | -0.1385 | Yes | ||
| 259 | CXADR | 22966 | -2.896 | -0.1366 | Yes | ||
| 260 | BNC2 | 23030 | -2.988 | -0.1362 | Yes | ||
| 261 | DBP | 23195 | -3.235 | -0.1396 | Yes | ||
| 262 | RBM39 | 23196 | -3.237 | -0.1368 | Yes | ||
| 263 | LAMP2 | 23209 | -3.261 | -0.1344 | Yes | ||
| 264 | RSL1D1 | 23287 | -3.399 | -0.1345 | Yes | ||
| 265 | THBD | 23462 | -3.718 | -0.1384 | Yes | ||
| 266 | VAV3 | 23486 | -3.765 | -0.1364 | Yes | ||
| 267 | HES1 | 23488 | -3.766 | -0.1336 | Yes | ||
| 268 | CAT | 23497 | -3.775 | -0.1311 | Yes | ||
| 269 | GRB14 | 23520 | -3.829 | -0.1291 | Yes | ||
| 270 | HGD | 23539 | -3.866 | -0.1269 | Yes | ||
| 271 | HK2 | 23634 | -4.085 | -0.1277 | Yes | ||
| 272 | XRN1 | 23652 | -4.122 | -0.1255 | Yes | ||
| 273 | NDUFS1 | 23713 | -4.266 | -0.1249 | Yes | ||
| 274 | CUEDC1 | 23756 | -4.364 | -0.1237 | Yes | ||
| 275 | MAST4 | 23769 | -4.402 | -0.1213 | Yes | ||
| 276 | GDPD1 | 23829 | -4.533 | -0.1207 | Yes | ||
| 277 | TRAPPC6A | 23839 | -4.567 | -0.1182 | Yes | ||
| 278 | NBN | 23844 | -4.578 | -0.1156 | Yes | ||
| 279 | RXFP1 | 23865 | -4.630 | -0.1135 | Yes | ||
| 280 | SLC6A12 | 24020 | -5.097 | -0.1166 | Yes | ||
| 281 | MUC1 | 24081 | -5.263 | -0.1160 | Yes | ||
| 282 | MYEF2 | 24109 | -5.324 | -0.1142 | Yes | ||
| 283 | RNASET2 | 24112 | -5.334 | -0.1114 | Yes | ||
| 284 | MXI1 | 24115 | -5.336 | -0.1087 | Yes | ||
| 285 | PPP1R9A | 24132 | -5.386 | -0.1064 | Yes | ||
| 286 | BHLHE40 | 24138 | -5.407 | -0.1038 | Yes | ||
| 287 | MSI2 | 24160 | -5.460 | -0.1017 | Yes | ||
| 288 | TM7SF2 | 24200 | -5.591 | -0.1004 | Yes | ||
| 289 | PDK3 | 24206 | -5.606 | -0.0977 | Yes | ||
| 290 | PITPNC1 | 24287 | -5.945 | -0.0980 | Yes | ||
| 291 | MTDH | 24386 | -6.334 | -0.0989 | Yes | ||
| 292 | GALNT14 | 24416 | -6.458 | -0.0972 | Yes | ||
| 293 | ANTXR1 | 24491 | -6.714 | -0.0972 | Yes | ||
| 294 | IER2 | 24682 | -7.558 | -0.1016 | Yes | ||
| 295 | SLC9A1 | 24734 | -7.850 | -0.1007 | Yes | ||
| 296 | LMBRD2 | 24738 | -7.865 | -0.0980 | Yes | ||
| 297 | RBM47 | 24864 | -8.644 | -0.1000 | Yes | ||
| 298 | BBX | 24910 | -8.978 | -0.0989 | Yes | ||
| 299 | CAMK2N1 | 24943 | -9.206 | -0.0972 | Yes | ||
| 300 | SLC2A10 | 24994 | -9.695 | -0.0963 | Yes | ||
| 301 | IDO1 | 25004 | -9.739 | -0.0938 | Yes | ||
| 302 | TMC5 | 25067 | -10.284 | -0.0934 | Yes | ||
| 303 | N4BP2L2 | 25107 | -10.628 | -0.0920 | Yes | ||
| 304 | ASS1 | 25113 | -10.709 | -0.0893 | Yes | ||
| 305 | HINT3 | 25192 | -11.494 | -0.0895 | Yes | ||
| 306 | SLC1A1 | 25224 | -11.828 | -0.0878 | Yes | ||
| 307 | DUSP5 | 25225 | -11.830 | -0.0850 | Yes | ||
| 308 | MET | 25240 | -11.954 | -0.0827 | Yes | ||
| 309 | P4HA1 | 25269 | -12.284 | -0.0809 | Yes | ||
| 310 | TMEM37 | 25270 | -12.296 | -0.0781 | Yes | ||
| 311 | RAB3IP | 25282 | -12.453 | -0.0756 | Yes | ||
| 312 | BST2 | 25313 | -12.832 | -0.0739 | Yes | ||
| 313 | PGPEP1 | 25427 | -14.337 | -0.0755 | Yes | ||
| 314 | DYNC2H1 | 25469 | -14.894 | -0.0742 | Yes | ||
| 315 | H6PD | 25493 | -15.251 | -0.0722 | Yes | ||
| 316 | ZNF226 | 25508 | -15.546 | -0.0699 | Yes | ||
| 317 | CDH1 | 25535 | -16.009 | -0.0681 | Yes | ||
| 318 | IFI16 | 25559 | -16.487 | -0.0661 | Yes | ||
| 319 | EEF2K | 25597 | -17.681 | -0.0647 | Yes | ||
| 320 | PIK3R1 | 25599 | -17.722 | -0.0619 | Yes | ||
| 321 | DUSP6 | 25608 | -17.968 | -0.0593 | Yes | ||
| 322 | TTC3 | 25625 | -18.237 | -0.0571 | Yes | ||
| 323 | AK4 | 25634 | -18.360 | -0.0545 | Yes | ||
| 324 | CNTNAP3 | 25654 | -18.791 | -0.0524 | Yes | ||
| 325 | PRUNE2 | 25661 | -18.954 | -0.0498 | Yes | ||
| 326 | ARSD | 25711 | -20.262 | -0.0489 | Yes | ||
| 327 | SERPINB1 | 25756 | -21.646 | -0.0477 | Yes | ||
| 328 | SMOC2 | 25848 | -25.146 | -0.0484 | Yes | ||
| 329 | APOOL | 25866 | -25.666 | -0.0462 | Yes | ||
| 330 | CCDC50 | 25945 | -30.328 | -0.0463 | Yes | ||
| 331 | RABL3 | 25947 | -30.339 | -0.0435 | Yes | ||
| 332 | PIGP | 25949 | -30.433 | -0.0407 | Yes | ||
| 333 | FAM107B | 25950 | -30.439 | -0.0378 | Yes | ||
| 334 | CD55 | 25987 | -33.140 | -0.0364 | Yes | ||
| 335 | SYTL2 | 25998 | -33.868 | -0.0339 | Yes | ||
| 336 | YPEL5 | 26050 | -38.933 | -0.0330 | Yes | ||
| 337 | PARP8 | 26051 | -38.977 | -0.0302 | Yes | ||
| 338 | MRAP2 | 26072 | -40.772 | -0.0281 | Yes | ||
| 339 | ETV5 | 26107 | -44.619 | -0.0266 | Yes | ||
| 340 | SLC7A11 | 26126 | -47.089 | -0.0244 | Yes | ||
| 341 | KRCC1 | 26142 | -50.080 | -0.0221 | Yes | ||
| 342 | ADM | 26168 | -55.394 | -0.0202 | Yes | ||
| 343 | PHLDA1 | 26170 | -55.684 | -0.0174 | Yes | ||
| 344 | VLDLR | 26190 | -59.843 | -0.0153 | Yes | ||
| 345 | SLC30A1 | 26223 | -73.941 | -0.0137 | Yes | ||
| 346 | SMARCA2 | 26237 | -82.253 | -0.0114 | Yes | ||
| 347 | SNTB1 | 26245 | -87.394 | -0.0088 | Yes | ||
| 348 | VEGFA | 26251 | -89.987 | -0.0061 | Yes | ||
| 349 | CHGA | 26292 | -180.006 | -0.0048 | Yes | ||
| 350 | DDIT4 | 26311 | -299.207 | -0.0027 | Yes | ||
| 351 | TSC22D3 | 26317 | -299.207 | -0.0000 | Yes |