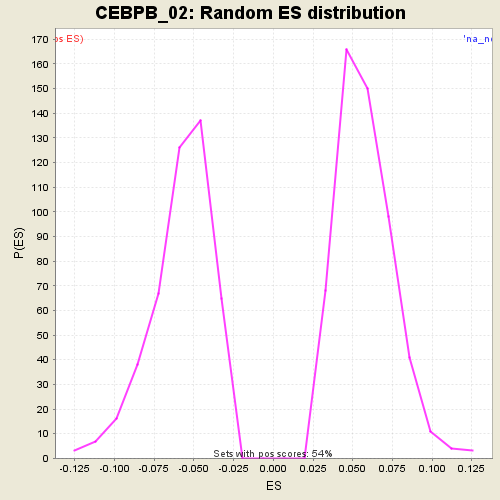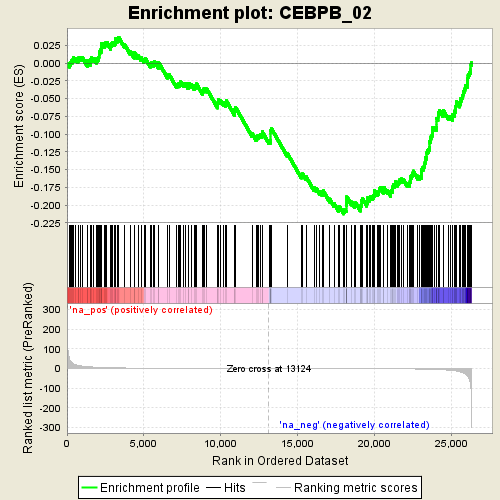
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | CEBPB_02 |
| Enrichment Score (ES) | -0.21265095 |
| Normalized Enrichment Score (NES) | -3.6552086 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PCDH7 | 146 | 52.996 | -0.0010 | No | ||
| 2 | EIF4A2 | 197 | 41.168 | 0.0016 | No | ||
| 3 | NFKBIZ | 270 | 32.545 | 0.0034 | No | ||
| 4 | PRR14L | 357 | 25.970 | 0.0046 | No | ||
| 5 | TOB1 | 385 | 24.709 | 0.0081 | No | ||
| 6 | EFNA5 | 553 | 19.297 | 0.0063 | No | ||
| 7 | VCAN | 712 | 16.078 | 0.0048 | No | ||
| 8 | HNRNPR | 762 | 15.243 | 0.0074 | No | ||
| 9 | MRC2 | 867 | 13.788 | 0.0080 | No | ||
| 10 | ONECUT2 | 974 | 12.436 | 0.0085 | No | ||
| 11 | PER1 | 1311 | 9.538 | 0.0001 | No | ||
| 12 | SPIB | 1336 | 9.361 | 0.0038 | No | ||
| 13 | SMAD6 | 1512 | 8.227 | 0.0016 | No | ||
| 14 | PITX2 | 1540 | 8.144 | 0.0051 | No | ||
| 15 | SLC12A1 | 1573 | 7.912 | 0.0084 | No | ||
| 16 | C1QL1 | 1730 | 7.140 | 0.0070 | No | ||
| 17 | TMEM104 | 1914 | 6.348 | 0.0045 | No | ||
| 18 | KPNA3 | 1991 | 6.089 | 0.0062 | No | ||
| 19 | H2AFZ | 2051 | 5.890 | 0.0085 | No | ||
| 20 | ACAN | 2078 | 5.784 | 0.0120 | No | ||
| 21 | MTF1 | 2092 | 5.745 | 0.0161 | No | ||
| 22 | DYRK3 | 2142 | 5.596 | 0.0187 | No | ||
| 23 | PDCL | 2228 | 5.317 | 0.0200 | No | ||
| 24 | ARF6 | 2236 | 5.288 | 0.0243 | No | ||
| 25 | LEMD2 | 2252 | 5.246 | 0.0283 | No | ||
| 26 | ACLY | 2430 | 4.768 | 0.0260 | No | ||
| 27 | MAP4K4 | 2482 | 4.661 | 0.0286 | No | ||
| 28 | BCL11A | 2595 | 4.435 | 0.0289 | No | ||
| 29 | MAP3K3 | 2855 | 3.857 | 0.0235 | No | ||
| 30 | SYNCRIP | 2873 | 3.833 | 0.0274 | No | ||
| 31 | PRDX3 | 2939 | 3.721 | 0.0294 | No | ||
| 32 | PDGFRL | 3071 | 3.498 | 0.0290 | No | ||
| 33 | KCNH7 | 3130 | 3.384 | 0.0313 | No | ||
| 34 | PHLDB1 | 3144 | 3.363 | 0.0353 | No | ||
| 35 | PAFAH2 | 3294 | 3.150 | 0.0342 | No | ||
| 36 | THRA | 3361 | 3.043 | 0.0362 | No | ||
| 37 | METTL9 | 3742 | 2.583 | 0.0262 | No | ||
| 38 | RASAL2 | 4125 | 2.201 | 0.0161 | No | ||
| 39 | SERPINF1 | 4362 | 1.962 | 0.0116 | No | ||
| 40 | ARID1B | 4378 | 1.947 | 0.0156 | No | ||
| 41 | EDN2 | 4622 | 1.762 | 0.0108 | No | ||
| 42 | H3F3B | 4816 | 1.613 | 0.0079 | No | ||
| 43 | PCTP | 5003 | 1.472 | 0.0054 | No | ||
| 44 | CPNE4 | 5089 | 1.413 | 0.0067 | No | ||
| 45 | LUZP1 | 5411 | 1.236 | -0.0011 | No | ||
| 46 | FBXW7 | 5460 | 1.204 | 0.0016 | No | ||
| 47 | RPA3 | 5617 | 1.112 | 0.0002 | No | ||
| 48 | TOP1 | 5684 | 1.076 | 0.0022 | No | ||
| 49 | ID3 | 5929 | 0.958 | -0.0026 | No | ||
| 50 | ITPR3 | 5950 | 0.948 | 0.0012 | No | ||
| 51 | FRMD5 | 6533 | 0.752 | -0.0166 | No | ||
| 52 | CSMD3 | 6644 | 0.712 | -0.0163 | No | ||
| 53 | GYS1 | 7128 | 0.569 | -0.0302 | No | ||
| 54 | RAB44 | 7218 | 0.546 | -0.0291 | No | ||
| 55 | NAT9 | 7336 | 0.520 | -0.0290 | No | ||
| 56 | ARNT | 7379 | 0.511 | -0.0261 | No | ||
| 57 | SOBP | 7549 | 0.475 | -0.0280 | No | ||
| 58 | SBF2 | 7689 | 0.449 | -0.0288 | No | ||
| 59 | SYNJ1 | 7879 | 0.416 | -0.0315 | No | ||
| 60 | C1orf122 | 7908 | 0.411 | -0.0280 | No | ||
| 61 | HOXC13 | 8083 | 0.383 | -0.0301 | No | ||
| 62 | IP6K2 | 8284 | 0.357 | -0.0333 | No | ||
| 63 | NDRG1 | 8342 | 0.347 | -0.0309 | No | ||
| 64 | EIF4A1 | 8417 | 0.335 | -0.0292 | No | ||
| 65 | FCER1G | 8825 | 0.281 | -0.0402 | No | ||
| 66 | HIST1H1C | 8844 | 0.279 | -0.0364 | No | ||
| 67 | DCAF6 | 8940 | 0.267 | -0.0355 | No | ||
| 68 | ADRB2 | 9079 | 0.251 | -0.0362 | No | ||
| 69 | SMARCAD1 | 9786 | 0.179 | -0.0587 | No | ||
| 70 | ATP13A4 | 9808 | 0.178 | -0.0550 | No | ||
| 71 | RUVBL2 | 9827 | 0.176 | -0.0511 | No | ||
| 72 | SMARCA1 | 9993 | 0.163 | -0.0529 | No | ||
| 73 | SLC19A3 | 10181 | 0.146 | -0.0555 | No | ||
| 74 | FMO2 | 10334 | 0.135 | -0.0568 | No | ||
| 75 | PPL | 10349 | 0.134 | -0.0528 | No | ||
| 76 | SIRPA | 10895 | 0.098 | -0.0691 | No | ||
| 77 | ERF | 10903 | 0.097 | -0.0649 | No | ||
| 78 | DLX1 | 10953 | 0.095 | -0.0622 | No | ||
| 79 | RPS21 | 12030 | 0.043 | -0.0989 | No | ||
| 80 | RAB2A | 12297 | 0.031 | -0.1045 | No | ||
| 81 | TRIB1 | 12362 | 0.028 | -0.1024 | No | ||
| 82 | PCF11 | 12469 | 0.023 | -0.1019 | No | ||
| 83 | SLC6A4 | 12551 | 0.020 | -0.1005 | No | ||
| 84 | REXO2 | 12678 | 0.015 | -0.1008 | No | ||
| 85 | RIN3 | 12685 | 0.015 | -0.0965 | No | ||
| 86 | PPARG | 13135 | -0.000 | -0.1091 | No | ||
| 87 | MYO1C | 13233 | -0.004 | -0.1083 | No | ||
| 88 | PPP1R3D | 13242 | -0.004 | -0.1041 | No | ||
| 89 | DDR2 | 13243 | -0.004 | -0.0995 | No | ||
| 90 | SLC25A12 | 13246 | -0.005 | -0.0950 | No | ||
| 91 | ARHGEF38 | 13296 | -0.006 | -0.0924 | No | ||
| 92 | MAP2K6 | 14326 | -0.045 | -0.1273 | No | ||
| 93 | RC3H2 | 15233 | -0.086 | -0.1574 | No | ||
| 94 | ESR1 | 15312 | -0.091 | -0.1559 | No | ||
| 95 | SHKBP1 | 15541 | -0.105 | -0.1601 | No | ||
| 96 | MPPED2 | 16066 | -0.138 | -0.1756 | No | ||
| 97 | ALDH1A2 | 16212 | -0.148 | -0.1766 | No | ||
| 98 | SMAD1 | 16440 | -0.165 | -0.1808 | No | ||
| 99 | SP8 | 16594 | -0.179 | -0.1821 | No | ||
| 100 | MPP6 | 16656 | -0.186 | -0.1799 | No | ||
| 101 | UBQLN1 | 17074 | -0.225 | -0.1913 | No | ||
| 102 | CHRM1 | 17367 | -0.259 | -0.1979 | No | ||
| 103 | WNT5A | 17667 | -0.294 | -0.2049 | No | ||
| 104 | PDE3B | 17718 | -0.300 | -0.2022 | No | ||
| 105 | PDAP1 | 17991 | -0.335 | -0.2081 | Yes | ||
| 106 | CYP24A1 | 18057 | -0.343 | -0.2061 | Yes | ||
| 107 | PLA2G4A | 18148 | -0.355 | -0.2050 | Yes | ||
| 108 | ZEB2 | 18149 | -0.355 | -0.2004 | Yes | ||
| 109 | SOX5 | 18173 | -0.358 | -0.1967 | Yes | ||
| 110 | MED12L | 18185 | -0.361 | -0.1926 | Yes | ||
| 111 | SARNP | 18196 | -0.363 | -0.1885 | Yes | ||
| 112 | RFX4 | 18500 | -0.416 | -0.1955 | Yes | ||
| 113 | CSNK1E | 18713 | -0.454 | -0.1991 | Yes | ||
| 114 | FGF14 | 18774 | -0.467 | -0.1969 | Yes | ||
| 115 | BMF | 19079 | -0.530 | -0.2040 | Yes | ||
| 116 | P2RY4 | 19107 | -0.535 | -0.2004 | Yes | ||
| 117 | ZBTB20 | 19150 | -0.545 | -0.1975 | Yes | ||
| 118 | NLRP3 | 19155 | -0.546 | -0.1931 | Yes | ||
| 119 | STEAP2 | 19202 | -0.556 | -0.1903 | Yes | ||
| 120 | BDKRB1 | 19496 | -0.629 | -0.1970 | Yes | ||
| 121 | LCOR | 19537 | -0.644 | -0.1940 | Yes | ||
| 122 | TGFB3 | 19546 | -0.647 | -0.1898 | Yes | ||
| 123 | TMPRSS4 | 19672 | -0.681 | -0.1900 | Yes | ||
| 124 | SBSN | 19741 | -0.697 | -0.1881 | Yes | ||
| 125 | STEAP4 | 19863 | -0.738 | -0.1882 | Yes | ||
| 126 | SEPHS2 | 19936 | -0.761 | -0.1864 | Yes | ||
| 127 | BDNF | 19969 | -0.773 | -0.1831 | Yes | ||
| 128 | UGP2 | 19979 | -0.776 | -0.1789 | Yes | ||
| 129 | STX18 | 20195 | -0.846 | -0.1825 | Yes | ||
| 130 | DENND4A | 20275 | -0.877 | -0.1810 | Yes | ||
| 131 | STAT3 | 20290 | -0.880 | -0.1770 | Yes | ||
| 132 | FOXP1 | 20351 | -0.904 | -0.1748 | Yes | ||
| 133 | BAZ1A | 20575 | -0.991 | -0.1788 | Yes | ||
| 134 | RS1 | 20594 | -0.995 | -0.1749 | Yes | ||
| 135 | RAD23B | 20837 | -1.110 | -0.1796 | Yes | ||
| 136 | FIGN | 21064 | -1.230 | -0.1838 | Yes | ||
| 137 | CTDSPL2 | 21065 | -1.230 | -0.1792 | Yes | ||
| 138 | S100PBP | 21151 | -1.276 | -0.1779 | Yes | ||
| 139 | CP | 21179 | -1.293 | -0.1744 | Yes | ||
| 140 | CDKL5 | 21195 | -1.306 | -0.1704 | Yes | ||
| 141 | ANGPT1 | 21314 | -1.369 | -0.1704 | Yes | ||
| 142 | CASK | 21338 | -1.381 | -0.1668 | Yes | ||
| 143 | NCKAP5 | 21513 | -1.490 | -0.1689 | Yes | ||
| 144 | NFKBIA | 21582 | -1.532 | -0.1669 | Yes | ||
| 145 | ASAH2 | 21639 | -1.570 | -0.1645 | Yes | ||
| 146 | CITED4 | 21719 | -1.633 | -0.1630 | Yes | ||
| 147 | DUSP1 | 21853 | -1.729 | -0.1636 | Yes | ||
| 148 | ASCL2 | 22113 | -1.956 | -0.1689 | Yes | ||
| 149 | FST | 22258 | -2.085 | -0.1699 | Yes | ||
| 150 | PPP1CB | 22299 | -2.127 | -0.1669 | Yes | ||
| 151 | ITGA11 | 22335 | -2.161 | -0.1637 | Yes | ||
| 152 | BNIP3L | 22352 | -2.182 | -0.1598 | Yes | ||
| 153 | IL1RAPL2 | 22420 | -2.257 | -0.1578 | Yes | ||
| 154 | UBE2E2 | 22461 | -2.298 | -0.1548 | Yes | ||
| 155 | PRLR | 22507 | -2.339 | -0.1520 | Yes | ||
| 156 | RBPMS | 22810 | -2.696 | -0.1590 | Yes | ||
| 157 | SERTAD4 | 22944 | -2.878 | -0.1595 | Yes | ||
| 158 | BNC2 | 23030 | -2.988 | -0.1582 | Yes | ||
| 159 | LONRF3 | 23046 | -3.007 | -0.1543 | Yes | ||
| 160 | CACNB2 | 23060 | -3.025 | -0.1502 | Yes | ||
| 161 | ELAVL2 | 23107 | -3.094 | -0.1474 | Yes | ||
| 162 | RBM39 | 23196 | -3.237 | -0.1463 | Yes | ||
| 163 | TXLNG | 23221 | -3.277 | -0.1426 | Yes | ||
| 164 | CDKN1B | 23273 | -3.364 | -0.1401 | Yes | ||
| 165 | TRPM1 | 23301 | -3.420 | -0.1365 | Yes | ||
| 166 | ZADH2 | 23311 | -3.444 | -0.1323 | Yes | ||
| 167 | PDE4D | 23393 | -3.607 | -0.1309 | Yes | ||
| 168 | MOSPD2 | 23396 | -3.611 | -0.1264 | Yes | ||
| 169 | PIP5K1A | 23424 | -3.658 | -0.1229 | Yes | ||
| 170 | PFN2 | 23510 | -3.792 | -0.1216 | Yes | ||
| 171 | KCNE3 | 23548 | -3.887 | -0.1185 | Yes | ||
| 172 | NEK6 | 23573 | -3.949 | -0.1149 | Yes | ||
| 173 | SLIT3 | 23575 | -3.953 | -0.1104 | Yes | ||
| 174 | MEOX2 | 23615 | -4.040 | -0.1073 | Yes | ||
| 175 | TTC39B | 23650 | -4.118 | -0.1041 | Yes | ||
| 176 | KCNJ2 | 23725 | -4.291 | -0.1024 | Yes | ||
| 177 | CITED1 | 23740 | -4.329 | -0.0984 | Yes | ||
| 178 | CUEDC1 | 23756 | -4.364 | -0.0944 | Yes | ||
| 179 | DBH | 23786 | -4.435 | -0.0910 | Yes | ||
| 180 | PDGFRA | 23905 | -4.753 | -0.0909 | Yes | ||
| 181 | NFIL3 | 24002 | -5.017 | -0.0901 | Yes | ||
| 182 | ASGR1 | 24028 | -5.122 | -0.0865 | Yes | ||
| 183 | GOT1 | 24030 | -5.126 | -0.0820 | Yes | ||
| 184 | NHLH1 | 24032 | -5.132 | -0.0775 | Yes | ||
| 185 | ZNF217 | 24124 | -5.354 | -0.0764 | Yes | ||
| 186 | PTPN12 | 24128 | -5.376 | -0.0720 | Yes | ||
| 187 | MBNL1 | 24169 | -5.494 | -0.0690 | Yes | ||
| 188 | PKNOX2 | 24225 | -5.671 | -0.0665 | Yes | ||
| 189 | ARRDC3 | 24452 | -6.549 | -0.0706 | Yes | ||
| 190 | RRBP1 | 24465 | -6.594 | -0.0665 | Yes | ||
| 191 | H2AFJ | 24816 | -8.300 | -0.0754 | Yes | ||
| 192 | PPM1B | 24915 | -8.997 | -0.0746 | Yes | ||
| 193 | LYPD1 | 25075 | -10.360 | -0.0762 | Yes | ||
| 194 | RHOB | 25084 | -10.442 | -0.0719 | Yes | ||
| 195 | DENND2D | 25180 | -11.367 | -0.0710 | Yes | ||
| 196 | DMD | 25205 | -11.618 | -0.0674 | Yes | ||
| 197 | PRR15 | 25260 | -12.124 | -0.0649 | Yes | ||
| 198 | RAB3IP | 25282 | -12.453 | -0.0612 | Yes | ||
| 199 | COL4A3 | 25296 | -12.671 | -0.0571 | Yes | ||
| 200 | BCL6 | 25317 | -12.864 | -0.0534 | Yes | ||
| 201 | ATAD2 | 25540 | -16.130 | -0.0573 | Yes | ||
| 202 | COL4A4 | 25585 | -17.346 | -0.0545 | Yes | ||
| 203 | SPRED1 | 25596 | -17.652 | -0.0503 | Yes | ||
| 204 | LMO4 | 25682 | -19.607 | -0.0490 | Yes | ||
| 205 | NFE2L2 | 25715 | -20.354 | -0.0457 | Yes | ||
| 206 | SPRY4 | 25748 | -21.510 | -0.0424 | Yes | ||
| 207 | TCF12 | 25794 | -22.666 | -0.0395 | Yes | ||
| 208 | CHD2 | 25830 | -24.016 | -0.0363 | Yes | ||
| 209 | SOCS2 | 25879 | -26.186 | -0.0336 | Yes | ||
| 210 | CBX4 | 25927 | -29.298 | -0.0309 | Yes | ||
| 211 | CEP120 | 26015 | -35.580 | -0.0297 | Yes | ||
| 212 | CCND2 | 26028 | -36.907 | -0.0256 | Yes | ||
| 213 | PDGFB | 26055 | -39.111 | -0.0220 | Yes | ||
| 214 | TGIF1 | 26070 | -40.726 | -0.0180 | Yes | ||
| 215 | SLC7A11 | 26126 | -47.089 | -0.0156 | Yes | ||
| 216 | TAC1 | 26186 | -59.363 | -0.0133 | Yes | ||
| 217 | SMARCA2 | 26237 | -82.253 | -0.0107 | Yes | ||
| 218 | SULF1 | 26261 | -97.588 | -0.0070 | Yes | ||
| 219 | TSHZ3 | 26264 | -98.836 | -0.0026 | Yes | ||
| 220 | G0S2 | 26301 | -213.686 | 0.0006 | Yes |