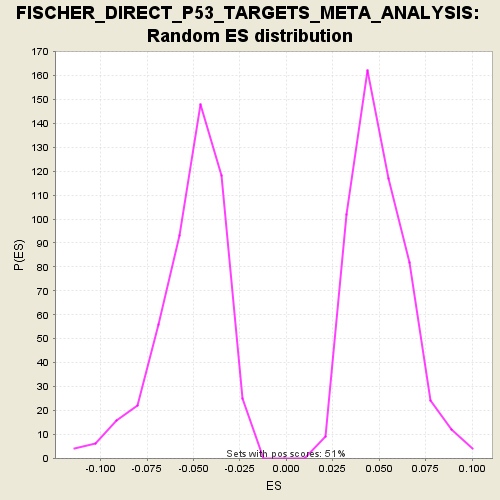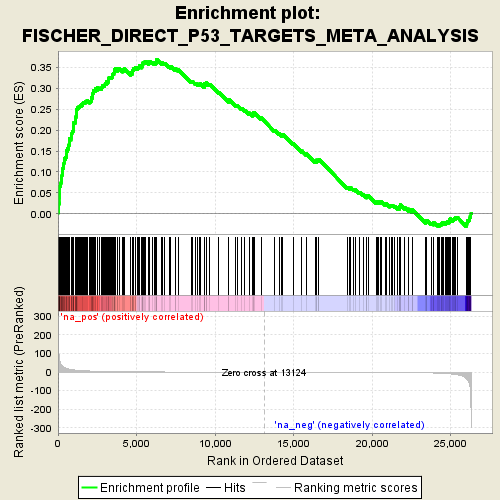
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | FISCHER_DIRECT_P53_TARGETS_META_ANALYSIS |
| Enrichment Score (ES) | 0.36791366 |
| Normalized Enrichment Score (NES) | 7.3137536 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CDKN1A | 2 | 299.207 | 0.0034 | Yes | ||
| 2 | TRIM22 | 9 | 242.455 | 0.0067 | Yes | ||
| 3 | PHLDA3 | 12 | 209.243 | 0.0101 | Yes | ||
| 4 | EDA2R | 25 | 129.226 | 0.0131 | Yes | ||
| 5 | SESN1 | 29 | 121.516 | 0.0165 | Yes | ||
| 6 | RPS27L | 40 | 106.999 | 0.0196 | Yes | ||
| 7 | BAX | 53 | 95.755 | 0.0226 | Yes | ||
| 8 | BTG2 | 60 | 90.877 | 0.0258 | Yes | ||
| 9 | POLH | 68 | 81.789 | 0.0291 | Yes | ||
| 10 | DDB2 | 70 | 81.275 | 0.0325 | Yes | ||
| 11 | VWCE | 71 | 80.787 | 0.0360 | Yes | ||
| 12 | TUBB6 | 76 | 77.923 | 0.0393 | Yes | ||
| 13 | SULF2 | 77 | 77.683 | 0.0428 | Yes | ||
| 14 | ZMAT3 | 82 | 75.551 | 0.0461 | Yes | ||
| 15 | TSPAN11 | 83 | 74.265 | 0.0496 | Yes | ||
| 16 | TNFRSF10B | 94 | 67.462 | 0.0527 | Yes | ||
| 17 | AEN | 98 | 66.791 | 0.0561 | Yes | ||
| 18 | GRHL3 | 99 | 66.649 | 0.0596 | Yes | ||
| 19 | FDXR | 106 | 65.175 | 0.0628 | Yes | ||
| 20 | TP53INP1 | 107 | 64.094 | 0.0663 | Yes | ||
| 21 | BBC3 | 149 | 52.488 | 0.0682 | Yes | ||
| 22 | MDM2 | 156 | 51.042 | 0.0715 | Yes | ||
| 23 | TMEM64 | 185 | 43.060 | 0.0739 | Yes | ||
| 24 | COBLL1 | 192 | 41.878 | 0.0771 | Yes | ||
| 25 | XPC | 204 | 40.284 | 0.0802 | Yes | ||
| 26 | SLC12A4 | 218 | 38.278 | 0.0832 | Yes | ||
| 27 | DRAM1 | 235 | 36.039 | 0.0860 | Yes | ||
| 28 | TRIAP1 | 239 | 35.685 | 0.0894 | Yes | ||
| 29 | RAP2B | 246 | 35.107 | 0.0927 | Yes | ||
| 30 | PCNA | 255 | 34.423 | 0.0958 | Yes | ||
| 31 | EI24 | 260 | 34.262 | 0.0992 | Yes | ||
| 32 | RRM2B | 281 | 31.368 | 0.1019 | Yes | ||
| 33 | GDF15 | 292 | 30.340 | 0.1050 | Yes | ||
| 34 | TNFRSF10D | 306 | 29.057 | 0.1080 | Yes | ||
| 35 | HEXIM1 | 324 | 27.700 | 0.1108 | Yes | ||
| 36 | SPATA18 | 325 | 27.585 | 0.1143 | Yes | ||
| 37 | TM7SF3 | 354 | 26.007 | 0.1167 | Yes | ||
| 38 | IKBIP | 371 | 25.423 | 0.1196 | Yes | ||
| 39 | LMNA | 381 | 24.893 | 0.1227 | Yes | ||
| 40 | TNFRSF10C | 390 | 24.454 | 0.1259 | Yes | ||
| 41 | ZC3HAV1 | 393 | 24.314 | 0.1293 | Yes | ||
| 42 | TNFRSF10A | 412 | 23.568 | 0.1321 | Yes | ||
| 43 | PLK2 | 461 | 21.783 | 0.1337 | Yes | ||
| 44 | GPC1 | 503 | 20.492 | 0.1356 | Yes | ||
| 45 | EPS8L2 | 524 | 19.973 | 0.1384 | Yes | ||
| 46 | CDH8 | 537 | 19.545 | 0.1414 | Yes | ||
| 47 | NTPCR | 550 | 19.341 | 0.1444 | Yes | ||
| 48 | PHPT1 | 551 | 19.325 | 0.1479 | Yes | ||
| 49 | CYFIP2 | 556 | 19.234 | 0.1512 | Yes | ||
| 50 | NYNRIN | 598 | 18.362 | 0.1531 | Yes | ||
| 51 | SFN | 603 | 18.262 | 0.1565 | Yes | ||
| 52 | RABGGTA | 654 | 17.240 | 0.1580 | Yes | ||
| 53 | DCP1B | 659 | 17.150 | 0.1614 | Yes | ||
| 54 | BRMS1L | 673 | 16.884 | 0.1643 | Yes | ||
| 55 | PLK3 | 703 | 16.241 | 0.1667 | Yes | ||
| 56 | CLP1 | 705 | 16.207 | 0.1702 | Yes | ||
| 57 | DDR1 | 711 | 16.106 | 0.1734 | Yes | ||
| 58 | VCAN | 712 | 16.078 | 0.1769 | Yes | ||
| 59 | ASCC3 | 736 | 15.646 | 0.1795 | Yes | ||
| 60 | RNF19B | 823 | 14.447 | 0.1797 | Yes | ||
| 61 | E2F7 | 830 | 14.296 | 0.1830 | Yes | ||
| 62 | CCNG1 | 836 | 14.184 | 0.1863 | Yes | ||
| 63 | RGMA | 877 | 13.682 | 0.1882 | Yes | ||
| 64 | CHST14 | 879 | 13.639 | 0.1916 | Yes | ||
| 65 | SAC3D1 | 890 | 13.487 | 0.1947 | Yes | ||
| 66 | DHRS3 | 933 | 12.896 | 0.1966 | Yes | ||
| 67 | UTP3 | 959 | 12.561 | 0.1991 | Yes | ||
| 68 | LIF | 967 | 12.489 | 0.2024 | Yes | ||
| 69 | ICOSLG | 977 | 12.412 | 0.2055 | Yes | ||
| 70 | DNAJB5 | 986 | 12.313 | 0.2087 | Yes | ||
| 71 | MARCH8 | 996 | 12.210 | 0.2118 | Yes | ||
| 72 | EPM2AIP1 | 1000 | 12.122 | 0.2152 | Yes | ||
| 73 | RAD51C | 1006 | 11.983 | 0.2185 | Yes | ||
| 74 | EML2 | 1080 | 11.261 | 0.2192 | Yes | ||
| 75 | RAB1A | 1098 | 11.114 | 0.2220 | Yes | ||
| 76 | GNAS | 1126 | 10.933 | 0.2244 | Yes | ||
| 77 | SLC4A11 | 1127 | 10.917 | 0.2279 | Yes | ||
| 78 | NINJ1 | 1134 | 10.869 | 0.2312 | Yes | ||
| 79 | TSKU | 1145 | 10.780 | 0.2343 | Yes | ||
| 80 | PANK1 | 1161 | 10.649 | 0.2372 | Yes | ||
| 81 | FAM84B | 1162 | 10.639 | 0.2407 | Yes | ||
| 82 | FBXO22 | 1174 | 10.526 | 0.2437 | Yes | ||
| 83 | MAP3K12 | 1175 | 10.523 | 0.2472 | Yes | ||
| 84 | RGL1 | 1193 | 10.435 | 0.2500 | Yes | ||
| 85 | ISYNA1 | 1242 | 9.978 | 0.2517 | Yes | ||
| 86 | TGS1 | 1254 | 9.898 | 0.2547 | Yes | ||
| 87 | PPM1D | 1329 | 9.415 | 0.2554 | Yes | ||
| 88 | FAS | 1372 | 9.028 | 0.2573 | Yes | ||
| 89 | ACER2 | 1407 | 8.821 | 0.2594 | Yes | ||
| 90 | NHLH2 | 1473 | 8.436 | 0.2604 | Yes | ||
| 91 | CCDC90B | 1548 | 8.076 | 0.2611 | Yes | ||
| 92 | PARD6G | 1580 | 7.862 | 0.2634 | Yes | ||
| 93 | ISCU | 1592 | 7.774 | 0.2664 | Yes | ||
| 94 | TRAF4 | 1685 | 7.356 | 0.2664 | Yes | ||
| 95 | MGRN1 | 1739 | 7.123 | 0.2678 | Yes | ||
| 96 | LRPAP1 | 1792 | 6.868 | 0.2693 | Yes | ||
| 97 | DGKA | 1870 | 6.514 | 0.2698 | Yes | ||
| 98 | TMEM68 | 2014 | 5.998 | 0.2678 | Yes | ||
| 99 | POLR2H | 2080 | 5.778 | 0.2688 | Yes | ||
| 100 | APAF1 | 2103 | 5.711 | 0.2714 | Yes | ||
| 101 | HRAS | 2126 | 5.642 | 0.2741 | Yes | ||
| 102 | DYRK3 | 2142 | 5.596 | 0.2770 | Yes | ||
| 103 | PTP4A1 | 2168 | 5.519 | 0.2795 | Yes | ||
| 104 | SERPINB5 | 2201 | 5.403 | 0.2818 | Yes | ||
| 105 | SCRIB | 2207 | 5.381 | 0.2851 | Yes | ||
| 106 | MICALL1 | 2215 | 5.357 | 0.2883 | Yes | ||
| 107 | ETV7 | 2232 | 5.312 | 0.2912 | Yes | ||
| 108 | FAM98C | 2235 | 5.294 | 0.2946 | Yes | ||
| 109 | ACTA2 | 2320 | 5.036 | 0.2948 | Yes | ||
| 110 | PLXNB2 | 2377 | 4.904 | 0.2961 | Yes | ||
| 111 | SERTAD1 | 2388 | 4.874 | 0.2992 | Yes | ||
| 112 | PRKD2 | 2494 | 4.643 | 0.2987 | Yes | ||
| 113 | ASTN2 | 2496 | 4.643 | 0.3021 | Yes | ||
| 114 | FCHO2 | 2665 | 4.281 | 0.2992 | Yes | ||
| 115 | TMEM63B | 2753 | 4.074 | 0.2993 | Yes | ||
| 116 | SOCS4 | 2790 | 3.997 | 0.3014 | Yes | ||
| 117 | MLF2 | 2809 | 3.960 | 0.3042 | Yes | ||
| 118 | TP53I3 | 2846 | 3.881 | 0.3063 | Yes | ||
| 119 | F5 | 2917 | 3.765 | 0.3071 | Yes | ||
| 120 | LNPEP | 2986 | 3.643 | 0.3080 | Yes | ||
| 121 | COL7A1 | 2997 | 3.632 | 0.3111 | Yes | ||
| 122 | RETSAT | 3069 | 3.503 | 0.3118 | Yes | ||
| 123 | SRA1 | 3076 | 3.483 | 0.3151 | Yes | ||
| 124 | ZNF385A | 3164 | 3.340 | 0.3152 | Yes | ||
| 125 | RRAD | 3188 | 3.309 | 0.3178 | Yes | ||
| 126 | IER5 | 3192 | 3.300 | 0.3212 | Yes | ||
| 127 | SFXN5 | 3208 | 3.261 | 0.3241 | Yes | ||
| 128 | TRIM35 | 3260 | 3.200 | 0.3256 | Yes | ||
| 129 | TRIM32 | 3358 | 3.048 | 0.3254 | Yes | ||
| 130 | TNFAIP8 | 3430 | 2.949 | 0.3262 | Yes | ||
| 131 | NADSYN1 | 3448 | 2.934 | 0.3290 | Yes | ||
| 132 | HSPA4L | 3458 | 2.917 | 0.3321 | Yes | ||
| 133 | EPHA2 | 3497 | 2.873 | 0.3341 | Yes | ||
| 134 | WDR81 | 3562 | 2.787 | 0.3352 | Yes | ||
| 135 | PRDM1 | 3563 | 2.787 | 0.3387 | Yes | ||
| 136 | RCBTB1 | 3585 | 2.761 | 0.3413 | Yes | ||
| 137 | WDR63 | 3590 | 2.757 | 0.3447 | Yes | ||
| 138 | CROT | 3637 | 2.718 | 0.3464 | Yes | ||
| 139 | IGFBP7 | 3794 | 2.533 | 0.3439 | Yes | ||
| 140 | ARFGEF2 | 3797 | 2.532 | 0.3473 | Yes | ||
| 141 | GABARAPL2 | 3882 | 2.434 | 0.3475 | Yes | ||
| 142 | ZNF219 | 4124 | 2.202 | 0.3418 | Yes | ||
| 143 | AMZ2 | 4169 | 2.132 | 0.3436 | Yes | ||
| 144 | MON2 | 4200 | 2.098 | 0.3459 | Yes | ||
| 145 | POU3F1 | 4593 | 1.779 | 0.3343 | Yes | ||
| 146 | EDN2 | 4622 | 1.762 | 0.3367 | Yes | ||
| 147 | GSS | 4711 | 1.683 | 0.3368 | Yes | ||
| 148 | MRPL49 | 4721 | 1.676 | 0.3400 | Yes | ||
| 149 | SYTL1 | 4760 | 1.649 | 0.3420 | Yes | ||
| 150 | SMAD5 | 4783 | 1.633 | 0.3446 | Yes | ||
| 151 | EFCAB10 | 4820 | 1.611 | 0.3467 | Yes | ||
| 152 | SNX2 | 4899 | 1.546 | 0.3472 | Yes | ||
| 153 | TANC1 | 4917 | 1.535 | 0.3501 | Yes | ||
| 154 | USP15 | 5077 | 1.420 | 0.3474 | Yes | ||
| 155 | POU2F2 | 5125 | 1.394 | 0.3491 | Yes | ||
| 156 | CMBL | 5156 | 1.377 | 0.3514 | Yes | ||
| 157 | LCE1E | 5194 | 1.357 | 0.3535 | Yes | ||
| 158 | RPS19 | 5342 | 1.269 | 0.3513 | Yes | ||
| 159 | NKAIN4 | 5370 | 1.250 | 0.3538 | Yes | ||
| 160 | PML | 5388 | 1.243 | 0.3566 | Yes | ||
| 161 | GPX1 | 5395 | 1.241 | 0.3599 | Yes | ||
| 162 | FBXW7 | 5460 | 1.204 | 0.3609 | Yes | ||
| 163 | CD82 | 5515 | 1.169 | 0.3623 | Yes | ||
| 164 | FLRT2 | 5580 | 1.131 | 0.3633 | Yes | ||
| 165 | TLCD1 | 5767 | 1.036 | 0.3597 | Yes | ||
| 166 | PARD6B | 5812 | 1.009 | 0.3615 | Yes | ||
| 167 | SMAD3 | 5839 | 0.995 | 0.3639 | Yes | ||
| 168 | PRRG2 | 5984 | 0.937 | 0.3619 | Yes | ||
| 169 | GBE1 | 6131 | 0.886 | 0.3598 | Yes | ||
| 170 | AK3 | 6201 | 0.858 | 0.3606 | Yes | ||
| 171 | SCN2A | 6245 | 0.840 | 0.3624 | Yes | ||
| 172 | TLR3 | 6260 | 0.836 | 0.3654 | Yes | ||
| 173 | FAM198B | 6286 | 0.827 | 0.3679 | Yes | ||
| 174 | CEL | 6587 | 0.731 | 0.3599 | No | ||
| 175 | KRT15 | 6668 | 0.699 | 0.3603 | No | ||
| 176 | RFK | 6774 | 0.666 | 0.3597 | No | ||
| 177 | EID2B | 7082 | 0.580 | 0.3514 | No | ||
| 178 | PBLD | 7187 | 0.552 | 0.3509 | No | ||
| 179 | CNOT6 | 7450 | 0.498 | 0.3443 | No | ||
| 180 | FCHSD2 | 7457 | 0.496 | 0.3476 | No | ||
| 181 | ZNF561 | 7650 | 0.454 | 0.3437 | No | ||
| 182 | AKAP9 | 8470 | 0.326 | 0.3157 | No | ||
| 183 | S100A2 | 8541 | 0.317 | 0.3165 | No | ||
| 184 | NDUFV3 | 8750 | 0.292 | 0.3120 | No | ||
| 185 | BLCAP | 8902 | 0.270 | 0.3097 | No | ||
| 186 | TRIP6 | 8988 | 0.263 | 0.3099 | No | ||
| 187 | SLC25A45 | 9039 | 0.256 | 0.3115 | No | ||
| 188 | ZNF79 | 9309 | 0.225 | 0.3046 | No | ||
| 189 | SYNC | 9314 | 0.225 | 0.3080 | No | ||
| 190 | BTG1 | 9324 | 0.224 | 0.3111 | No | ||
| 191 | AGBL5 | 9450 | 0.212 | 0.3098 | No | ||
| 192 | PRKAB2 | 9465 | 0.210 | 0.3127 | No | ||
| 193 | YAP1 | 9651 | 0.192 | 0.3091 | No | ||
| 194 | C17orf82 | 10238 | 0.141 | 0.2901 | No | ||
| 195 | CSNK1G1 | 10856 | 0.099 | 0.2699 | No | ||
| 196 | SEMA3B | 10871 | 0.099 | 0.2728 | No | ||
| 197 | ITPKC | 11305 | 0.076 | 0.2597 | No | ||
| 198 | ABCA12 | 11410 | 0.069 | 0.2591 | No | ||
| 199 | TMEM131 | 11691 | 0.056 | 0.2519 | No | ||
| 200 | GADD45A | 11891 | 0.047 | 0.2477 | No | ||
| 201 | ALDH1A3 | 12164 | 0.036 | 0.2407 | No | ||
| 202 | TRIB1 | 12362 | 0.028 | 0.2367 | No | ||
| 203 | BCL9L | 12424 | 0.026 | 0.2378 | No | ||
| 204 | PADI4 | 12429 | 0.025 | 0.2411 | No | ||
| 205 | ZNF337 | 12497 | 0.022 | 0.2420 | No | ||
| 206 | BLOC1S2 | 12926 | 0.006 | 0.2291 | No | ||
| 207 | C12orf45 | 13793 | -0.024 | 0.1993 | No | ||
| 208 | DUSP14 | 14069 | -0.035 | 0.1922 | No | ||
| 209 | RALGAPB | 14259 | -0.043 | 0.1884 | No | ||
| 210 | HHAT | 14320 | -0.045 | 0.1896 | No | ||
| 211 | ANXA4 | 14971 | -0.071 | 0.1681 | No | ||
| 212 | EFNB1 | 15532 | -0.104 | 0.1501 | No | ||
| 213 | KCTD1 | 15800 | -0.120 | 0.1433 | No | ||
| 214 | PDE4C | 16362 | -0.158 | 0.1253 | No | ||
| 215 | ZNF564 | 16449 | -0.166 | 0.1254 | No | ||
| 216 | ANKRA2 | 16458 | -0.167 | 0.1286 | No | ||
| 217 | MBD5 | 16562 | -0.175 | 0.1282 | No | ||
| 218 | ARAP1 | 16612 | -0.181 | 0.1298 | No | ||
| 219 | KLHL12 | 18419 | -0.403 | 0.0639 | No | ||
| 220 | ZNF37A | 18561 | -0.426 | 0.0619 | No | ||
| 221 | KIAA1217 | 18638 | -0.441 | 0.0625 | No | ||
| 222 | BCL10 | 18811 | -0.474 | 0.0594 | No | ||
| 223 | LRP1 | 18940 | -0.503 | 0.0579 | No | ||
| 224 | ZNF423 | 19207 | -0.557 | 0.0512 | No | ||
| 225 | ABCB6 | 19428 | -0.613 | 0.0462 | No | ||
| 226 | PSTPIP2 | 19645 | -0.674 | 0.0414 | No | ||
| 227 | OBSL1 | 19670 | -0.680 | 0.0440 | No | ||
| 228 | MCC | 19736 | -0.694 | 0.0450 | No | ||
| 229 | STAT3 | 20290 | -0.880 | 0.0272 | No | ||
| 230 | TEAD3 | 20321 | -0.891 | 0.0295 | No | ||
| 231 | CSF1 | 20430 | -0.932 | 0.0289 | No | ||
| 232 | STX6 | 20551 | -0.981 | 0.0278 | No | ||
| 233 | MFAP3L | 20566 | -0.987 | 0.0307 | No | ||
| 234 | ARHGAP42 | 20840 | -1.114 | 0.0237 | No | ||
| 235 | PPP1R3C | 20907 | -1.143 | 0.0246 | No | ||
| 236 | NCEH1 | 21123 | -1.260 | 0.0199 | No | ||
| 237 | HIST2H2BE | 21206 | -1.312 | 0.0202 | No | ||
| 238 | PTEN | 21288 | -1.359 | 0.0206 | No | ||
| 239 | PPFIBP1 | 21445 | -1.449 | 0.0181 | No | ||
| 240 | ORAI3 | 21635 | -1.567 | 0.0143 | No | ||
| 241 | APOBEC3C | 21718 | -1.633 | 0.0146 | No | ||
| 242 | PRKAB1 | 21750 | -1.651 | 0.0169 | No | ||
| 243 | REV3L | 21780 | -1.669 | 0.0193 | No | ||
| 244 | BCL2L1 | 21802 | -1.687 | 0.0220 | No | ||
| 245 | ITGA3 | 22089 | -1.937 | 0.0145 | No | ||
| 246 | TET2 | 22334 | -2.161 | 0.0086 | No | ||
| 247 | SUPT7L | 22342 | -2.171 | 0.0118 | No | ||
| 248 | LPXN | 22548 | -2.375 | 0.0074 | No | ||
| 249 | KCNN4 | 22561 | -2.396 | 0.0104 | No | ||
| 250 | GNAI1 | 23408 | -3.628 | -0.0186 | No | ||
| 251 | BTBD10 | 23438 | -3.679 | -0.0162 | No | ||
| 252 | HES1 | 23488 | -3.766 | -0.0146 | No | ||
| 253 | MAST4 | 23769 | -4.402 | -0.0219 | No | ||
| 254 | TRUB1 | 23889 | -4.708 | -0.0230 | No | ||
| 255 | GREB1 | 23915 | -4.774 | -0.0205 | No | ||
| 256 | BHLHE40 | 24138 | -5.407 | -0.0255 | No | ||
| 257 | CFLAR | 24227 | -5.676 | -0.0254 | No | ||
| 258 | CDC42SE1 | 24321 | -6.075 | -0.0255 | No | ||
| 259 | SERPINE1 | 24389 | -6.348 | -0.0246 | No | ||
| 260 | CPEB2 | 24429 | -6.500 | -0.0226 | No | ||
| 261 | SLC46A1 | 24482 | -6.662 | -0.0211 | No | ||
| 262 | BBS2 | 24529 | -6.878 | -0.0194 | No | ||
| 263 | CPSF4 | 24665 | -7.513 | -0.0211 | No | ||
| 264 | SLC9A1 | 24734 | -7.850 | -0.0202 | No | ||
| 265 | GCH1 | 24774 | -8.103 | -0.0182 | No | ||
| 266 | MFGE8 | 24873 | -8.686 | -0.0185 | No | ||
| 267 | IER3 | 24909 | -8.968 | -0.0164 | No | ||
| 268 | NUPR1 | 24923 | -9.037 | -0.0134 | No | ||
| 269 | RNF144B | 24962 | -9.378 | -0.0114 | No | ||
| 270 | KRT8 | 25119 | -10.758 | -0.0139 | No | ||
| 271 | KLHDC7A | 25193 | -11.514 | -0.0132 | No | ||
| 272 | MRPL39 | 25226 | -11.847 | -0.0109 | No | ||
| 273 | TP53 | 25300 | -12.686 | -0.0102 | No | ||
| 274 | BCL6 | 25317 | -12.864 | -0.0074 | No | ||
| 275 | RIN1 | 25419 | -14.202 | -0.0078 | No | ||
| 276 | IRF2BPL | 25985 | -33.000 | -0.0260 | No | ||
| 277 | FBN2 | 26004 | -34.322 | -0.0232 | No | ||
| 278 | IGDCC4 | 26042 | -38.110 | -0.0211 | No | ||
| 279 | PMAIP1 | 26053 | -38.997 | -0.0180 | No | ||
| 280 | LIMA1 | 26094 | -42.892 | -0.0161 | No | ||
| 281 | NEFL | 26149 | -51.235 | -0.0147 | No | ||
| 282 | SARS | 26192 | -60.122 | -0.0128 | No | ||
| 283 | SLC30A1 | 26223 | -73.941 | -0.0105 | No | ||
| 284 | RGS16 | 26231 | -76.235 | -0.0073 | No | ||
| 285 | SESN2 | 26273 | -109.055 | -0.0054 | No | ||
| 286 | TXNIP | 26293 | -180.286 | -0.0026 | No | ||
| 287 | ATF3 | 26295 | -192.665 | 0.0008 | No |