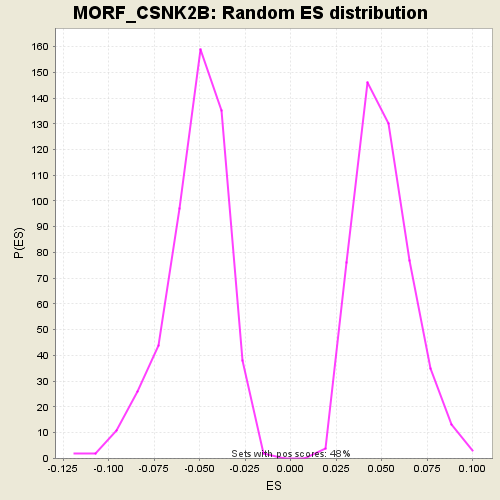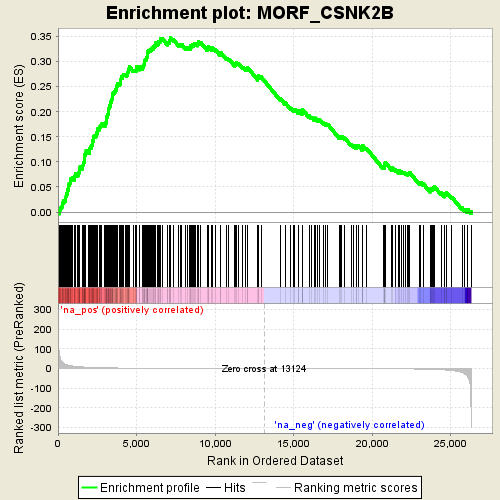
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | MORF_CSNK2B |
| Enrichment Score (ES) | 0.34815443 |
| Normalized Enrichment Score (NES) | 6.8069606 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BZW1 | 102 | 66.257 | -0.0004 | Yes | ||
| 2 | TUBB2A | 105 | 65.807 | 0.0031 | Yes | ||
| 3 | ARHGDIA | 126 | 58.390 | 0.0059 | Yes | ||
| 4 | KPNA2 | 160 | 49.616 | 0.0082 | Yes | ||
| 5 | EIF4E2 | 201 | 40.595 | 0.0102 | Yes | ||
| 6 | PCNA | 255 | 34.423 | 0.0118 | Yes | ||
| 7 | EI24 | 260 | 34.262 | 0.0152 | Yes | ||
| 8 | LSS | 273 | 32.374 | 0.0183 | Yes | ||
| 9 | UBAP2L | 327 | 27.414 | 0.0198 | Yes | ||
| 10 | DHCR7 | 331 | 27.123 | 0.0232 | Yes | ||
| 11 | SEC24C | 429 | 23.105 | 0.0231 | Yes | ||
| 12 | FEN1 | 473 | 21.511 | 0.0250 | Yes | ||
| 13 | LSM3 | 500 | 20.523 | 0.0275 | Yes | ||
| 14 | HCCS | 502 | 20.507 | 0.0310 | Yes | ||
| 15 | MAEA | 511 | 20.287 | 0.0343 | Yes | ||
| 16 | TMED9 | 539 | 19.512 | 0.0368 | Yes | ||
| 17 | DDX39A | 596 | 18.368 | 0.0382 | Yes | ||
| 18 | ATP5G1 | 609 | 18.106 | 0.0413 | Yes | ||
| 19 | EIF4G1 | 621 | 17.869 | 0.0445 | Yes | ||
| 20 | GLO1 | 669 | 16.943 | 0.0462 | Yes | ||
| 21 | TARDBP | 678 | 16.822 | 0.0495 | Yes | ||
| 22 | SF3A1 | 683 | 16.727 | 0.0529 | Yes | ||
| 23 | CALM3 | 693 | 16.496 | 0.0561 | Yes | ||
| 24 | RBBP4 | 755 | 15.378 | 0.0573 | Yes | ||
| 25 | PRPS2 | 770 | 15.143 | 0.0603 | Yes | ||
| 26 | G3BP2 | 805 | 14.830 | 0.0626 | Yes | ||
| 27 | RBM14 | 808 | 14.771 | 0.0661 | Yes | ||
| 28 | NDUFB3 | 878 | 13.682 | 0.0670 | Yes | ||
| 29 | SEC13 | 928 | 12.984 | 0.0686 | Yes | ||
| 30 | RNPEP | 1057 | 11.527 | 0.0673 | Yes | ||
| 31 | ATP5G3 | 1063 | 11.489 | 0.0706 | Yes | ||
| 32 | SNRPG | 1076 | 11.321 | 0.0737 | Yes | ||
| 33 | PPP2R1A | 1078 | 11.290 | 0.0773 | Yes | ||
| 34 | SUMO2 | 1250 | 9.927 | 0.0743 | Yes | ||
| 35 | TRAPPC3 | 1309 | 9.579 | 0.0756 | Yes | ||
| 36 | DYNLL1 | 1319 | 9.478 | 0.0788 | Yes | ||
| 37 | HSP90AA1 | 1345 | 9.282 | 0.0814 | Yes | ||
| 38 | UBE2L3 | 1351 | 9.211 | 0.0848 | Yes | ||
| 39 | FUS | 1383 | 8.968 | 0.0871 | Yes | ||
| 40 | LMNB2 | 1395 | 8.889 | 0.0903 | Yes | ||
| 41 | CBX3 | 1527 | 8.187 | 0.0888 | Yes | ||
| 42 | EIF4B | 1567 | 7.940 | 0.0909 | Yes | ||
| 43 | PUF60 | 1584 | 7.822 | 0.0938 | Yes | ||
| 44 | NSDHL | 1589 | 7.797 | 0.0972 | Yes | ||
| 45 | PRMT1 | 1642 | 7.527 | 0.0988 | Yes | ||
| 46 | TBL3 | 1667 | 7.437 | 0.1014 | Yes | ||
| 47 | COX6B1 | 1668 | 7.436 | 0.1050 | Yes | ||
| 48 | SYPL1 | 1682 | 7.364 | 0.1080 | Yes | ||
| 49 | SSBP1 | 1691 | 7.316 | 0.1113 | Yes | ||
| 50 | DOCK3 | 1694 | 7.294 | 0.1148 | Yes | ||
| 51 | SNRPD2 | 1742 | 7.092 | 0.1165 | Yes | ||
| 52 | HDAC1 | 1775 | 6.955 | 0.1188 | Yes | ||
| 53 | HNRNPA2B1 | 1777 | 6.941 | 0.1224 | Yes | ||
| 54 | SNRPE | 1908 | 6.371 | 0.1209 | Yes | ||
| 55 | HNRNPAB | 2011 | 6.002 | 0.1206 | Yes | ||
| 56 | ARF3 | 2015 | 5.993 | 0.1240 | Yes | ||
| 57 | RBMX | 2025 | 5.955 | 0.1272 | Yes | ||
| 58 | H2AFZ | 2051 | 5.890 | 0.1298 | Yes | ||
| 59 | COMMD4 | 2120 | 5.661 | 0.1308 | Yes | ||
| 60 | PTOV1 | 2140 | 5.605 | 0.1336 | Yes | ||
| 61 | HNRNPUL1 | 2169 | 5.509 | 0.1361 | Yes | ||
| 62 | PRPF31 | 2187 | 5.438 | 0.1390 | Yes | ||
| 63 | CYCS | 2189 | 5.436 | 0.1425 | Yes | ||
| 64 | ALG8 | 2241 | 5.280 | 0.1441 | Yes | ||
| 65 | YWHAE | 2245 | 5.274 | 0.1476 | Yes | ||
| 66 | VDAC1 | 2272 | 5.196 | 0.1501 | Yes | ||
| 67 | HNRNPA3 | 2318 | 5.038 | 0.1519 | Yes | ||
| 68 | PTGES3 | 2391 | 4.867 | 0.1527 | Yes | ||
| 69 | ACLY | 2430 | 4.768 | 0.1548 | Yes | ||
| 70 | FAM120A | 2445 | 4.736 | 0.1579 | Yes | ||
| 71 | BANF1 | 2505 | 4.630 | 0.1591 | Yes | ||
| 72 | MRPL9 | 2515 | 4.602 | 0.1624 | Yes | ||
| 73 | CS | 2517 | 4.592 | 0.1659 | Yes | ||
| 74 | KHDRBS1 | 2608 | 4.398 | 0.1660 | Yes | ||
| 75 | SNRNP200 | 2630 | 4.347 | 0.1687 | Yes | ||
| 76 | PSMB2 | 2666 | 4.280 | 0.1710 | Yes | ||
| 77 | RARS | 2726 | 4.131 | 0.1722 | Yes | ||
| 78 | UQCRH | 2750 | 4.082 | 0.1749 | Yes | ||
| 79 | SF3A2 | 2795 | 3.979 | 0.1768 | Yes | ||
| 80 | PRDX3 | 2939 | 3.721 | 0.1749 | Yes | ||
| 81 | SRSF1 | 3027 | 3.569 | 0.1751 | Yes | ||
| 82 | FIBP | 3044 | 3.537 | 0.1780 | Yes | ||
| 83 | LAGE3 | 3061 | 3.515 | 0.1810 | Yes | ||
| 84 | UBA1 | 3077 | 3.481 | 0.1839 | Yes | ||
| 85 | AFG3L2 | 3083 | 3.476 | 0.1873 | Yes | ||
| 86 | MCM7 | 3087 | 3.466 | 0.1908 | Yes | ||
| 87 | GPI | 3139 | 3.369 | 0.1924 | Yes | ||
| 88 | UQCRC1 | 3167 | 3.337 | 0.1949 | Yes | ||
| 89 | AP2M1 | 3189 | 3.303 | 0.1976 | Yes | ||
| 90 | CCT3 | 3201 | 3.286 | 0.2008 | Yes | ||
| 91 | SLC25A3 | 3213 | 3.255 | 0.2039 | Yes | ||
| 92 | SCAMP3 | 3230 | 3.233 | 0.2068 | Yes | ||
| 93 | AP2S1 | 3247 | 3.210 | 0.2098 | Yes | ||
| 94 | MFAP1 | 3269 | 3.183 | 0.2125 | Yes | ||
| 95 | SDHB | 3349 | 3.069 | 0.2131 | Yes | ||
| 96 | ATP5J2 | 3356 | 3.050 | 0.2164 | Yes | ||
| 97 | BAG6 | 3365 | 3.038 | 0.2196 | Yes | ||
| 98 | SKP1 | 3377 | 3.018 | 0.2228 | Yes | ||
| 99 | PARK7 | 3404 | 2.984 | 0.2253 | Yes | ||
| 100 | CAP1 | 3442 | 2.937 | 0.2275 | Yes | ||
| 101 | ATP5J | 3466 | 2.901 | 0.2302 | Yes | ||
| 102 | RAN | 3473 | 2.894 | 0.2335 | Yes | ||
| 103 | HAT1 | 3476 | 2.890 | 0.2370 | Yes | ||
| 104 | TRIM28 | 3532 | 2.820 | 0.2384 | Yes | ||
| 105 | TMEM147 | 3582 | 2.764 | 0.2401 | Yes | ||
| 106 | PSMD9 | 3627 | 2.733 | 0.2420 | Yes | ||
| 107 | MBTPS1 | 3639 | 2.718 | 0.2451 | Yes | ||
| 108 | PSMB4 | 3707 | 2.631 | 0.2461 | Yes | ||
| 109 | ATIC | 3716 | 2.617 | 0.2493 | Yes | ||
| 110 | AP3S1 | 3749 | 2.576 | 0.2517 | Yes | ||
| 111 | SUMO1 | 3763 | 2.565 | 0.2547 | Yes | ||
| 112 | VPS26A | 3800 | 2.530 | 0.2569 | Yes | ||
| 113 | ATP5D | 3906 | 2.402 | 0.2564 | Yes | ||
| 114 | PSMB7 | 3958 | 2.358 | 0.2580 | Yes | ||
| 115 | FBL | 3973 | 2.344 | 0.2610 | Yes | ||
| 116 | ERH | 3990 | 2.325 | 0.2640 | Yes | ||
| 117 | SLC4A2 | 4007 | 2.309 | 0.2669 | Yes | ||
| 118 | XPO7 | 4026 | 2.299 | 0.2698 | Yes | ||
| 119 | ILF3 | 4120 | 2.204 | 0.2698 | Yes | ||
| 120 | CHERP | 4132 | 2.190 | 0.2729 | Yes | ||
| 121 | NDUFAB1 | 4167 | 2.135 | 0.2752 | Yes | ||
| 122 | IDH3G | 4268 | 2.043 | 0.2749 | Yes | ||
| 123 | TECR | 4373 | 1.956 | 0.2745 | Yes | ||
| 124 | HMGN1 | 4411 | 1.921 | 0.2766 | Yes | ||
| 125 | SRSF9 | 4444 | 1.892 | 0.2789 | Yes | ||
| 126 | AKR7A2 | 4487 | 1.858 | 0.2809 | Yes | ||
| 127 | PDHB | 4493 | 1.855 | 0.2842 | Yes | ||
| 128 | YWHAB | 4514 | 1.842 | 0.2870 | Yes | ||
| 129 | CDK4 | 4534 | 1.825 | 0.2899 | Yes | ||
| 130 | PPP1CA | 4801 | 1.621 | 0.2832 | Yes | ||
| 131 | NHP2 | 4903 | 1.544 | 0.2829 | Yes | ||
| 132 | SF3B2 | 4967 | 1.495 | 0.2840 | Yes | ||
| 133 | BCAP31 | 4974 | 1.492 | 0.2873 | Yes | ||
| 134 | YWHAQ | 5008 | 1.472 | 0.2896 | Yes | ||
| 135 | NDUFS4 | 5176 | 1.368 | 0.2868 | Yes | ||
| 136 | COX6A1 | 5180 | 1.364 | 0.2902 | Yes | ||
| 137 | COX5B | 5352 | 1.262 | 0.2872 | Yes | ||
| 138 | SRSF3 | 5380 | 1.246 | 0.2897 | Yes | ||
| 139 | LSM2 | 5438 | 1.216 | 0.2911 | Yes | ||
| 140 | C1QBP | 5468 | 1.198 | 0.2936 | Yes | ||
| 141 | SMG7 | 5479 | 1.189 | 0.2967 | Yes | ||
| 142 | RALY | 5504 | 1.175 | 0.2994 | Yes | ||
| 143 | COPE | 5520 | 1.165 | 0.3023 | Yes | ||
| 144 | PSMD8 | 5541 | 1.152 | 0.3051 | Yes | ||
| 145 | IMPDH1 | 5601 | 1.119 | 0.3064 | Yes | ||
| 146 | NDUFS2 | 5642 | 1.099 | 0.3085 | Yes | ||
| 147 | NONO | 5669 | 1.083 | 0.3110 | Yes | ||
| 148 | PSMB1 | 5677 | 1.079 | 0.3143 | Yes | ||
| 149 | TMBIM6 | 5680 | 1.077 | 0.3178 | Yes | ||
| 150 | CHD4 | 5715 | 1.057 | 0.3200 | Yes | ||
| 151 | TUG1 | 5764 | 1.037 | 0.3218 | Yes | ||
| 152 | COX4I1 | 5820 | 1.005 | 0.3232 | Yes | ||
| 153 | EIF4H | 5908 | 0.968 | 0.3234 | Yes | ||
| 154 | SET | 5935 | 0.954 | 0.3260 | Yes | ||
| 155 | LSM4 | 5982 | 0.937 | 0.3278 | Yes | ||
| 156 | GANAB | 6086 | 0.901 | 0.3274 | Yes | ||
| 157 | GNG5 | 6091 | 0.899 | 0.3308 | Yes | ||
| 158 | ATP5H | 6160 | 0.873 | 0.3317 | Yes | ||
| 159 | PPM1G | 6188 | 0.863 | 0.3342 | Yes | ||
| 160 | PPP1R7 | 6202 | 0.856 | 0.3373 | Yes | ||
| 161 | ACOT7 | 6361 | 0.804 | 0.3348 | Yes | ||
| 162 | COX5A | 6384 | 0.796 | 0.3375 | Yes | ||
| 163 | DRG1 | 6387 | 0.795 | 0.3410 | Yes | ||
| 164 | CCT7 | 6486 | 0.767 | 0.3408 | Yes | ||
| 165 | CTDNEP1 | 6536 | 0.751 | 0.3425 | Yes | ||
| 166 | EIF3B | 6547 | 0.749 | 0.3456 | Yes | ||
| 167 | HSPD1 | 6645 | 0.711 | 0.3455 | Yes | ||
| 168 | EIF3I | 6971 | 0.612 | 0.3365 | Yes | ||
| 169 | CDC123 | 6998 | 0.604 | 0.3391 | Yes | ||
| 170 | XRCC6 | 7077 | 0.582 | 0.3397 | Yes | ||
| 171 | STARD7 | 7115 | 0.571 | 0.3418 | Yes | ||
| 172 | SDHA | 7134 | 0.567 | 0.3447 | Yes | ||
| 173 | POLR2I | 7137 | 0.567 | 0.3482 | Yes | ||
| 174 | CLPP | 7333 | 0.521 | 0.3442 | No | ||
| 175 | NDUFS5 | 7681 | 0.449 | 0.3345 | No | ||
| 176 | CLTA | 7765 | 0.433 | 0.3348 | No | ||
| 177 | RHEB | 7866 | 0.418 | 0.3345 | No | ||
| 178 | MPV17 | 8136 | 0.375 | 0.3278 | No | ||
| 179 | HUWE1 | 8212 | 0.366 | 0.3284 | No | ||
| 180 | CANX | 8348 | 0.345 | 0.3268 | No | ||
| 181 | DGUOK | 8411 | 0.336 | 0.3280 | No | ||
| 182 | EIF4A1 | 8417 | 0.335 | 0.3314 | No | ||
| 183 | SOD1 | 8489 | 0.323 | 0.3322 | No | ||
| 184 | EIF3K | 8552 | 0.315 | 0.3334 | No | ||
| 185 | CCT5 | 8648 | 0.304 | 0.3333 | No | ||
| 186 | CCT2 | 8673 | 0.302 | 0.3359 | No | ||
| 187 | GPX4 | 8740 | 0.292 | 0.3369 | No | ||
| 188 | H2AFV | 8893 | 0.272 | 0.3347 | No | ||
| 189 | PHB2 | 8924 | 0.269 | 0.3371 | No | ||
| 190 | EIF3C | 8949 | 0.266 | 0.3397 | No | ||
| 191 | LYPLA1 | 9080 | 0.251 | 0.3383 | No | ||
| 192 | COPS5 | 9514 | 0.206 | 0.3252 | No | ||
| 193 | PRRC2C | 9557 | 0.203 | 0.3272 | No | ||
| 194 | HNRNPC | 9576 | 0.199 | 0.3300 | No | ||
| 195 | COX6C | 9775 | 0.180 | 0.3260 | No | ||
| 196 | RUVBL2 | 9827 | 0.176 | 0.3276 | No | ||
| 197 | VPS52 | 10028 | 0.160 | 0.3235 | No | ||
| 198 | DCTN2 | 10337 | 0.135 | 0.3152 | No | ||
| 199 | ERP29 | 10341 | 0.134 | 0.3186 | No | ||
| 200 | VBP1 | 10734 | 0.107 | 0.3071 | No | ||
| 201 | CSNK2B | 10883 | 0.098 | 0.3050 | No | ||
| 202 | SNRPA1 | 11250 | 0.079 | 0.2945 | No | ||
| 203 | DHX16 | 11296 | 0.076 | 0.2963 | No | ||
| 204 | NDUFV1 | 11344 | 0.074 | 0.2981 | No | ||
| 205 | ATP5O | 11462 | 0.067 | 0.2972 | No | ||
| 206 | DEK | 11769 | 0.052 | 0.2890 | No | ||
| 207 | NDUFC1 | 11935 | 0.046 | 0.2862 | No | ||
| 208 | AP3D1 | 12055 | 0.042 | 0.2852 | No | ||
| 209 | RER1 | 12078 | 0.041 | 0.2879 | No | ||
| 210 | UQCRFS1 | 12726 | 0.014 | 0.2666 | No | ||
| 211 | SMARCD2 | 12757 | 0.012 | 0.2690 | No | ||
| 212 | VDAC2 | 12764 | 0.012 | 0.2723 | No | ||
| 213 | MAP2K2 | 12922 | 0.007 | 0.2699 | No | ||
| 214 | GMPS | 14135 | -0.038 | 0.2269 | No | ||
| 215 | SNX3 | 14460 | -0.050 | 0.2180 | No | ||
| 216 | URM1 | 14825 | -0.063 | 0.2076 | No | ||
| 217 | GNB2 | 15018 | -0.074 | 0.2037 | No | ||
| 218 | PTPN11 | 15070 | -0.077 | 0.2053 | No | ||
| 219 | UQCRB | 15308 | -0.090 | 0.1998 | No | ||
| 220 | ANP32B | 15328 | -0.092 | 0.2026 | No | ||
| 221 | DDOST | 15536 | -0.104 | 0.1982 | No | ||
| 222 | NAA10 | 15551 | -0.106 | 0.2013 | No | ||
| 223 | RAC1 | 15577 | -0.107 | 0.2039 | No | ||
| 224 | ESPL1 | 15985 | -0.133 | 0.1918 | No | ||
| 225 | ATP6AP1 | 16142 | -0.143 | 0.1894 | No | ||
| 226 | NDUFS3 | 16303 | -0.154 | 0.1868 | No | ||
| 227 | EEF1E1 | 16374 | -0.158 | 0.1876 | No | ||
| 228 | KARS | 16544 | -0.173 | 0.1847 | No | ||
| 229 | DDX49 | 16657 | -0.186 | 0.1840 | No | ||
| 230 | NDUFV2 | 16912 | -0.210 | 0.1778 | No | ||
| 231 | HDGF | 17029 | -0.220 | 0.1769 | No | ||
| 232 | GPN1 | 17156 | -0.235 | 0.1756 | No | ||
| 233 | KDELR1 | 17905 | -0.323 | 0.1504 | No | ||
| 234 | PDAP1 | 17991 | -0.335 | 0.1507 | No | ||
| 235 | TAF11 | 18072 | -0.344 | 0.1512 | No | ||
| 236 | CYC1 | 18254 | -0.373 | 0.1478 | No | ||
| 237 | KIF2A | 18670 | -0.449 | 0.1354 | No | ||
| 238 | ATP5A1 | 18824 | -0.477 | 0.1331 | No | ||
| 239 | HDDC2 | 18990 | -0.511 | 0.1303 | No | ||
| 240 | GGCT | 19002 | -0.513 | 0.1335 | No | ||
| 241 | HADHB | 19125 | -0.541 | 0.1323 | No | ||
| 242 | EIF2B4 | 19360 | -0.594 | 0.1269 | No | ||
| 243 | UQCRC2 | 19378 | -0.599 | 0.1298 | No | ||
| 244 | HADHA | 19391 | -0.605 | 0.1329 | No | ||
| 245 | PDCD6 | 19623 | -0.665 | 0.1276 | No | ||
| 246 | ACP1 | 20702 | -1.045 | 0.0898 | No | ||
| 247 | DDX1 | 20762 | -1.072 | 0.0910 | No | ||
| 248 | ATXN10 | 20784 | -1.083 | 0.0938 | No | ||
| 249 | DAP | 20798 | -1.089 | 0.0969 | No | ||
| 250 | RAD23B | 20837 | -1.110 | 0.0990 | No | ||
| 251 | TUFM | 21247 | -1.336 | 0.0868 | No | ||
| 252 | ANAPC5 | 21308 | -1.367 | 0.0881 | No | ||
| 253 | ACOT13 | 21488 | -1.477 | 0.0847 | No | ||
| 254 | DAP3 | 21675 | -1.597 | 0.0812 | No | ||
| 255 | JTB | 21732 | -1.645 | 0.0826 | No | ||
| 256 | TCEA1 | 21870 | -1.744 | 0.0809 | No | ||
| 257 | NDUFS8 | 21998 | -1.864 | 0.0795 | No | ||
| 258 | CDC23 | 22136 | -1.977 | 0.0778 | No | ||
| 259 | UBA2 | 22250 | -2.077 | 0.0771 | No | ||
| 260 | DARS | 22324 | -2.154 | 0.0778 | No | ||
| 261 | RAD23A | 22403 | -2.224 | 0.0784 | No | ||
| 262 | BUB3 | 23035 | -2.995 | 0.0577 | No | ||
| 263 | MDH1 | 23098 | -3.088 | 0.0589 | No | ||
| 264 | SRP9 | 23252 | -3.333 | 0.0566 | No | ||
| 265 | RFC2 | 23687 | -4.196 | 0.0435 | No | ||
| 266 | NDUFS1 | 23713 | -4.266 | 0.0461 | No | ||
| 267 | HSPA9 | 23764 | -4.390 | 0.0477 | No | ||
| 268 | CTBP1 | 23872 | -4.662 | 0.0471 | No | ||
| 269 | GNB1 | 23909 | -4.766 | 0.0493 | No | ||
| 270 | DDX19B | 23940 | -4.850 | 0.0517 | No | ||
| 271 | MTDH | 24386 | -6.334 | 0.0382 | No | ||
| 272 | SNRPA | 24600 | -7.237 | 0.0336 | No | ||
| 273 | MRPS18B | 24611 | -7.295 | 0.0367 | No | ||
| 274 | NARS | 24711 | -7.711 | 0.0365 | No | ||
| 275 | RAD21 | 24731 | -7.837 | 0.0393 | No | ||
| 276 | SLC3A2 | 25078 | -10.370 | 0.0296 | No | ||
| 277 | GLB1 | 25723 | -20.684 | 0.0084 | No | ||
| 278 | LARP1 | 25910 | -28.139 | 0.0048 | No | ||
| 279 | HAX1 | 26046 | -38.579 | 0.0032 | No | ||
| 280 | SLC1A5 | 26086 | -42.020 | 0.0053 | No | ||
| 281 | IFRD1 | 26303 | -230.785 | 0.0005 | No |