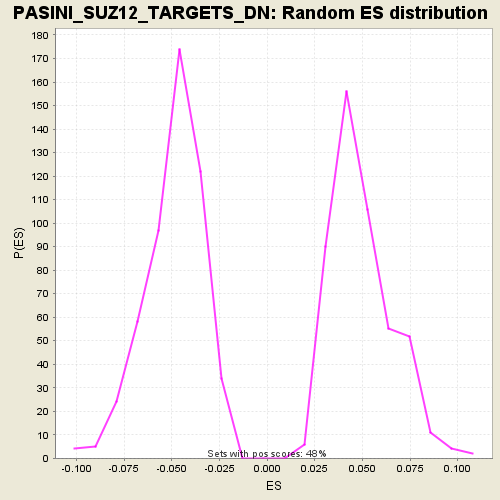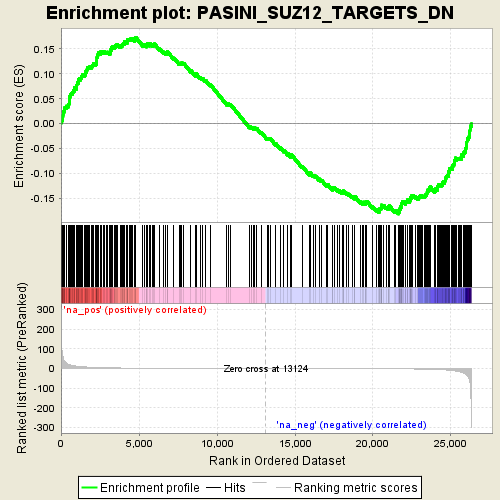
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | PASINI_SUZ12_TARGETS_DN |
| Enrichment Score (ES) | -0.18094389 |
| Normalized Enrichment Score (NES) | -3.7133865 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CDKN1A | 2 | 299.207 | 0.0032 | No | ||
| 2 | FLNC | 35 | 112.692 | 0.0052 | No | ||
| 3 | BTG2 | 60 | 90.877 | 0.0076 | No | ||
| 4 | TUBB6 | 76 | 77.923 | 0.0103 | No | ||
| 5 | IDI1 | 88 | 70.501 | 0.0131 | No | ||
| 6 | DPYSL2 | 110 | 63.090 | 0.0156 | No | ||
| 7 | COL3A1 | 124 | 59.189 | 0.0183 | No | ||
| 8 | BMP4 | 130 | 58.085 | 0.0214 | No | ||
| 9 | CITED2 | 143 | 54.021 | 0.0242 | No | ||
| 10 | SDC1 | 189 | 42.210 | 0.0258 | No | ||
| 11 | LRP8 | 208 | 39.535 | 0.0283 | No | ||
| 12 | RBP1 | 229 | 37.474 | 0.0308 | No | ||
| 13 | RAP2B | 246 | 35.107 | 0.0335 | No | ||
| 14 | KCTD10 | 348 | 26.459 | 0.0329 | No | ||
| 15 | CMTM7 | 365 | 25.490 | 0.0355 | No | ||
| 16 | ARL4C | 445 | 22.258 | 0.0358 | No | ||
| 17 | PLK2 | 461 | 21.783 | 0.0384 | No | ||
| 18 | AHNAK | 507 | 20.432 | 0.0400 | No | ||
| 19 | TCEAL8 | 522 | 20.067 | 0.0427 | No | ||
| 20 | TUBA1A | 529 | 19.799 | 0.0458 | No | ||
| 21 | SYT11 | 548 | 19.376 | 0.0483 | No | ||
| 22 | EFNA5 | 553 | 19.297 | 0.0514 | No | ||
| 23 | CYFIP2 | 556 | 19.234 | 0.0546 | No | ||
| 24 | MLLT11 | 588 | 18.479 | 0.0567 | No | ||
| 25 | SFN | 603 | 18.262 | 0.0594 | No | ||
| 26 | NCS1 | 642 | 17.461 | 0.0612 | No | ||
| 27 | VCAN | 712 | 16.078 | 0.0619 | No | ||
| 28 | ACTN1 | 757 | 15.349 | 0.0634 | No | ||
| 29 | COL5A1 | 771 | 15.118 | 0.0662 | No | ||
| 30 | CCNG1 | 836 | 14.184 | 0.0670 | No | ||
| 31 | CAV1 | 863 | 13.935 | 0.0693 | No | ||
| 32 | CRYAB | 869 | 13.777 | 0.0724 | No | ||
| 33 | COL1A1 | 966 | 12.528 | 0.0719 | No | ||
| 34 | CRLF1 | 985 | 12.338 | 0.0745 | No | ||
| 35 | SOAT1 | 994 | 12.245 | 0.0775 | No | ||
| 36 | MMP14 | 1020 | 11.858 | 0.0798 | No | ||
| 37 | TIMP2 | 1029 | 11.763 | 0.0827 | No | ||
| 38 | SCD | 1102 | 11.098 | 0.0832 | No | ||
| 39 | METRNL | 1118 | 10.991 | 0.0859 | No | ||
| 40 | GNAS | 1126 | 10.933 | 0.0889 | No | ||
| 41 | FAM84B | 1162 | 10.639 | 0.0909 | No | ||
| 42 | ELOVL1 | 1253 | 9.905 | 0.0907 | No | ||
| 43 | DBN1 | 1280 | 9.747 | 0.0929 | No | ||
| 44 | TPM1 | 1322 | 9.453 | 0.0946 | No | ||
| 45 | PERP | 1343 | 9.287 | 0.0971 | No | ||
| 46 | SMO | 1375 | 8.997 | 0.0992 | No | ||
| 47 | TPM2 | 1517 | 8.213 | 0.0970 | No | ||
| 48 | PITX2 | 1540 | 8.144 | 0.0995 | No | ||
| 49 | TMEM47 | 1564 | 7.957 | 0.1018 | No | ||
| 50 | KDELC1 | 1582 | 7.831 | 0.1045 | No | ||
| 51 | RAB34 | 1629 | 7.594 | 0.1060 | No | ||
| 52 | SP5 | 1661 | 7.460 | 0.1080 | No | ||
| 53 | MYADM | 1674 | 7.417 | 0.1108 | No | ||
| 54 | SOX4 | 1692 | 7.315 | 0.1135 | No | ||
| 55 | TMEM30A | 1783 | 6.901 | 0.1133 | No | ||
| 56 | MYH10 | 1847 | 6.647 | 0.1141 | No | ||
| 57 | EFEMP1 | 1926 | 6.296 | 0.1144 | No | ||
| 58 | ACTC1 | 1983 | 6.115 | 0.1155 | No | ||
| 59 | FLNA | 2016 | 5.990 | 0.1175 | No | ||
| 60 | PKP2 | 2059 | 5.870 | 0.1192 | No | ||
| 61 | EID1 | 2086 | 5.754 | 0.1215 | No | ||
| 62 | LGALS1 | 2213 | 5.365 | 0.1199 | No | ||
| 63 | HMGA2 | 2268 | 5.208 | 0.1211 | No | ||
| 64 | CTXN1 | 2269 | 5.207 | 0.1243 | No | ||
| 65 | VIM | 2293 | 5.120 | 0.1267 | No | ||
| 66 | ETS1 | 2298 | 5.112 | 0.1298 | No | ||
| 67 | KLHL9 | 2299 | 5.112 | 0.1331 | No | ||
| 68 | ACTA2 | 2320 | 5.036 | 0.1356 | No | ||
| 69 | CNN1 | 2347 | 4.991 | 0.1379 | No | ||
| 70 | FRMD4B | 2368 | 4.923 | 0.1404 | No | ||
| 71 | ERRFI1 | 2394 | 4.861 | 0.1427 | No | ||
| 72 | DUSP4 | 2513 | 4.608 | 0.1414 | No | ||
| 73 | GNG3 | 2534 | 4.568 | 0.1439 | No | ||
| 74 | SPSB1 | 2604 | 4.410 | 0.1445 | No | ||
| 75 | CORO1C | 2725 | 4.138 | 0.1432 | No | ||
| 76 | COL11A1 | 2781 | 4.021 | 0.1443 | No | ||
| 77 | ANXA5 | 2892 | 3.799 | 0.1434 | No | ||
| 78 | DYRK2 | 2969 | 3.658 | 0.1437 | No | ||
| 79 | KLF7 | 3118 | 3.421 | 0.1413 | No | ||
| 80 | PPP1R18 | 3162 | 3.341 | 0.1429 | No | ||
| 81 | CD151 | 3170 | 3.333 | 0.1459 | No | ||
| 82 | IER5 | 3192 | 3.300 | 0.1484 | No | ||
| 83 | APP | 3222 | 3.246 | 0.1505 | No | ||
| 84 | TNFRSF19 | 3250 | 3.207 | 0.1528 | No | ||
| 85 | CAV2 | 3275 | 3.178 | 0.1551 | No | ||
| 86 | TGFB2 | 3406 | 2.979 | 0.1534 | No | ||
| 87 | CAP1 | 3442 | 2.937 | 0.1553 | No | ||
| 88 | WISP1 | 3499 | 2.869 | 0.1564 | No | ||
| 89 | RTN3 | 3543 | 2.806 | 0.1580 | No | ||
| 90 | CD276 | 3599 | 2.752 | 0.1592 | No | ||
| 91 | CLIC1 | 3784 | 2.545 | 0.1554 | No | ||
| 92 | MTPN | 3843 | 2.477 | 0.1564 | No | ||
| 93 | ACSL4 | 3911 | 2.397 | 0.1571 | No | ||
| 94 | DDX6 | 3934 | 2.383 | 0.1595 | No | ||
| 95 | IRS1 | 3974 | 2.342 | 0.1613 | No | ||
| 96 | COLEC12 | 4067 | 2.247 | 0.1610 | No | ||
| 97 | PRSS23 | 4069 | 2.246 | 0.1643 | No | ||
| 98 | THBS1 | 4168 | 2.135 | 0.1638 | No | ||
| 99 | ABTB2 | 4231 | 2.076 | 0.1646 | No | ||
| 100 | PLOD2 | 4236 | 2.069 | 0.1678 | No | ||
| 101 | PRPF40A | 4277 | 2.035 | 0.1695 | No | ||
| 102 | CMTM3 | 4372 | 1.957 | 0.1691 | No | ||
| 103 | T | 4437 | 1.894 | 0.1699 | No | ||
| 104 | PIK3R3 | 4491 | 1.856 | 0.1712 | No | ||
| 105 | POU3F1 | 4593 | 1.779 | 0.1706 | No | ||
| 106 | VAT1 | 4737 | 1.664 | 0.1683 | No | ||
| 107 | STEAP1 | 4752 | 1.651 | 0.1711 | No | ||
| 108 | RCN2 | 4792 | 1.626 | 0.1728 | No | ||
| 109 | S100A6 | 5248 | 1.327 | 0.1586 | No | ||
| 110 | MARCKS | 5326 | 1.277 | 0.1589 | No | ||
| 111 | CYR61 | 5463 | 1.203 | 0.1570 | No | ||
| 112 | ANXA1 | 5495 | 1.180 | 0.1590 | No | ||
| 113 | SERPINH1 | 5549 | 1.146 | 0.1603 | No | ||
| 114 | ZMYND8 | 5671 | 1.083 | 0.1589 | No | ||
| 115 | CALD1 | 5718 | 1.056 | 0.1604 | No | ||
| 116 | TPM4 | 5857 | 0.985 | 0.1583 | No | ||
| 117 | REEP3 | 5932 | 0.957 | 0.1588 | No | ||
| 118 | AXL | 5986 | 0.935 | 0.1600 | No | ||
| 119 | CDKN2B | 6331 | 0.813 | 0.1500 | No | ||
| 120 | S100A10 | 6578 | 0.733 | 0.1438 | No | ||
| 121 | VCL | 6710 | 0.684 | 0.1421 | No | ||
| 122 | MSN | 6800 | 0.660 | 0.1419 | No | ||
| 123 | PDLIM7 | 6822 | 0.652 | 0.1444 | No | ||
| 124 | PARVA | 7235 | 0.541 | 0.1318 | No | ||
| 125 | SOX11 | 7600 | 0.463 | 0.1211 | No | ||
| 126 | PXDN | 7673 | 0.452 | 0.1216 | No | ||
| 127 | CGN | 7749 | 0.437 | 0.1220 | No | ||
| 128 | CLDN12 | 7855 | 0.420 | 0.1212 | No | ||
| 129 | KIF5C | 8327 | 0.351 | 0.1064 | No | ||
| 130 | HMGN3 | 8625 | 0.306 | 0.0982 | No | ||
| 131 | SLC39A6 | 8652 | 0.303 | 0.1005 | No | ||
| 132 | SEMA3C | 8932 | 0.268 | 0.0930 | No | ||
| 133 | FLRT3 | 9066 | 0.252 | 0.0912 | No | ||
| 134 | NR6A1 | 9262 | 0.231 | 0.0870 | No | ||
| 135 | GMFB | 9562 | 0.202 | 0.0787 | No | ||
| 136 | BMP1 | 10639 | 0.114 | 0.0406 | No | ||
| 137 | FARP1 | 10728 | 0.108 | 0.0405 | No | ||
| 138 | CSNK1G1 | 10856 | 0.099 | 0.0389 | No | ||
| 139 | LOXL2 | 12093 | 0.040 | -0.0053 | No | ||
| 140 | ILK | 12227 | 0.034 | -0.0072 | No | ||
| 141 | TRIB1 | 12362 | 0.028 | -0.0091 | No | ||
| 142 | LPAR4 | 12403 | 0.027 | -0.0073 | No | ||
| 143 | PHIP | 12537 | 0.021 | -0.0092 | No | ||
| 144 | IRS2 | 12823 | 0.009 | -0.0169 | No | ||
| 145 | MYO1C | 13233 | -0.004 | -0.0293 | No | ||
| 146 | NPNT | 13308 | -0.006 | -0.0289 | No | ||
| 147 | LPP | 13403 | -0.008 | -0.0293 | No | ||
| 148 | SHB | 13773 | -0.023 | -0.0402 | No | ||
| 149 | DUSP14 | 14069 | -0.035 | -0.0482 | No | ||
| 150 | TMEM43 | 14291 | -0.044 | -0.0535 | No | ||
| 151 | ITGB1 | 14539 | -0.052 | -0.0597 | No | ||
| 152 | SCOC | 14740 | -0.059 | -0.0641 | No | ||
| 153 | KPNA4 | 14759 | -0.060 | -0.0615 | No | ||
| 154 | IGFBP4 | 15456 | -0.099 | -0.0850 | No | ||
| 155 | LIMD1 | 15927 | -0.129 | -0.0998 | No | ||
| 156 | KLHDC2 | 15970 | -0.131 | -0.0982 | No | ||
| 157 | SYNPO2L | 16175 | -0.146 | -0.1028 | No | ||
| 158 | GNG2 | 16311 | -0.154 | -0.1047 | No | ||
| 159 | SOCS6 | 16552 | -0.174 | -0.1106 | No | ||
| 160 | BNIP2 | 16702 | -0.189 | -0.1131 | No | ||
| 161 | GBP2 | 17046 | -0.222 | -0.1230 | No | ||
| 162 | MALAT1 | 17089 | -0.227 | -0.1214 | No | ||
| 163 | ITGAV | 17402 | -0.264 | -0.1301 | No | ||
| 164 | GBP1 | 17409 | -0.264 | -0.1271 | No | ||
| 165 | DSP | 17506 | -0.275 | -0.1275 | No | ||
| 166 | WNT4 | 17725 | -0.300 | -0.1326 | No | ||
| 167 | PHLDB2 | 17848 | -0.317 | -0.1340 | No | ||
| 168 | ARAP3 | 18025 | -0.339 | -0.1375 | No | ||
| 169 | SUZ12 | 18093 | -0.347 | -0.1368 | No | ||
| 170 | ANXA2 | 18120 | -0.351 | -0.1345 | No | ||
| 171 | LRRFIP1 | 18317 | -0.384 | -0.1388 | No | ||
| 172 | TTYH3 | 18453 | -0.409 | -0.1407 | No | ||
| 173 | ZDHHC9 | 18658 | -0.447 | -0.1453 | No | ||
| 174 | B2M | 18808 | -0.474 | -0.1478 | No | ||
| 175 | CA3 | 18822 | -0.476 | -0.1450 | No | ||
| 176 | STEAP2 | 19202 | -0.556 | -0.1563 | No | ||
| 177 | AKT1 | 19355 | -0.593 | -0.1589 | No | ||
| 178 | SH3BGRL | 19388 | -0.604 | -0.1568 | No | ||
| 179 | SEMA3E | 19544 | -0.646 | -0.1595 | No | ||
| 180 | YAF2 | 19551 | -0.649 | -0.1565 | No | ||
| 181 | FN1 | 19610 | -0.662 | -0.1555 | No | ||
| 182 | BDNF | 19969 | -0.773 | -0.1660 | No | ||
| 183 | CFL2 | 20208 | -0.852 | -0.1718 | No | ||
| 184 | FOXP1 | 20351 | -0.904 | -0.1740 | No | ||
| 185 | CSF1 | 20430 | -0.932 | -0.1738 | No | ||
| 186 | ZNF266 | 20435 | -0.935 | -0.1706 | No | ||
| 187 | WNK1 | 20487 | -0.956 | -0.1693 | No | ||
| 188 | F3 | 20529 | -0.973 | -0.1676 | No | ||
| 189 | STX6 | 20551 | -0.981 | -0.1652 | No | ||
| 190 | BAZ1A | 20575 | -0.991 | -0.1628 | No | ||
| 191 | HBEGF | 20687 | -1.038 | -0.1638 | No | ||
| 192 | TMBIM1 | 20872 | -1.127 | -0.1676 | No | ||
| 193 | GAP43 | 20993 | -1.186 | -0.1690 | No | ||
| 194 | CDH2 | 21016 | -1.199 | -0.1665 | No | ||
| 195 | COL1A2 | 21039 | -1.211 | -0.1641 | No | ||
| 196 | AMMECR1L | 21409 | -1.426 | -0.1750 | No | ||
| 197 | TAGLN2 | 21447 | -1.451 | -0.1732 | No | ||
| 198 | FILIP1L | 21650 | -1.578 | -0.1777 | Yes | ||
| 199 | VGLL3 | 21679 | -1.602 | -0.1755 | Yes | ||
| 200 | TROVE2 | 21694 | -1.616 | -0.1728 | Yes | ||
| 201 | PHC2 | 21729 | -1.640 | -0.1708 | Yes | ||
| 202 | LIMD2 | 21769 | -1.661 | -0.1690 | Yes | ||
| 203 | AMOTL2 | 21778 | -1.669 | -0.1661 | Yes | ||
| 204 | PPBP | 21835 | -1.716 | -0.1649 | Yes | ||
| 205 | CDK6 | 21848 | -1.723 | -0.1621 | Yes | ||
| 206 | DUSP1 | 21853 | -1.729 | -0.1590 | Yes | ||
| 207 | LPGAT1 | 21867 | -1.740 | -0.1563 | Yes | ||
| 208 | KRT19 | 21947 | -1.819 | -0.1560 | Yes | ||
| 209 | ITGA3 | 22089 | -1.937 | -0.1582 | Yes | ||
| 210 | ZYX | 22093 | -1.938 | -0.1550 | Yes | ||
| 211 | RPS6KA3 | 22191 | -2.024 | -0.1555 | Yes | ||
| 212 | LTB | 22199 | -2.030 | -0.1525 | Yes | ||
| 213 | IGFBP5 | 22359 | -2.187 | -0.1553 | Yes | ||
| 214 | PDLIM3 | 22376 | -2.196 | -0.1527 | Yes | ||
| 215 | DOCK11 | 22400 | -2.220 | -0.1503 | Yes | ||
| 216 | WWTR1 | 22417 | -2.253 | -0.1476 | Yes | ||
| 217 | UBE2J1 | 22476 | -2.310 | -0.1466 | Yes | ||
| 218 | GHR | 22496 | -2.324 | -0.1441 | Yes | ||
| 219 | RDH10 | 22550 | -2.377 | -0.1428 | Yes | ||
| 220 | NT5E | 22714 | -2.579 | -0.1458 | Yes | ||
| 221 | GPRC5A | 22859 | -2.784 | -0.1481 | Yes | ||
| 222 | SERTAD4 | 22944 | -2.878 | -0.1481 | Yes | ||
| 223 | CXADR | 22966 | -2.896 | -0.1456 | Yes | ||
| 224 | GNG12 | 23018 | -2.969 | -0.1443 | Yes | ||
| 225 | GSN | 23103 | -3.090 | -0.1443 | Yes | ||
| 226 | GLIPR2 | 23201 | -3.247 | -0.1447 | Yes | ||
| 227 | KLF6 | 23308 | -3.435 | -0.1455 | Yes | ||
| 228 | SLC25A24 | 23341 | -3.497 | -0.1435 | Yes | ||
| 229 | NUAK1 | 23392 | -3.606 | -0.1421 | Yes | ||
| 230 | RYK | 23457 | -3.713 | -0.1413 | Yes | ||
| 231 | PEA15 | 23464 | -3.719 | -0.1383 | Yes | ||
| 232 | TNFRSF12A | 23483 | -3.763 | -0.1357 | Yes | ||
| 233 | ID2 | 23493 | -3.773 | -0.1328 | Yes | ||
| 234 | AMFR | 23576 | -3.961 | -0.1327 | Yes | ||
| 235 | AIF1L | 23603 | -4.006 | -0.1304 | Yes | ||
| 236 | CNN2 | 23635 | -4.087 | -0.1283 | Yes | ||
| 237 | PKIA | 23666 | -4.143 | -0.1262 | Yes | ||
| 238 | TSPAN2 | 23977 | -4.952 | -0.1349 | Yes | ||
| 239 | COL4A5 | 23997 | -5.007 | -0.1323 | Yes | ||
| 240 | TAGLN | 24035 | -5.137 | -0.1305 | Yes | ||
| 241 | PTPN12 | 24128 | -5.376 | -0.1308 | Yes | ||
| 242 | BHLHE40 | 24138 | -5.407 | -0.1278 | Yes | ||
| 243 | MARVELD2 | 24147 | -5.433 | -0.1249 | Yes | ||
| 244 | GLI3 | 24171 | -5.497 | -0.1225 | Yes | ||
| 245 | CPE | 24236 | -5.705 | -0.1217 | Yes | ||
| 246 | GPC4 | 24304 | -6.010 | -0.1210 | Yes | ||
| 247 | PLIN2 | 24409 | -6.435 | -0.1217 | Yes | ||
| 248 | FOSL2 | 24470 | -6.617 | -0.1208 | Yes | ||
| 249 | CYP1B1 | 24480 | -6.654 | -0.1178 | Yes | ||
| 250 | PAWR | 24519 | -6.853 | -0.1160 | Yes | ||
| 251 | PDLIM5 | 24618 | -7.323 | -0.1165 | Yes | ||
| 252 | PRTG | 24621 | -7.344 | -0.1133 | Yes | ||
| 253 | TSPAN9 | 24636 | -7.401 | -0.1106 | Yes | ||
| 254 | ADK | 24669 | -7.527 | -0.1086 | Yes | ||
| 255 | RRAS2 | 24687 | -7.594 | -0.1060 | Yes | ||
| 256 | SKIL | 24743 | -7.924 | -0.1048 | Yes | ||
| 257 | ABHD2 | 24791 | -8.195 | -0.1033 | Yes | ||
| 258 | TMCC3 | 24847 | -8.524 | -0.1022 | Yes | ||
| 259 | EPHA1 | 24849 | -8.535 | -0.0990 | Yes | ||
| 260 | PMP22 | 24857 | -8.599 | -0.0960 | Yes | ||
| 261 | IER3 | 24909 | -8.968 | -0.0947 | Yes | ||
| 262 | TMEM123 | 24929 | -9.121 | -0.0921 | Yes | ||
| 263 | SH3GLB1 | 24930 | -9.123 | -0.0889 | Yes | ||
| 264 | WLS | 25009 | -9.776 | -0.0886 | Yes | ||
| 265 | RHOB | 25084 | -10.442 | -0.0882 | Yes | ||
| 266 | KRT8 | 25119 | -10.758 | -0.0862 | Yes | ||
| 267 | RHOBTB3 | 25121 | -10.806 | -0.0830 | Yes | ||
| 268 | RBMS1 | 25170 | -11.277 | -0.0815 | Yes | ||
| 269 | BMP2 | 25202 | -11.606 | -0.0795 | Yes | ||
| 270 | AMOTL1 | 25211 | -11.663 | -0.0765 | Yes | ||
| 271 | KRT18 | 25215 | -11.722 | -0.0734 | Yes | ||
| 272 | MUM1L1 | 25261 | -12.127 | -0.0718 | Yes | ||
| 273 | JUN | 25262 | -12.127 | -0.0686 | Yes | ||
| 274 | RND3 | 25387 | -13.780 | -0.0701 | Yes | ||
| 275 | UBE2E1 | 25456 | -14.805 | -0.0694 | Yes | ||
| 276 | PRNP | 25527 | -15.875 | -0.0688 | Yes | ||
| 277 | DUSP6 | 25608 | -17.968 | -0.0686 | Yes | ||
| 278 | JAG1 | 25675 | -19.325 | -0.0679 | Yes | ||
| 279 | STMN2 | 25677 | -19.359 | -0.0647 | Yes | ||
| 280 | BACH2 | 25702 | -19.987 | -0.0623 | Yes | ||
| 281 | TCF12 | 25794 | -22.666 | -0.0626 | Yes | ||
| 282 | HS6ST2 | 25812 | -23.055 | -0.0599 | Yes | ||
| 283 | CRMP1 | 25829 | -24.013 | -0.0573 | Yes | ||
| 284 | TMPRSS2 | 25888 | -26.668 | -0.0562 | Yes | ||
| 285 | FZD2 | 25914 | -28.452 | -0.0539 | Yes | ||
| 286 | GRB10 | 25941 | -30.158 | -0.0517 | Yes | ||
| 287 | FAM107B | 25950 | -30.439 | -0.0487 | Yes | ||
| 288 | PROM1 | 25970 | -31.719 | -0.0462 | Yes | ||
| 289 | PALLD | 25978 | -32.108 | -0.0432 | Yes | ||
| 290 | TGFB1I1 | 25990 | -33.266 | -0.0403 | Yes | ||
| 291 | IGFBP3 | 26019 | -35.792 | -0.0381 | Yes | ||
| 292 | ANXA3 | 26030 | -37.029 | -0.0353 | Yes | ||
| 293 | TES | 26033 | -37.078 | -0.0321 | Yes | ||
| 294 | PDGFB | 26055 | -39.111 | -0.0296 | Yes | ||
| 295 | SAMD4A | 26129 | -47.887 | -0.0291 | Yes | ||
| 296 | NEFL | 26149 | -51.235 | -0.0266 | Yes | ||
| 297 | SERPINB9 | 26165 | -53.978 | -0.0239 | Yes | ||
| 298 | PHLDA1 | 26170 | -55.684 | -0.0208 | Yes | ||
| 299 | SETD7 | 26173 | -56.730 | -0.0176 | Yes | ||
| 300 | CSRP1 | 26194 | -61.077 | -0.0151 | Yes | ||
| 301 | CAPN2 | 26220 | -71.517 | -0.0128 | Yes | ||
| 302 | MYC | 26225 | -75.265 | -0.0097 | Yes | ||
| 303 | PRICKLE1 | 26240 | -83.886 | -0.0070 | Yes | ||
| 304 | SULF1 | 26261 | -97.588 | -0.0045 | Yes | ||
| 305 | ACTA1 | 26309 | -299.207 | -0.0030 | Yes | ||
| 306 | MEG3 | 26313 | -299.207 | 0.0002 | Yes |