Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | NS50832_padj |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | VECCHI_GASTRIC_CANCER_EARLY_DN |
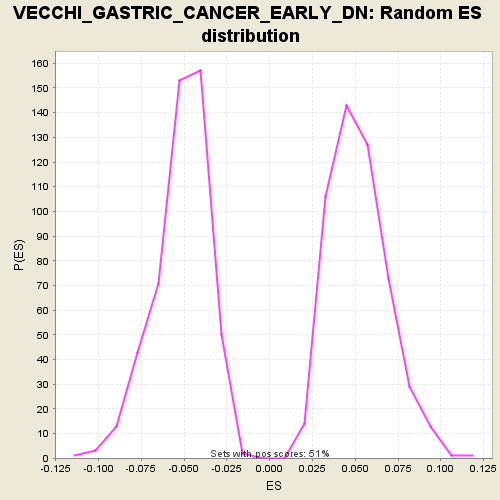
| Enrichment Score (ES) | -0.2122346 |
| Normalized Enrichment Score (NES) | -4.108077 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PLP1 | 31 | 117.747 | 0.0025 | No | ||
| 2 | GLUL | 84 | 72.529 | 0.0041 | No | ||
| 3 | CITED2 | 143 | 54.021 | 0.0055 | No | ||
| 4 | PCDH7 | 146 | 52.996 | 0.0091 | No | ||
| 5 | IRX2 | 176 | 44.698 | 0.0116 | No | ||
| 6 | C14orf132 | 195 | 41.676 | 0.0146 | No | ||
| 7 | FHL1 | 544 | 19.472 | 0.0049 | No | ||
| 8 | WIF1 | 644 | 17.424 | 0.0047 | No | ||
| 9 | INA | 875 | 13.684 | -0.0004 | No | ||
| 10 | HHIP | 1129 | 10.894 | -0.0065 | No | ||
| 11 | UNC5D | 1318 | 9.490 | -0.0101 | No | ||
| 12 | PRIMA1 | 1362 | 9.132 | -0.0081 | No | ||
| 13 | SOX2 | 1391 | 8.931 | -0.0055 | No | ||
| 14 | SYNM | 1396 | 8.885 | -0.0020 | No | ||
| 15 | TTLL7 | 1401 | 8.863 | 0.0015 | No | ||
| 16 | BEX2 | 1464 | 8.495 | 0.0028 | No | ||
| 17 | CHGB | 1472 | 8.437 | 0.0061 | No | ||
| 18 | TMEM47 | 1564 | 7.957 | 0.0063 | No | ||
| 19 | MAGEH1 | 1585 | 7.816 | 0.0092 | No | ||
| 20 | YIF1B | 1622 | 7.635 | 0.0114 | No | ||
| 21 | SEMA4A | 1641 | 7.532 | 0.0144 | No | ||
| 22 | ARMCX2 | 1849 | 6.638 | 0.0101 | No | ||
| 23 | IRX3 | 1911 | 6.352 | 0.0114 | No | ||
| 24 | EFEMP1 | 1926 | 6.296 | 0.0145 | No | ||
| 25 | FAM46C | 2099 | 5.720 | 0.0116 | No | ||
| 26 | VEPH1 | 2143 | 5.596 | 0.0136 | No | ||
| 27 | LRRC17 | 2205 | 5.393 | 0.0149 | No | ||
| 28 | ENPP2 | 2284 | 5.147 | 0.0155 | No | ||
| 29 | CNN1 | 2347 | 4.991 | 0.0168 | No | ||
| 30 | MAF | 2406 | 4.831 | 0.0182 | No | ||
| 31 | CLDND1 | 2469 | 4.682 | 0.0195 | No | ||
| 32 | SRPX | 2743 | 4.103 | 0.0127 | No | ||
| 33 | SSTR1 | 2810 | 3.957 | 0.0138 | No | ||
| 34 | AQP4 | 3001 | 3.616 | 0.0101 | No | ||
| 35 | GPX3 | 3011 | 3.596 | 0.0134 | No | ||
| 36 | SLC2A12 | 3012 | 3.591 | 0.0171 | No | ||
| 37 | CLU | 3249 | 3.207 | 0.0117 | No | ||
| 38 | DTNA | 3295 | 3.148 | 0.0136 | No | ||
| 39 | NTN4 | 3301 | 3.137 | 0.0170 | No | ||
| 40 | SVEP1 | 3302 | 3.134 | 0.0207 | No | ||
| 41 | DNER | 3305 | 3.131 | 0.0243 | No | ||
| 42 | NR3C2 | 3899 | 2.409 | 0.0052 | No | ||
| 43 | PCDH9 | 3903 | 2.405 | 0.0087 | No | ||
| 44 | RGS4 | 4025 | 2.299 | 0.0077 | No | ||
| 45 | SST | 4210 | 2.093 | 0.0043 | No | ||
| 46 | BEX5 | 4226 | 2.079 | 0.0073 | No | ||
| 47 | RAI2 | 4306 | 2.011 | 0.0080 | No | ||
| 48 | MANEA | 4370 | 1.958 | 0.0092 | No | ||
| 49 | LDHB | 4389 | 1.937 | 0.0122 | No | ||
| 50 | SIDT2 | 4896 | 1.549 | -0.0036 | No | ||
| 51 | SLC36A4 | 4909 | 1.539 | -0.0004 | No | ||
| 52 | PPP2R3A | 5017 | 1.464 | -0.0009 | No | ||
| 53 | LIFR | 5046 | 1.441 | 0.0017 | No | ||
| 54 | KIAA1324 | 5368 | 1.250 | -0.0070 | No | ||
| 55 | PDGFC | 5424 | 1.228 | -0.0055 | No | ||
| 56 | GRIP2 | 5543 | 1.151 | -0.0063 | No | ||
| 57 | VSTM2A | 5566 | 1.140 | -0.0035 | No | ||
| 58 | TMEFF2 | 5631 | 1.104 | -0.0023 | No | ||
| 59 | ST8SIA4 | 5769 | 1.034 | -0.0040 | No | ||
| 60 | FCRL5 | 5974 | 0.940 | -0.0081 | No | ||
| 61 | ENTPD3 | 6150 | 0.877 | -0.0112 | No | ||
| 62 | TMPRSS6 | 6348 | 0.807 | -0.0151 | No | ||
| 63 | RNLS | 6427 | 0.786 | -0.0145 | No | ||
| 64 | ACOX2 | 6529 | 0.753 | -0.0147 | No | ||
| 65 | PLIN1 | 6544 | 0.750 | -0.0116 | No | ||
| 66 | PGC | 6615 | 0.720 | -0.0106 | No | ||
| 67 | SFRP1 | 6633 | 0.715 | -0.0076 | No | ||
| 68 | TCEAL2 | 6687 | 0.691 | -0.0060 | No | ||
| 69 | ID4 | 6813 | 0.655 | -0.0072 | No | ||
| 70 | USP51 | 6818 | 0.653 | -0.0037 | No | ||
| 71 | GADD45B | 7123 | 0.569 | -0.0117 | No | ||
| 72 | PBLD | 7187 | 0.552 | -0.0105 | No | ||
| 73 | ATP4A | 7308 | 0.528 | -0.0114 | No | ||
| 74 | BEND5 | 7341 | 0.519 | -0.0090 | No | ||
| 75 | GNG7 | 7356 | 0.517 | -0.0059 | No | ||
| 76 | TMED6 | 7438 | 0.500 | -0.0053 | No | ||
| 77 | SOBP | 7549 | 0.475 | -0.0059 | No | ||
| 78 | CYBRD1 | 7823 | 0.424 | -0.0127 | No | ||
| 79 | MCTP1 | 7915 | 0.410 | -0.0126 | No | ||
| 80 | UBL3 | 8185 | 0.369 | -0.0193 | No | ||
| 81 | SCARA5 | 8199 | 0.368 | -0.0161 | No | ||
| 82 | NLRP7 | 8243 | 0.362 | -0.0141 | No | ||
| 83 | APOA1 | 8324 | 0.352 | -0.0135 | No | ||
| 84 | KIF5C | 8327 | 0.351 | -0.0100 | No | ||
| 85 | BNIP3 | 8337 | 0.349 | -0.0067 | No | ||
| 86 | GMPR | 8477 | 0.324 | -0.0084 | No | ||
| 87 | GATA5 | 8526 | 0.318 | -0.0065 | No | ||
| 88 | GSPT2 | 8627 | 0.306 | -0.0067 | No | ||
| 89 | ACE2 | 8733 | 0.293 | -0.0071 | No | ||
| 90 | SLC16A7 | 8846 | 0.279 | -0.0078 | No | ||
| 91 | MTTP | 8870 | 0.275 | -0.0050 | No | ||
| 92 | SLC28A2 | 8951 | 0.266 | -0.0044 | No | ||
| 93 | ADRB2 | 9079 | 0.251 | -0.0057 | No | ||
| 94 | METTL7A | 9196 | 0.240 | -0.0065 | No | ||
| 95 | ABAT | 9210 | 0.238 | -0.0033 | No | ||
| 96 | NME5 | 9428 | 0.214 | -0.0080 | No | ||
| 97 | HLA-DOB | 9701 | 0.186 | -0.0148 | No | ||
| 98 | PRKACB | 9831 | 0.176 | -0.0161 | No | ||
| 99 | SMARCA1 | 9993 | 0.163 | -0.0186 | No | ||
| 100 | TPCN2 | 10142 | 0.149 | -0.0207 | No | ||
| 101 | DNAJC6 | 10177 | 0.146 | -0.0183 | No | ||
| 102 | CYP2C18 | 10892 | 0.098 | -0.0421 | No | ||
| 103 | CYP3A4 | 10973 | 0.094 | -0.0415 | No | ||
| 104 | LYVE1 | 11148 | 0.084 | -0.0445 | No | ||
| 105 | LONRF2 | 11804 | 0.051 | -0.0660 | No | ||
| 106 | KCNJ15 | 11867 | 0.048 | -0.0648 | No | ||
| 107 | BEX1 | 12077 | 0.041 | -0.0691 | No | ||
| 108 | MAMDC2 | 12250 | 0.033 | -0.0721 | No | ||
| 109 | RBPMS2 | 12285 | 0.032 | -0.0697 | No | ||
| 110 | MYRIP | 12562 | 0.020 | -0.0767 | No | ||
| 111 | MFAP5 | 12936 | 0.006 | -0.0874 | No | ||
| 112 | ARHGAP24 | 13093 | 0.001 | -0.0897 | No | ||
| 113 | ENTPD5 | 13316 | -0.006 | -0.0946 | No | ||
| 114 | IDUA | 13424 | -0.009 | -0.0950 | No | ||
| 115 | FKBP5 | 13452 | -0.010 | -0.0924 | No | ||
| 116 | CKMT2 | 13563 | -0.015 | -0.0930 | No | ||
| 117 | RNF180 | 13660 | -0.018 | -0.0930 | No | ||
| 118 | PDK4 | 13791 | -0.024 | -0.0944 | No | ||
| 119 | MAP9 | 14381 | -0.047 | -0.1133 | No | ||
| 120 | SLC26A7 | 14536 | -0.052 | -0.1156 | No | ||
| 121 | AFF3 | 14572 | -0.053 | -0.1133 | No | ||
| 122 | CD302 | 14800 | -0.062 | -0.1184 | No | ||
| 123 | CD79A | 14923 | -0.068 | -0.1194 | No | ||
| 124 | CKAP2 | 14956 | -0.070 | -0.1170 | No | ||
| 125 | TMEM100 | 15021 | -0.074 | -0.1158 | No | ||
| 126 | KCNE2 | 15054 | -0.076 | -0.1134 | No | ||
| 127 | MT1M | 15243 | -0.087 | -0.1169 | No | ||
| 128 | NAP1L2 | 15347 | -0.094 | -0.1172 | No | ||
| 129 | ADAM28 | 15574 | -0.107 | -0.1223 | No | ||
| 130 | ST3GAL6 | 15813 | -0.121 | -0.1278 | No | ||
| 131 | MAL | 16028 | -0.136 | -0.1323 | No | ||
| 132 | ALDH1A2 | 16212 | -0.148 | -0.1357 | No | ||
| 133 | HPGD | 16300 | -0.154 | -0.1354 | No | ||
| 134 | PLLP | 16518 | -0.172 | -0.1401 | No | ||
| 135 | CPA2 | 16742 | -0.193 | -0.1450 | No | ||
| 136 | ARMCX1 | 16771 | -0.196 | -0.1424 | No | ||
| 137 | SPINK2 | 17353 | -0.257 | -0.1611 | No | ||
| 138 | RBP2 | 17512 | -0.276 | -0.1635 | No | ||
| 139 | GPR155 | 17536 | -0.279 | -0.1607 | No | ||
| 140 | SOSTDC1 | 17672 | -0.294 | -0.1623 | No | ||
| 141 | LEPR | 17902 | -0.323 | -0.1674 | No | ||
| 142 | MYOT | 18137 | -0.353 | -0.1727 | No | ||
| 143 | SOX6 | 18167 | -0.358 | -0.1702 | No | ||
| 144 | AGTR1 | 18281 | -0.377 | -0.1709 | No | ||
| 145 | FUT9 | 18366 | -0.393 | -0.1705 | No | ||
| 146 | GC | 18676 | -0.449 | -0.1787 | No | ||
| 147 | BMPR1B | 18895 | -0.493 | -0.1834 | No | ||
| 148 | HLF | 19420 | -0.612 | -0.1999 | No | ||
| 149 | PPOX | 19743 | -0.697 | -0.2086 | Yes | ||
| 150 | REG1A | 19744 | -0.698 | -0.2049 | Yes | ||
| 151 | GABRB3 | 19774 | -0.707 | -0.2024 | Yes | ||
| 152 | ANGPTL1 | 19818 | -0.725 | -0.2004 | Yes | ||
| 153 | GCNT2 | 19933 | -0.760 | -0.2011 | Yes | ||
| 154 | EBF1 | 19965 | -0.771 | -0.1987 | Yes | ||
| 155 | DAB1 | 19977 | -0.775 | -0.1954 | Yes | ||
| 156 | CCPG1 | 19990 | -0.782 | -0.1923 | Yes | ||
| 157 | SNRPN | 20021 | -0.789 | -0.1898 | Yes | ||
| 158 | SCUBE2 | 20091 | -0.806 | -0.1888 | Yes | ||
| 159 | CFL2 | 20208 | -0.852 | -0.1896 | Yes | ||
| 160 | ASPA | 20250 | -0.866 | -0.1875 | Yes | ||
| 161 | GIP | 20279 | -0.878 | -0.1849 | Yes | ||
| 162 | PLCXD3 | 20413 | -0.928 | -0.1864 | Yes | ||
| 163 | GREM2 | 20479 | -0.953 | -0.1852 | Yes | ||
| 164 | KCNMB2 | 20530 | -0.973 | -0.1835 | Yes | ||
| 165 | PTGER3 | 20721 | -1.056 | -0.1871 | Yes | ||
| 166 | SORBS2 | 20831 | -1.108 | -0.1877 | Yes | ||
| 167 | SYNPO2 | 20858 | -1.121 | -0.1850 | Yes | ||
| 168 | MAGI2 | 20938 | -1.159 | -0.1844 | Yes | ||
| 169 | FIGN | 21064 | -1.230 | -0.1855 | Yes | ||
| 170 | LAMA2 | 21082 | -1.242 | -0.1825 | Yes | ||
| 171 | PELI2 | 21101 | -1.249 | -0.1796 | Yes | ||
| 172 | RAB27A | 21149 | -1.274 | -0.1777 | Yes | ||
| 173 | KIT | 21165 | -1.287 | -0.1747 | Yes | ||
| 174 | GNAO1 | 21169 | -1.289 | -0.1711 | Yes | ||
| 175 | WIPF3 | 21305 | -1.365 | -0.1727 | Yes | ||
| 176 | ANGPT1 | 21314 | -1.369 | -0.1693 | Yes | ||
| 177 | KANK4 | 21320 | -1.372 | -0.1659 | Yes | ||
| 178 | ZNF385B | 21363 | -1.400 | -0.1638 | Yes | ||
| 179 | JAM2 | 21427 | -1.439 | -0.1626 | Yes | ||
| 180 | TMEM27 | 21440 | -1.446 | -0.1594 | Yes | ||
| 181 | C2orf40 | 21533 | -1.505 | -0.1593 | Yes | ||
| 182 | ALDH6A1 | 21603 | -1.547 | -0.1583 | Yes | ||
| 183 | THSD4 | 21616 | -1.554 | -0.1551 | Yes | ||
| 184 | SLC18A2 | 21624 | -1.560 | -0.1517 | Yes | ||
| 185 | SNX24 | 21648 | -1.576 | -0.1490 | Yes | ||
| 186 | CYTIP | 21723 | -1.636 | -0.1482 | Yes | ||
| 187 | PTPRZ1 | 21730 | -1.642 | -0.1447 | Yes | ||
| 188 | GAS1 | 21755 | -1.655 | -0.1420 | Yes | ||
| 189 | ASAH2B | 21766 | -1.659 | -0.1387 | Yes | ||
| 190 | PRKD3 | 21969 | -1.838 | -0.1428 | Yes | ||
| 191 | ESRRG | 22069 | -1.918 | -0.1430 | Yes | ||
| 192 | RBMS3 | 22109 | -1.951 | -0.1408 | Yes | ||
| 193 | CGNL1 | 22208 | -2.038 | -0.1410 | Yes | ||
| 194 | PRDM16 | 22321 | -2.152 | -0.1416 | Yes | ||
| 195 | UBE2J1 | 22476 | -2.310 | -0.1439 | Yes | ||
| 196 | GHR | 22496 | -2.324 | -0.1410 | Yes | ||
| 197 | RAB9B | 22500 | -2.328 | -0.1374 | Yes | ||
| 198 | FAM3B | 22579 | -2.419 | -0.1368 | Yes | ||
| 199 | GPR27 | 22740 | -2.608 | -0.1393 | Yes | ||
| 200 | RCAN2 | 22804 | -2.692 | -0.1380 | Yes | ||
| 201 | SCN7A | 22862 | -2.788 | -0.1366 | Yes | ||
| 202 | KLF15 | 22876 | -2.802 | -0.1334 | Yes | ||
| 203 | C7 | 22906 | -2.827 | -0.1309 | Yes | ||
| 204 | AKR1C2 | 22988 | -2.925 | -0.1303 | Yes | ||
| 205 | MZB1 | 23000 | -2.943 | -0.1271 | Yes | ||
| 206 | ADA | 23091 | -3.070 | -0.1269 | Yes | ||
| 207 | RASSF10 | 23144 | -3.145 | -0.1253 | Yes | ||
| 208 | RGMB | 23260 | -3.343 | -0.1260 | Yes | ||
| 209 | MT1H | 23299 | -3.419 | -0.1238 | Yes | ||
| 210 | NLRX1 | 23326 | -3.465 | -0.1212 | Yes | ||
| 211 | SLC15A1 | 23389 | -3.602 | -0.1199 | Yes | ||
| 212 | SETBP1 | 23585 | -3.977 | -0.1238 | Yes | ||
| 213 | SLC1A2 | 23754 | -4.357 | -0.1266 | Yes | ||
| 214 | SH3GL2 | 23763 | -4.389 | -0.1232 | Yes | ||
| 215 | PDGFD | 23772 | -4.405 | -0.1199 | Yes | ||
| 216 | ALDH1A1 | 23818 | -4.510 | -0.1180 | Yes | ||
| 217 | RPS6KA6 | 23855 | -4.592 | -0.1157 | Yes | ||
| 218 | CHRDL1 | 23901 | -4.737 | -0.1138 | Yes | ||
| 219 | SYT4 | 24069 | -5.238 | -0.1165 | Yes | ||
| 220 | IL6R | 24075 | -5.248 | -0.1131 | Yes | ||
| 221 | ITIH5 | 24085 | -5.267 | -0.1098 | Yes | ||
| 222 | DTNB | 24149 | -5.434 | -0.1085 | Yes | ||
| 223 | PKIB | 24187 | -5.562 | -0.1063 | Yes | ||
| 224 | CPE | 24236 | -5.705 | -0.1045 | Yes | ||
| 225 | KLF9 | 24326 | -6.110 | -0.1043 | Yes | ||
| 226 | CYP1B1 | 24480 | -6.654 | -0.1065 | Yes | ||
| 227 | RORA | 24535 | -6.926 | -0.1049 | Yes | ||
| 228 | ECHDC3 | 24539 | -6.952 | -0.1014 | Yes | ||
| 229 | LDB3 | 24563 | -7.067 | -0.0986 | Yes | ||
| 230 | FAM20A | 24577 | -7.134 | -0.0955 | Yes | ||
| 231 | C16orf89 | 24712 | -7.718 | -0.0970 | Yes | ||
| 232 | CLDN10 | 24736 | -7.854 | -0.0942 | Yes | ||
| 233 | ITM2A | 24748 | -7.957 | -0.0910 | Yes | ||
| 234 | HMGN5 | 24926 | -9.077 | -0.0941 | Yes | ||
| 235 | CHRM3 | 25024 | -9.924 | -0.0942 | Yes | ||
| 236 | FAM107A | 25030 | -10.032 | -0.0907 | Yes | ||
| 237 | CD36 | 25052 | -10.178 | -0.0879 | Yes | ||
| 238 | DMD | 25205 | -11.618 | -0.0901 | Yes | ||
| 239 | DUSP5 | 25225 | -11.830 | -0.0872 | Yes | ||
| 240 | NDNF | 25228 | -11.869 | -0.0836 | Yes | ||
| 241 | GPM6B | 25244 | -11.974 | -0.0805 | Yes | ||
| 242 | MUM1L1 | 25261 | -12.127 | -0.0775 | Yes | ||
| 243 | CNTN1 | 25362 | -13.471 | -0.0777 | Yes | ||
| 244 | UBE2QL1 | 25363 | -13.491 | -0.0740 | Yes | ||
| 245 | CXCL12 | 25372 | -13.621 | -0.0707 | Yes | ||
| 246 | SUSD4 | 25416 | -14.162 | -0.0687 | Yes | ||
| 247 | COL4A6 | 25422 | -14.248 | -0.0652 | Yes | ||
| 248 | SERPINI1 | 25428 | -14.339 | -0.0618 | Yes | ||
| 249 | PID1 | 25462 | -14.835 | -0.0594 | Yes | ||
| 250 | STOX2 | 25537 | -16.055 | -0.0586 | Yes | ||
| 251 | ZNF844 | 25616 | -18.049 | -0.0579 | Yes | ||
| 252 | LARP6 | 25627 | -18.258 | -0.0546 | Yes | ||
| 253 | MAP7D2 | 25707 | -20.168 | -0.0540 | Yes | ||
| 254 | MT1E | 25733 | -20.906 | -0.0513 | Yes | ||
| 255 | F13A1 | 25739 | -21.210 | -0.0479 | Yes | ||
| 256 | MAOA | 25755 | -21.638 | -0.0448 | Yes | ||
| 257 | ELL2 | 25886 | -26.509 | -0.0461 | Yes | ||
| 258 | NTRK3 | 25906 | -27.988 | -0.0432 | Yes | ||
| 259 | SHISA3 | 25921 | -28.773 | -0.0401 | Yes | ||
| 260 | NR3C1 | 25926 | -29.242 | -0.0366 | Yes | ||
| 261 | NCAM1 | 25964 | -31.402 | -0.0344 | Yes | ||
| 262 | SLIT2 | 25966 | -31.492 | -0.0308 | Yes | ||
| 263 | BCHE | 26012 | -34.895 | -0.0289 | Yes | ||
| 264 | HSPB8 | 26026 | -36.409 | -0.0257 | Yes | ||
| 265 | ATP1A2 | 26137 | -49.083 | -0.0263 | Yes | ||
| 266 | RIMS3 | 26155 | -52.334 | -0.0233 | Yes | ||
| 267 | VLDLR | 26190 | -59.843 | -0.0209 | Yes | ||
| 268 | ARRDC4 | 26217 | -70.088 | -0.0183 | Yes | ||
| 269 | MDFIC | 26218 | -70.415 | -0.0146 | Yes | ||
| 270 | MT1F | 26271 | -107.226 | -0.0130 | Yes | ||
| 271 | ALDH1L2 | 26287 | -152.864 | -0.0099 | Yes | ||
| 272 | CHGA | 26292 | -180.006 | -0.0064 | Yes | ||
| 273 | MT1X | 26307 | -290.514 | -0.0033 | Yes | ||
| 274 | TSC22D3 | 26317 | -299.207 | -0.0000 | Yes |