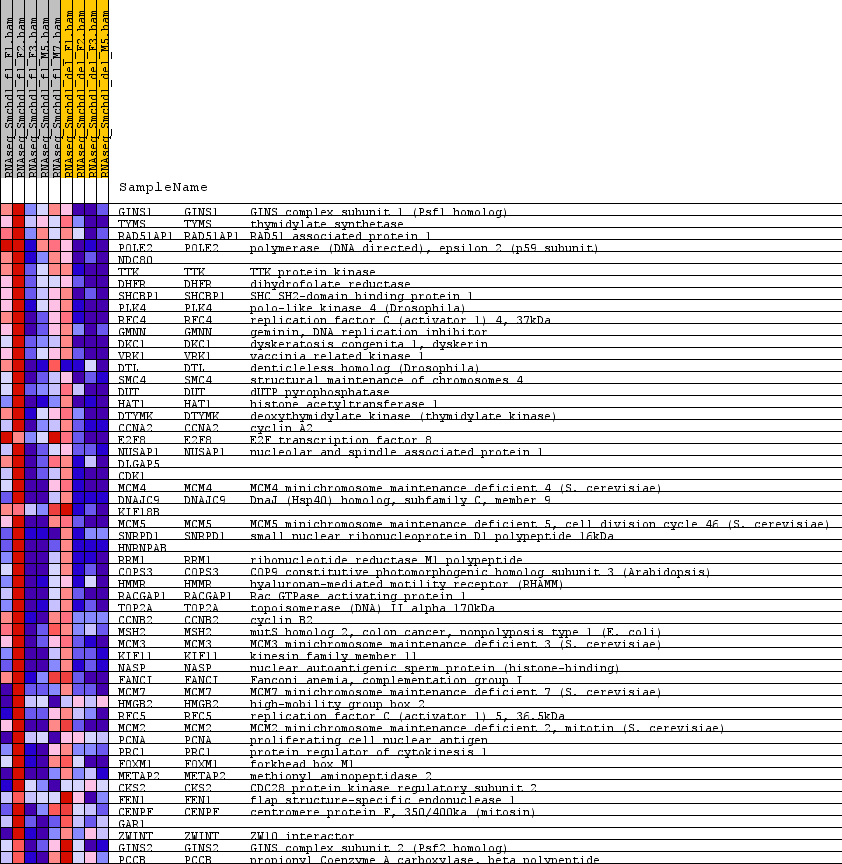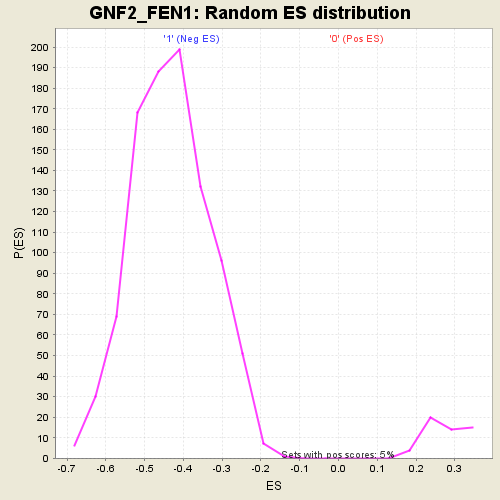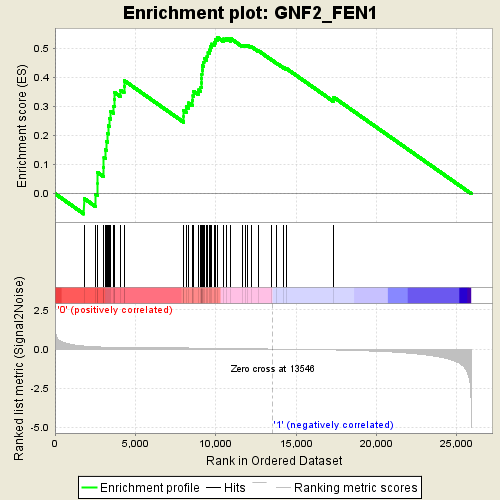
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 0 |
| GeneSet | GNF2_FEN1 |
| Enrichment Score (ES) | 0.5364209 |
| Normalized Enrichment Score (NES) | 1.9170101 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.21794935 |
| FWER p-Value | 0.706 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GINS1 | GINS1 Entrez, Source | GINS complex subunit 1 (Psf1 homolog) | 1800 | 0.195 | -0.0161 | Yes |
| 2 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 2505 | 0.150 | -0.0020 | Yes |
| 3 | RAD51AP1 | RAD51AP1 Entrez, Source | RAD51 associated protein 1 | 2626 | 0.146 | 0.0336 | Yes |
| 4 | POLE2 | POLE2 Entrez, Source | polymerase (DNA directed), epsilon 2 (p59 subunit) | 2630 | 0.146 | 0.0736 | Yes |
| 5 | NDC80 | 3031 | 0.120 | 0.0911 | Yes | ||
| 6 | TTK | TTK Entrez, Source | TTK protein kinase | 3042 | 0.119 | 0.1234 | Yes |
| 7 | DHFR | DHFR Entrez, Source | dihydrofolate reductase | 3124 | 0.114 | 0.1517 | Yes |
| 8 | SHCBP1 | SHCBP1 Entrez, Source | SHC SH2-domain binding protein 1 | 3202 | 0.110 | 0.1790 | Yes |
| 9 | PLK4 | PLK4 Entrez, Source | polo-like kinase 4 (Drosophila) | 3251 | 0.108 | 0.2068 | Yes |
| 10 | RFC4 | RFC4 Entrez, Source | replication factor C (activator 1) 4, 37kDa | 3299 | 0.105 | 0.2339 | Yes |
| 11 | GMNN | GMNN Entrez, Source | geminin, DNA replication inhibitor | 3370 | 0.102 | 0.2592 | Yes |
| 12 | DKC1 | DKC1 Entrez, Source | dyskeratosis congenita 1, dyskerin | 3456 | 0.098 | 0.2830 | Yes |
| 13 | VRK1 | VRK1 Entrez, Source | vaccinia related kinase 1 | 3639 | 0.091 | 0.3011 | Yes |
| 14 | DTL | DTL Entrez, Source | denticleless homolog (Drosophila) | 3696 | 0.089 | 0.3234 | Yes |
| 15 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 3717 | 0.088 | 0.3469 | Yes |
| 16 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 4087 | 0.082 | 0.3553 | Yes |
| 17 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 4293 | 0.075 | 0.3681 | Yes |
| 18 | DTYMK | DTYMK Entrez, Source | deoxythymidylate kinase (thymidylate kinase) | 4302 | 0.075 | 0.3886 | Yes |
| 19 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 7999 | 0.073 | 0.2657 | Yes |
| 20 | E2F8 | E2F8 Entrez, Source | E2F transcription factor 8 | 8004 | 0.073 | 0.2856 | Yes |
| 21 | NUSAP1 | NUSAP1 Entrez, Source | nucleolar and spindle associated protein 1 | 8179 | 0.071 | 0.2983 | Yes |
| 22 | DLGAP5 | 8319 | 0.068 | 0.3116 | Yes | ||
| 23 | CDK1 | 8548 | 0.063 | 0.3201 | Yes | ||
| 24 | MCM4 | MCM4 Entrez, Source | MCM4 minichromosome maintenance deficient 4 (S. cerevisiae) | 8582 | 0.062 | 0.3359 | Yes |
| 25 | DNAJC9 | DNAJC9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 9 | 8623 | 0.061 | 0.3512 | Yes |
| 26 | KIF18B | 8919 | 0.060 | 0.3562 | Yes | ||
| 27 | MCM5 | MCM5 Entrez, Source | MCM5 minichromosome maintenance deficient 5, cell division cycle 46 (S. cerevisiae) | 9075 | 0.057 | 0.3659 | Yes |
| 28 | SNRPD1 | SNRPD1 Entrez, Source | small nuclear ribonucleoprotein D1 polypeptide 16kDa | 9105 | 0.056 | 0.3803 | Yes |
| 29 | HNRNPAB | 9112 | 0.056 | 0.3955 | Yes | ||
| 30 | RRM1 | RRM1 Entrez, Source | ribonucleotide reductase M1 polypeptide | 9137 | 0.056 | 0.4099 | Yes |
| 31 | COPS3 | COPS3 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 3 (Arabidopsis) | 9146 | 0.056 | 0.4248 | Yes |
| 32 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 9173 | 0.055 | 0.4390 | Yes |
| 33 | RACGAP1 | RACGAP1 Entrez, Source | Rac GTPase activating protein 1 | 9253 | 0.055 | 0.4509 | Yes |
| 34 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9270 | 0.054 | 0.4652 | Yes |
| 35 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 9453 | 0.051 | 0.4723 | Yes |
| 36 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 9516 | 0.050 | 0.4837 | Yes |
| 37 | MCM3 | MCM3 Entrez, Source | MCM3 minichromosome maintenance deficient 3 (S. cerevisiae) | 9611 | 0.049 | 0.4934 | Yes |
| 38 | KIF11 | KIF11 Entrez, Source | kinesin family member 11 | 9652 | 0.048 | 0.5052 | Yes |
| 39 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 9757 | 0.047 | 0.5140 | Yes |
| 40 | FANCI | FANCI Entrez, Source | Fanconi anemia, complementation group I | 9924 | 0.045 | 0.5199 | Yes |
| 41 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 9993 | 0.044 | 0.5292 | Yes |
| 42 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 10107 | 0.042 | 0.5364 | Yes |
| 43 | RFC5 | RFC5 Entrez, Source | replication factor C (activator 1) 5, 36.5kDa | 10501 | 0.037 | 0.5313 | No |
| 44 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 10680 | 0.034 | 0.5338 | No |
| 45 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 10939 | 0.031 | 0.5324 | No |
| 46 | PRC1 | PRC1 Entrez, Source | protein regulator of cytokinesis 1 | 11666 | 0.023 | 0.5105 | No |
| 47 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 11822 | 0.021 | 0.5103 | No |
| 48 | METAP2 | METAP2 Entrez, Source | methionyl aminopeptidase 2 | 11990 | 0.019 | 0.5090 | No |
| 49 | CKS2 | CKS2 Entrez, Source | CDC28 protein kinase regulatory subunit 2 | 12217 | 0.016 | 0.5047 | No |
| 50 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 12646 | 0.011 | 0.4911 | No |
| 51 | CENPF | CENPF Entrez, Source | centromere protein F, 350/400ka (mitosin) | 13466 | 0.001 | 0.4597 | No |
| 52 | GAR1 | 13801 | -0.003 | 0.4476 | No | ||
| 53 | ZWINT | ZWINT Entrez, Source | ZW10 interactor | 14235 | -0.008 | 0.4331 | No |
| 54 | GINS2 | GINS2 Entrez, Source | GINS complex subunit 2 (Psf2 homolog) | 14397 | -0.011 | 0.4298 | No |
| 55 | PCCB | PCCB Entrez, Source | propionyl Coenzyme A carboxylase, beta polypeptide | 17337 | -0.055 | 0.3313 | No |