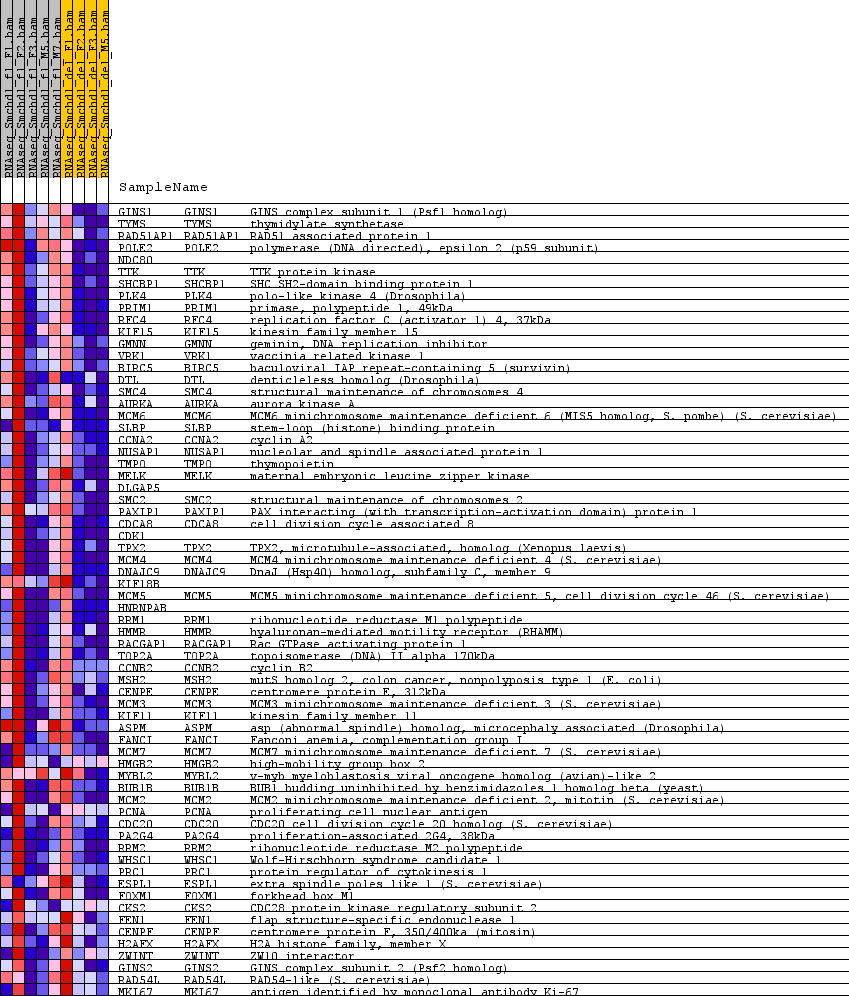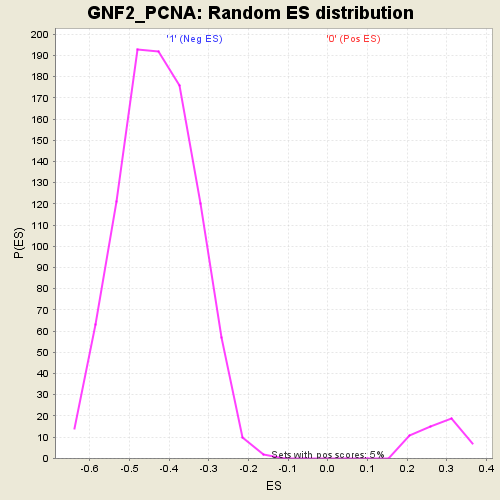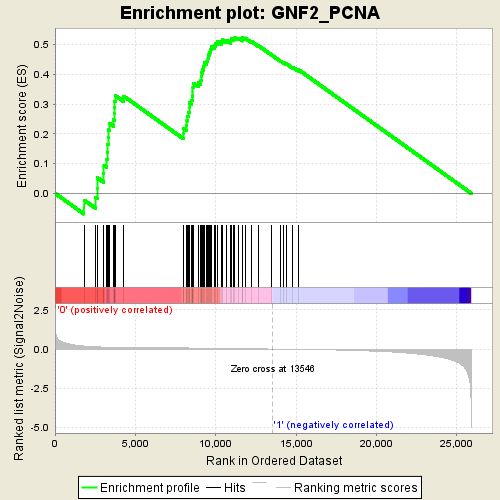
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 0 |
| GeneSet | GNF2_PCNA |
| Enrichment Score (ES) | 0.52445 |
| Normalized Enrichment Score (NES) | 1.8589573 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.2283703 |
| FWER p-Value | 0.881 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GINS1 | GINS1 Entrez, Source | GINS complex subunit 1 (Psf1 homolog) | 1800 | 0.195 | -0.0226 | Yes |
| 2 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 2505 | 0.150 | -0.0135 | Yes |
| 3 | RAD51AP1 | RAD51AP1 Entrez, Source | RAD51 associated protein 1 | 2626 | 0.146 | 0.0172 | Yes |
| 4 | POLE2 | POLE2 Entrez, Source | polymerase (DNA directed), epsilon 2 (p59 subunit) | 2630 | 0.146 | 0.0524 | Yes |
| 5 | NDC80 | 3031 | 0.120 | 0.0659 | Yes | ||
| 6 | TTK | TTK Entrez, Source | TTK protein kinase | 3042 | 0.119 | 0.0943 | Yes |
| 7 | SHCBP1 | SHCBP1 Entrez, Source | SHC SH2-domain binding protein 1 | 3202 | 0.110 | 0.1148 | Yes |
| 8 | PLK4 | PLK4 Entrez, Source | polo-like kinase 4 (Drosophila) | 3251 | 0.108 | 0.1390 | Yes |
| 9 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 3268 | 0.107 | 0.1642 | Yes |
| 10 | RFC4 | RFC4 Entrez, Source | replication factor C (activator 1) 4, 37kDa | 3299 | 0.105 | 0.1884 | Yes |
| 11 | KIF15 | KIF15 Entrez, Source | kinesin family member 15 | 3309 | 0.105 | 0.2134 | Yes |
| 12 | GMNN | GMNN Entrez, Source | geminin, DNA replication inhibitor | 3370 | 0.102 | 0.2357 | Yes |
| 13 | VRK1 | VRK1 Entrez, Source | vaccinia related kinase 1 | 3639 | 0.091 | 0.2474 | Yes |
| 14 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 3691 | 0.090 | 0.2671 | Yes |
| 15 | DTL | DTL Entrez, Source | denticleless homolog (Drosophila) | 3696 | 0.089 | 0.2885 | Yes |
| 16 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 3717 | 0.088 | 0.3091 | Yes |
| 17 | AURKA | AURKA Entrez, Source | aurora kinase A | 3750 | 0.087 | 0.3290 | Yes |
| 18 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 4276 | 0.076 | 0.3270 | Yes |
| 19 | SLBP | SLBP Entrez, Source | stem-loop (histone) binding protein | 7995 | 0.073 | 0.2008 | Yes |
| 20 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 7999 | 0.073 | 0.2184 | Yes |
| 21 | NUSAP1 | NUSAP1 Entrez, Source | nucleolar and spindle associated protein 1 | 8179 | 0.071 | 0.2285 | Yes |
| 22 | TMPO | TMPO Entrez, Source | thymopoietin | 8207 | 0.070 | 0.2443 | Yes |
| 23 | MELK | MELK Entrez, Source | maternal embryonic leucine zipper kinase | 8236 | 0.069 | 0.2600 | Yes |
| 24 | DLGAP5 | 8319 | 0.068 | 0.2733 | Yes | ||
| 25 | SMC2 | SMC2 Entrez, Source | structural maintenance of chromosomes 2 | 8352 | 0.067 | 0.2883 | Yes |
| 26 | PAXIP1 | PAXIP1 Entrez, Source | PAX interacting (with transcription-activation domain) protein 1 | 8356 | 0.067 | 0.3043 | Yes |
| 27 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 8511 | 0.064 | 0.3138 | Yes |
| 28 | CDK1 | 8548 | 0.063 | 0.3277 | Yes | ||
| 29 | TPX2 | TPX2 Entrez, Source | TPX2, microtubule-associated, homolog (Xenopus laevis) | 8581 | 0.062 | 0.3415 | Yes |
| 30 | MCM4 | MCM4 Entrez, Source | MCM4 minichromosome maintenance deficient 4 (S. cerevisiae) | 8582 | 0.062 | 0.3565 | Yes |
| 31 | DNAJC9 | DNAJC9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 9 | 8623 | 0.061 | 0.3698 | Yes |
| 32 | KIF18B | 8919 | 0.060 | 0.3728 | Yes | ||
| 33 | MCM5 | MCM5 Entrez, Source | MCM5 minichromosome maintenance deficient 5, cell division cycle 46 (S. cerevisiae) | 9075 | 0.057 | 0.3806 | Yes |
| 34 | HNRNPAB | 9112 | 0.056 | 0.3927 | Yes | ||
| 35 | RRM1 | RRM1 Entrez, Source | ribonucleotide reductase M1 polypeptide | 9137 | 0.056 | 0.4053 | Yes |
| 36 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 9173 | 0.055 | 0.4173 | Yes |
| 37 | RACGAP1 | RACGAP1 Entrez, Source | Rac GTPase activating protein 1 | 9253 | 0.055 | 0.4274 | Yes |
| 38 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9270 | 0.054 | 0.4399 | Yes |
| 39 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 9453 | 0.051 | 0.4452 | Yes |
| 40 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 9516 | 0.050 | 0.4549 | Yes |
| 41 | CENPE | CENPE Entrez, Source | centromere protein E, 312kDa | 9548 | 0.050 | 0.4658 | Yes |
| 42 | MCM3 | MCM3 Entrez, Source | MCM3 minichromosome maintenance deficient 3 (S. cerevisiae) | 9611 | 0.049 | 0.4752 | Yes |
| 43 | KIF11 | KIF11 Entrez, Source | kinesin family member 11 | 9652 | 0.048 | 0.4853 | Yes |
| 44 | ASPM | ASPM Entrez, Source | asp (abnormal spindle) homolog, microcephaly associated (Drosophila) | 9711 | 0.047 | 0.4945 | Yes |
| 45 | FANCI | FANCI Entrez, Source | Fanconi anemia, complementation group I | 9924 | 0.045 | 0.4971 | Yes |
| 46 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 9993 | 0.044 | 0.5050 | Yes |
| 47 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 10107 | 0.042 | 0.5108 | Yes |
| 48 | MYBL2 | MYBL2 Entrez, Source | v-myb myeloblastosis viral oncogene homolog (avian)-like 2 | 10368 | 0.039 | 0.5101 | Yes |
| 49 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 10437 | 0.037 | 0.5165 | Yes |
| 50 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 10680 | 0.034 | 0.5154 | Yes |
| 51 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 10939 | 0.031 | 0.5129 | Yes |
| 52 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 10950 | 0.031 | 0.5200 | Yes |
| 53 | PA2G4 | PA2G4 Entrez, Source | proliferation-associated 2G4, 38kDa | 11100 | 0.029 | 0.5213 | Yes |
| 54 | RRM2 | RRM2 Entrez, Source | ribonucleotide reductase M2 polypeptide | 11196 | 0.028 | 0.5245 | Yes |
| 55 | WHSC1 | WHSC1 Entrez, Source | Wolf-Hirschhorn syndrome candidate 1 | 11405 | 0.026 | 0.5226 | No |
| 56 | PRC1 | PRC1 Entrez, Source | protein regulator of cytokinesis 1 | 11666 | 0.023 | 0.5181 | No |
| 57 | ESPL1 | ESPL1 Entrez, Source | extra spindle poles like 1 (S. cerevisiae) | 11672 | 0.023 | 0.5234 | No |
| 58 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 11822 | 0.021 | 0.5227 | No |
| 59 | CKS2 | CKS2 Entrez, Source | CDC28 protein kinase regulatory subunit 2 | 12217 | 0.016 | 0.5113 | No |
| 60 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 12646 | 0.011 | 0.4973 | No |
| 61 | CENPF | CENPF Entrez, Source | centromere protein F, 350/400ka (mitosin) | 13466 | 0.001 | 0.4659 | No |
| 62 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 14049 | -0.006 | 0.4447 | No |
| 63 | ZWINT | ZWINT Entrez, Source | ZW10 interactor | 14235 | -0.008 | 0.4396 | No |
| 64 | GINS2 | GINS2 Entrez, Source | GINS complex subunit 2 (Psf2 homolog) | 14397 | -0.011 | 0.4359 | No |
| 65 | RAD54L | RAD54L Entrez, Source | RAD54-like (S. cerevisiae) | 14784 | -0.016 | 0.4249 | No |
| 66 | MKI67 | MKI67 Entrez, Source | antigen identified by monoclonal antibody Ki-67 | 15149 | -0.022 | 0.4161 | No |