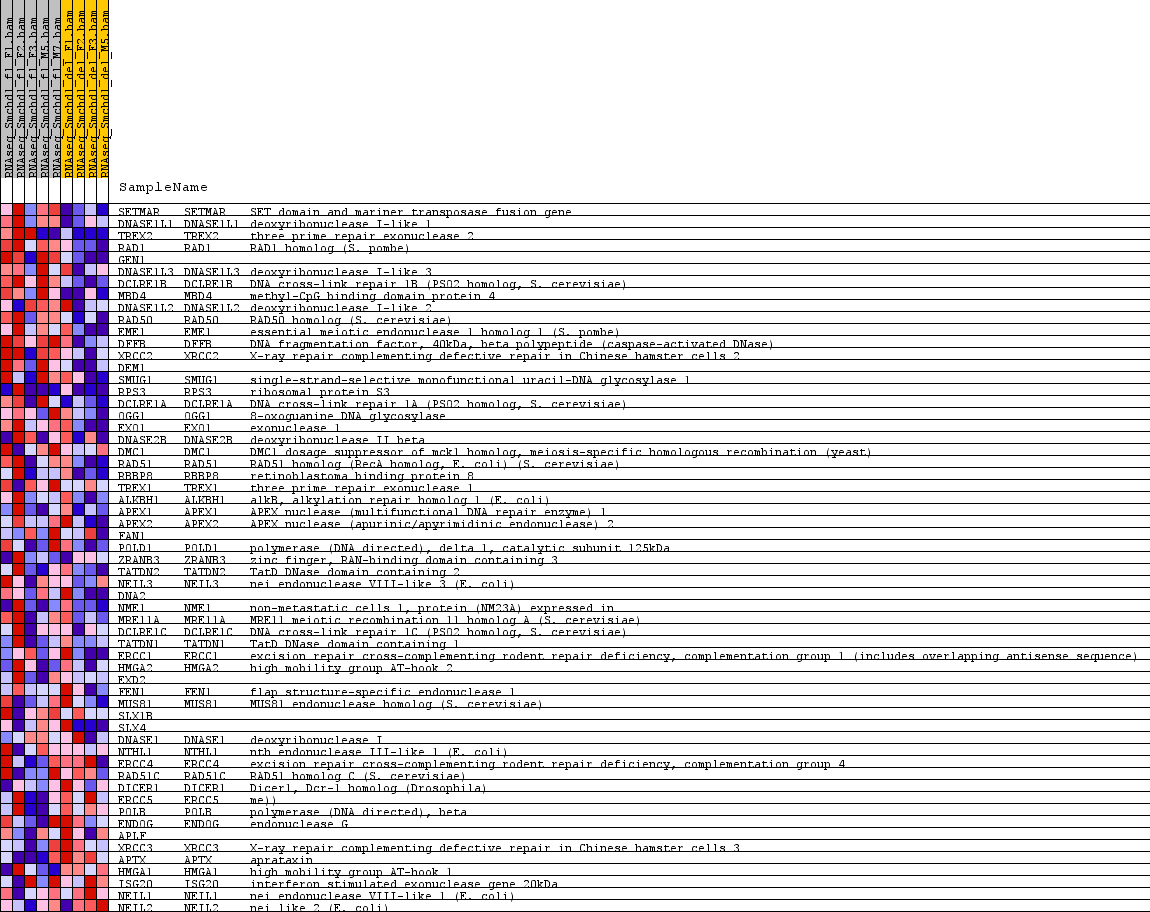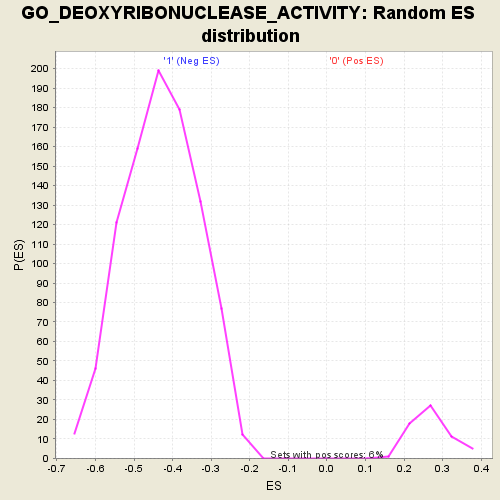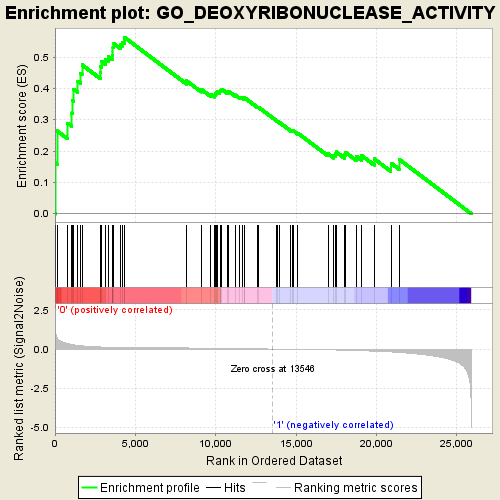
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 0 |
| GeneSet | GO_DEOXYRIBONUCLEASE_ACTIVITY |
| Enrichment Score (ES) | 0.5626565 |
| Normalized Enrichment Score (NES) | 2.0547664 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14603247 |
| FWER p-Value | 0.203 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SETMAR | SETMAR Entrez, Source | SET domain and mariner transposase fusion gene | 28 | 1.113 | 0.1621 | Yes |
| 2 | DNASE1L1 | DNASE1L1 Entrez, Source | deoxyribonuclease I-like 1 | 125 | 0.724 | 0.2644 | Yes |
| 3 | TREX2 | TREX2 Entrez, Source | three prime repair exonuclease 2 | 782 | 0.341 | 0.2891 | Yes |
| 4 | RAD1 | RAD1 Entrez, Source | RAD1 homolog (S. pombe) | 1045 | 0.290 | 0.3215 | Yes |
| 5 | GEN1 | 1095 | 0.280 | 0.3606 | Yes | ||
| 6 | DNASE1L3 | DNASE1L3 Entrez, Source | deoxyribonuclease I-like 3 | 1158 | 0.271 | 0.3980 | Yes |
| 7 | DCLRE1B | DCLRE1B Entrez, Source | DNA cross-link repair 1B (PSO2 homolog, S. cerevisiae) | 1421 | 0.239 | 0.4229 | Yes |
| 8 | MBD4 | MBD4 Entrez, Source | methyl-CpG binding domain protein 4 | 1590 | 0.217 | 0.4482 | Yes |
| 9 | DNASE1L2 | DNASE1L2 Entrez, Source | deoxyribonuclease I-like 2 | 1680 | 0.207 | 0.4751 | Yes |
| 10 | RAD50 | RAD50 Entrez, Source | RAD50 homolog (S. cerevisiae) | 2833 | 0.133 | 0.4500 | Yes |
| 11 | EME1 | EME1 Entrez, Source | essential meiotic endonuclease 1 homolog 1 (S. pombe) | 2841 | 0.132 | 0.4691 | Yes |
| 12 | DFFB | DFFB Entrez, Source | DNA fragmentation factor, 40kDa, beta polypeptide (caspase-activated DNase) | 2890 | 0.129 | 0.4862 | Yes |
| 13 | XRCC2 | XRCC2 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 2 | 3138 | 0.114 | 0.4933 | Yes |
| 14 | DEM1 | 3340 | 0.103 | 0.5007 | Yes | ||
| 15 | SMUG1 | SMUG1 Entrez, Source | single-strand-selective monofunctional uracil-DNA glycosylase 1 | 3552 | 0.095 | 0.5064 | Yes |
| 16 | RPS3 | RPS3 Entrez, Source | ribosomal protein S3 | 3602 | 0.093 | 0.5181 | Yes |
| 17 | DCLRE1A | DCLRE1A Entrez, Source | DNA cross-link repair 1A (PSO2 homolog, S. cerevisiae) | 3603 | 0.093 | 0.5317 | Yes |
| 18 | OGG1 | OGG1 Entrez, Source | 8-oxoguanine DNA glycosylase | 3661 | 0.091 | 0.5428 | Yes |
| 19 | EXO1 | EXO1 Entrez, Source | exonuclease 1 | 4079 | 0.083 | 0.5387 | Yes |
| 20 | DNASE2B | DNASE2B Entrez, Source | deoxyribonuclease II beta | 4171 | 0.079 | 0.5468 | Yes |
| 21 | DMC1 | DMC1 Entrez, Source | DMC1 dosage suppressor of mck1 homolog, meiosis-specific homologous recombination (yeast) | 4301 | 0.075 | 0.5529 | Yes |
| 22 | RAD51 | RAD51 Entrez, Source | RAD51 homolog (RecA homolog, E. coli) (S. cerevisiae) | 4332 | 0.074 | 0.5627 | Yes |
| 23 | RBBP8 | RBBP8 Entrez, Source | retinoblastoma binding protein 8 | 8178 | 0.071 | 0.4242 | No |
| 24 | TREX1 | TREX1 Entrez, Source | three prime repair exonuclease 1 | 9102 | 0.056 | 0.3968 | No |
| 25 | ALKBH1 | ALKBH1 Entrez, Source | alkB, alkylation repair homolog 1 (E. coli) | 9690 | 0.048 | 0.3810 | No |
| 26 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 9914 | 0.045 | 0.3790 | No |
| 27 | APEX2 | APEX2 Entrez, Source | APEX nuclease (apurinic/apyrimidinic endonuclease) 2 | 9990 | 0.044 | 0.3825 | No |
| 28 | FAN1 | 10019 | 0.043 | 0.3877 | No | ||
| 29 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10086 | 0.042 | 0.3914 | No |
| 30 | ZRANB3 | ZRANB3 Entrez, Source | zinc finger, RAN-binding domain containing 3 | 10264 | 0.040 | 0.3904 | No |
| 31 | TATDN2 | TATDN2 Entrez, Source | TatD DNase domain containing 2 | 10307 | 0.039 | 0.3945 | No |
| 32 | NEIL3 | NEIL3 Entrez, Source | nei endonuclease VIII-like 3 (E. coli) | 10382 | 0.038 | 0.3973 | No |
| 33 | DNA2 | 10747 | 0.033 | 0.3881 | No | ||
| 34 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 10797 | 0.033 | 0.3910 | No |
| 35 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 11214 | 0.028 | 0.3790 | No |
| 36 | DCLRE1C | DCLRE1C Entrez, Source | DNA cross-link repair 1C (PSO2 homolog, S. cerevisiae) | 11483 | 0.025 | 0.3723 | No |
| 37 | TATDN1 | TATDN1 Entrez, Source | TatD DNase domain containing 1 | 11639 | 0.023 | 0.3697 | No |
| 38 | ERCC1 | ERCC1 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 1 (includes overlapping antisense sequence) | 11774 | 0.022 | 0.3677 | No |
| 39 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 11790 | 0.021 | 0.3702 | No |
| 40 | EXD2 | 12616 | 0.011 | 0.3399 | No | ||
| 41 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 12646 | 0.011 | 0.3404 | No |
| 42 | MUS81 | MUS81 Entrez, Source | MUS81 endonuclease homolog (S. cerevisiae) | 13797 | -0.003 | 0.2964 | No |
| 43 | SLX1B | 13841 | -0.003 | 0.2952 | No | ||
| 44 | SLX4 | 13997 | -0.005 | 0.2900 | No | ||
| 45 | DNASE1 | DNASE1 Entrez, Source | deoxyribonuclease I | 14678 | -0.015 | 0.2658 | No |
| 46 | NTHL1 | NTHL1 Entrez, Source | nth endonuclease III-like 1 (E. coli) | 14759 | -0.016 | 0.2650 | No |
| 47 | ERCC4 | ERCC4 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 4 | 14817 | -0.017 | 0.2653 | No |
| 48 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 15118 | -0.021 | 0.2568 | No |
| 49 | DICER1 | DICER1 Entrez, Source | Dicer1, Dcr-1 homolog (Drosophila) | 16988 | -0.048 | 0.1916 | No |
| 50 | ERCC5 | ERCC5 Entrez, Source | me)) | 17348 | -0.055 | 0.1858 | No |
| 51 | POLB | POLB Entrez, Source | polymerase (DNA directed), beta | 17433 | -0.057 | 0.1909 | No |
| 52 | ENDOG | ENDOG Entrez, Source | endonuclease G | 17494 | -0.059 | 0.1972 | No |
| 53 | APLF | 18013 | -0.071 | 0.1876 | No | ||
| 54 | XRCC3 | XRCC3 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 3 | 18052 | -0.072 | 0.1967 | No |
| 55 | APTX | APTX Entrez, Source | aprataxin | 18735 | -0.092 | 0.1837 | No |
| 56 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 19069 | -0.103 | 0.1859 | No |
| 57 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 19862 | -0.134 | 0.1750 | No |
| 58 | NEIL1 | NEIL1 Entrez, Source | nei endonuclease VIII-like 1 (E. coli) | 20907 | -0.183 | 0.1614 | No |
| 59 | NEIL2 | NEIL2 Entrez, Source | nei like 2 (E. coli) | 21412 | -0.216 | 0.1736 | No |