Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 1 |
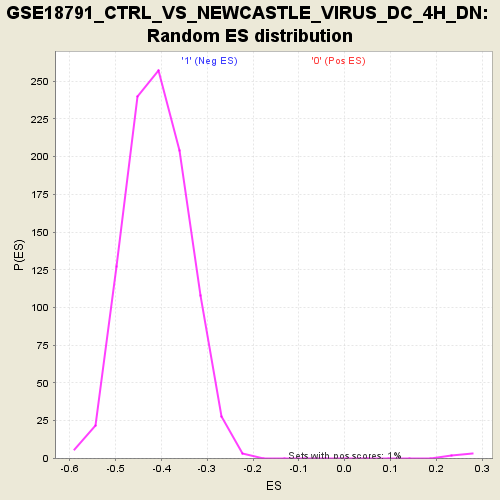
| GeneSet | GSE18791_CTRL_VS_NEWCASTLE_VIRUS_DC_4H_DN |
| Enrichment Score (ES) | -0.7391456 |
| Normalized Enrichment Score (NES) | -1.8040872 |
| Nominal p-value | 0.0 |
| FDR q-value | 9.1727683E-4 |
| FWER p-Value | 0.04 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HIST1H1T | HIST1H1T Entrez, Source | histone cluster 1, H1t | 75 | 0.869 | 0.0105 | No |
| 2 | SCN11A | SCN11A Entrez, Source | sodium channel, voltage-gated, type XI, alpha | 275 | 0.547 | 0.0111 | No |
| 3 | TRIM21 | TRIM21 Entrez, Source | tripartite motif-containing 21 | 618 | 0.389 | 0.0038 | No |
| 4 | IFITM1 | IFITM1 Entrez, Source | interferon induced transmembrane protein 1 (9-27) | 995 | 0.299 | -0.0062 | No |
| 5 | GFOD1 | GFOD1 Entrez, Source | glucose-fructose oxidoreductase domain containing 1 | 1027 | 0.294 | -0.0028 | No |
| 6 | TTC9B | TTC9B Entrez, Source | tetratricopeptide repeat domain 9B | 1136 | 0.274 | -0.0028 | No |
| 7 | GRASP | GRASP Entrez, Source | GRP1 (general receptor for phosphoinositides 1)-associated scaffold protein | 1559 | 0.222 | -0.0158 | No |
| 8 | LRRC48 | LRRC48 Entrez, Source | leucine rich repeat containing 48 | 1737 | 0.200 | -0.0196 | No |
| 9 | PRX | PRX Entrez, Source | periaxin | 1789 | 0.196 | -0.0186 | No |
| 10 | SLC1A6 | SLC1A6 Entrez, Source | solute carrier family 1 (high affinity aspartate/glutamate transporter), member 6 | 2398 | 0.160 | -0.0397 | No |
| 11 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 2436 | 0.157 | -0.0387 | No |
| 12 | PGAP1 | PGAP1 Entrez, Source | - | 2785 | 0.135 | -0.0502 | No |
| 13 | MYADM | MYADM Entrez, Source | myeloid-associated differentiation marker | 2849 | 0.132 | -0.0506 | No |
| 14 | MIS18BP1 | 2886 | 0.129 | -0.0500 | No | ||
| 15 | NDC80 | 3031 | 0.120 | -0.0537 | No | ||
| 16 | SPC24 | 3195 | 0.111 | -0.0584 | No | ||
| 17 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 3204 | 0.110 | -0.0570 | No |
| 18 | STAT2 | STAT2 Entrez, Source | signal transducer and activator of transcription 2, 113kDa | 3441 | 0.099 | -0.0646 | No |
| 19 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 3743 | 0.087 | -0.0750 | No |
| 20 | ZBP1 | ZBP1 Entrez, Source | Z-DNA binding protein 1 | 4016 | 0.085 | -0.0842 | No |
| 21 | PROC | PROC Entrez, Source | protein C (inactivator of coagulation factors Va and VIIIa) | 4095 | 0.082 | -0.0860 | No |
| 22 | MASTL | MASTL Entrez, Source | microtubule associated serine/threonine kinase-like | 4210 | 0.078 | -0.0892 | No |
| 23 | HOMER1 | HOMER1 Entrez, Source | homer homolog 1 (Drosophila) | 4328 | 0.075 | -0.0926 | No |
| 24 | BPIFA2 | 4833 | 0.074 | -0.1111 | No | ||
| 25 | HESX1 | HESX1 Entrez, Source | homeobox, ES cell expressed 1 | 5122 | 0.074 | -0.1211 | No |
| 26 | CCL8 | CCL8 Entrez, Source | chemokine (C-C motif) ligand 8 | 5906 | 0.074 | -0.1504 | No |
| 27 | NT5C3 | NT5C3 Entrez, Source | 5'-nucleotidase, cytosolic III | 7985 | 0.074 | -0.2300 | No |
| 28 | TRIM31 | TRIM31 Entrez, Source | tripartite motif-containing 31 | 8378 | 0.067 | -0.2442 | No |
| 29 | B3GAT2 | B3GAT2 Entrez, Source | beta-1,3-glucuronyltransferase 2 (glucuronosyltransferase S) | 8399 | 0.066 | -0.2439 | No |
| 30 | C1GALT1 | C1GALT1 Entrez, Source | core 1 synthase, glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase, 1 | 8900 | 0.060 | -0.2624 | No |
| 31 | ADM | ADM Entrez, Source | adrenomedullin | 8976 | 0.058 | -0.2644 | No |
| 32 | TMEM212 | 8984 | 0.058 | -0.2638 | No | ||
| 33 | PELI1 | PELI1 Entrez, Source | pellino homolog 1 (Drosophila) | 9096 | 0.057 | -0.2672 | No |
| 34 | OAS3 | OAS3 Entrez, Source | 2'-5'-oligoadenylate synthetase 3, 100kDa | 9242 | 0.055 | -0.2720 | No |
| 35 | PPM1K | PPM1K Entrez, Source | protein phosphatase 1K (PP2C domain containing) | 9477 | 0.051 | -0.2803 | No |
| 36 | UTY | UTY Entrez, Source | ubiquitously transcribed tetratricopeptide repeat gene, Y-linked | 9546 | 0.050 | -0.2822 | No |
| 37 | PELO | PELO Entrez, Source | pelota homolog (Drosophila) | 9763 | 0.047 | -0.2899 | No |
| 38 | CD80 | CD80 Entrez, Source | CD80 molecule | 9947 | 0.044 | -0.2963 | No |
| 39 | TLK2 | TLK2 Entrez, Source | tousled-like kinase 2 | 9957 | 0.044 | -0.2960 | No |
| 40 | PLA1A | PLA1A Entrez, Source | phospholipase A1 member A | 10085 | 0.042 | -0.3003 | No |
| 41 | NAMPT | 10235 | 0.040 | -0.3054 | No | ||
| 42 | TNFSF10 | TNFSF10 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 10 | 10244 | 0.040 | -0.3051 | No |
| 43 | RLF | RLF Entrez, Source | rearranged L-myc fusion | 10543 | 0.036 | -0.3161 | No |
| 44 | PAPOLG | PAPOLG Entrez, Source | poly(A) polymerase gamma | 10722 | 0.034 | -0.3225 | No |
| 45 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 10904 | 0.031 | -0.3291 | No |
| 46 | USP42 | USP42 Entrez, Source | ubiquitin specific peptidase 42 | 11009 | 0.030 | -0.3327 | No |
| 47 | WDR33 | WDR33 Entrez, Source | WD repeat domain 33 | 11600 | 0.024 | -0.3552 | No |
| 48 | PAPD7 | 11863 | 0.021 | -0.3651 | No | ||
| 49 | HSPA4L | HSPA4L Entrez, Source | heat shock 70kDa protein 4-like | 11927 | 0.020 | -0.3672 | No |
| 50 | TRIM56 | TRIM56 Entrez, Source | tripartite motif-containing 56 | 11967 | 0.019 | -0.3684 | No |
| 51 | DNAJC27 | 12108 | 0.018 | -0.3736 | No | ||
| 52 | PATL1 | 12138 | 0.017 | -0.3745 | No | ||
| 53 | PNPT1 | PNPT1 Entrez, Source | polyribonucleotide nucleotidyltransferase 1 | 12329 | 0.015 | -0.3816 | No |
| 54 | FAM46A | FAM46A Entrez, Source | family with sequence similarity 46, member A | 12429 | 0.013 | -0.3853 | No |
| 55 | MYEF2 | MYEF2 Entrez, Source | myelin expression factor 2 | 12514 | 0.012 | -0.3883 | No |
| 56 | DCP1A | DCP1A Entrez, Source | DCP1 decapping enzyme homolog A (S. cerevisiae) | 12620 | 0.011 | -0.3923 | No |
| 57 | CCDC57 | CCDC57 Entrez, Source | coiled-coil domain containing 57 | 12972 | 0.007 | -0.4058 | No |
| 58 | FNBP4 | FNBP4 Entrez, Source | formin binding protein 4 | 13043 | 0.006 | -0.4084 | No |
| 59 | ARL5B | ARL5B Entrez, Source | ADP-ribosylation factor-like 5B | 13127 | 0.005 | -0.4115 | No |
| 60 | FAM125A | FAM125A Entrez, Source | family with sequence similarity 125, member A | 13884 | -0.004 | -0.4409 | No |
| 61 | CLK1 | CLK1 Entrez, Source | CDC-like kinase 1 | 14245 | -0.008 | -0.4547 | No |
| 62 | CLK4 | CLK4 Entrez, Source | CDC-like kinase 4 | 14268 | -0.009 | -0.4554 | No |
| 63 | SNAPC1 | SNAPC1 Entrez, Source | small nuclear RNA activating complex, polypeptide 1, 43kDa | 14407 | -0.011 | -0.4606 | No |
| 64 | DOPEY1 | DOPEY1 Entrez, Source | dopey family member 1 | 14562 | -0.013 | -0.4664 | No |
| 65 | ZBTB43 | ZBTB43 Entrez, Source | zinc finger and BTB domain containing 43 | 14704 | -0.015 | -0.4716 | No |
| 66 | ADAR | ADAR Entrez, Source | adenosine deaminase, RNA-specific | 14749 | -0.016 | -0.4731 | No |
| 67 | GTPBP2 | GTPBP2 Entrez, Source | GTP binding protein 2 | 14768 | -0.016 | -0.4736 | No |
| 68 | SORBS3 | SORBS3 Entrez, Source | sorbin and SH3 domain containing 3 | 15088 | -0.021 | -0.4856 | No |
| 69 | SRC | SRC Entrez, Source | v-src sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog (avian) | 15130 | -0.021 | -0.4869 | No |
| 70 | PHF11 | PHF11 Entrez, Source | PHD finger protein 11 | 15262 | -0.024 | -0.4916 | No |
| 71 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 15487 | -0.026 | -0.4999 | No |
| 72 | PARP6 | PARP6 Entrez, Source | poly (ADP-ribose) polymerase family, member 6 | 15537 | -0.027 | -0.5014 | No |
| 73 | THAP3 | THAP3 Entrez, Source | THAP domain containing, apoptosis associated protein 3 | 15763 | -0.030 | -0.5097 | No |
| 74 | MYO10 | MYO10 Entrez, Source | myosin X | 16053 | -0.035 | -0.5204 | No |
| 75 | BATF2 | BATF2 Entrez, Source | basic leucine zipper transcription factor, ATF-like 2 | 16492 | -0.039 | -0.5368 | No |
| 76 | KDM6B | 16501 | -0.039 | -0.5365 | No | ||
| 77 | NFKB2 | NFKB2 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) | 16578 | -0.040 | -0.5388 | No |
| 78 | PATL2 | 16798 | -0.045 | -0.5467 | No | ||
| 79 | CSRNP1 | 16989 | -0.048 | -0.5533 | No | ||
| 80 | ANKS1B | ANKS1B Entrez, Source | ankyrin repeat and sterile alpha motif domain containing 1B | 17081 | -0.050 | -0.5561 | No |
| 81 | DAND5 | DAND5 Entrez, Source | DAN domain family, member 5 | 17138 | -0.050 | -0.5575 | No |
| 82 | FBXO34 | FBXO34 Entrez, Source | F-box protein 34 | 17206 | -0.052 | -0.5593 | No |
| 83 | DNAJB5 | DNAJB5 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 5 | 17795 | -0.065 | -0.5811 | No |
| 84 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 17931 | -0.069 | -0.5853 | No |
| 85 | DENND2C | DENND2C Entrez, Source | DENN/MADD domain containing 2C | 18075 | -0.073 | -0.5897 | No |
| 86 | NR2E1 | NR2E1 Entrez, Source | nuclear receptor subfamily 2, group E, member 1 | 18204 | -0.076 | -0.5935 | No |
| 87 | PMAIP1 | PMAIP1 Entrez, Source | phorbol-12-myristate-13-acetate-induced protein 1 | 18228 | -0.077 | -0.5932 | No |
| 88 | USPL1 | USPL1 Entrez, Source | ubiquitin specific peptidase like 1 | 18252 | -0.077 | -0.5929 | No |
| 89 | RICTOR | RICTOR Entrez, Source | - | 18292 | -0.078 | -0.5932 | No |
| 90 | MFN1 | MFN1 Entrez, Source | mitofusin 1 | 18329 | -0.080 | -0.5934 | No |
| 91 | PPP3CC | PPP3CC Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, gamma isoform (calcineurin A gamma) | 18409 | -0.082 | -0.5952 | No |
| 92 | SETD4 | SETD4 Entrez, Source | SET domain containing 4 | 18463 | -0.084 | -0.5960 | No |
| 93 | TGIF1 | 18493 | -0.085 | -0.5958 | No | ||
| 94 | SLC25A28 | SLC25A28 Entrez, Source | solute carrier family 25, member 28 | 19038 | -0.102 | -0.6154 | No |
| 95 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 19108 | -0.104 | -0.6164 | No |
| 96 | ARID3A | ARID3A Entrez, Source | AT rich interactive domain 3A (BRIGHT- like) | 19150 | -0.106 | -0.6164 | No |
| 97 | TMEM217 | 19213 | -0.108 | -0.6171 | No | ||
| 98 | B4GALT5 | B4GALT5 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 5 | 19418 | -0.116 | -0.6233 | No |
| 99 | TNNT1 | TNNT1 Entrez, Source | troponin T type 1 (skeletal, slow) | 19419 | -0.117 | -0.6215 | No |
| 100 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 19862 | -0.134 | -0.6366 | No |
| 101 | ZNFX1 | ZNFX1 Entrez, Source | zinc finger, NFX1-type containing 1 | 19866 | -0.134 | -0.6346 | No |
| 102 | SLC31A2 | SLC31A2 Entrez, Source | solute carrier family 31 (copper transporters), member 2 | 20026 | -0.140 | -0.6387 | No |
| 103 | PML | PML Entrez, Source | promyelocytic leukemia | 20690 | -0.171 | -0.6618 | No |
| 104 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 21171 | -0.200 | -0.6773 | No |
| 105 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 21174 | -0.200 | -0.6744 | No |
| 106 | FAM69B | FAM69B Entrez, Source | family with sequence similarity 69, member B | 21638 | -0.234 | -0.6887 | No |
| 107 | HERC6 | HERC6 Entrez, Source | hect domain and RLD 6 | 21820 | -0.248 | -0.6919 | No |
| 108 | ERRFI1 | ERRFI1 Entrez, Source | ERBB receptor feedback inhibitor 1 | 22047 | -0.267 | -0.6966 | No |
| 109 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 22778 | -0.338 | -0.7198 | No |
| 110 | CMPK2 | 23278 | -0.403 | -0.7330 | Yes | ||
| 111 | SPATA18 | SPATA18 Entrez, Source | spermatogenesis associated 18 homolog (rat) | 23357 | -0.414 | -0.7296 | Yes |
| 112 | ARHGAP8 | ARHGAP8 Entrez, Source | Rho GTPase activating protein 8 | 23427 | -0.424 | -0.7258 | Yes |
| 113 | FUT4 | FUT4 Entrez, Source | fucosyltransferase 4 (alpha (1,3) fucosyltransferase, myeloid-specific) | 23500 | -0.434 | -0.7219 | Yes |
| 114 | EPSTI1 | EPSTI1 Entrez, Source | epithelial stromal interaction 1 (breast) | 23538 | -0.439 | -0.7166 | Yes |
| 115 | MAK | MAK Entrez, Source | male germ cell-associated kinase | 23634 | -0.455 | -0.7133 | Yes |
| 116 | RNF213 | 23919 | -0.506 | -0.7165 | Yes | ||
| 117 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 23939 | -0.510 | -0.7094 | Yes |
| 118 | UNC93B1 | UNC93B1 Entrez, Source | unc-93 homolog B1 (C. elegans) | 23979 | -0.517 | -0.7029 | Yes |
| 119 | DDX58 | DDX58 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 58 | 24122 | -0.545 | -0.7001 | Yes |
| 120 | PARP9 | PARP9 Entrez, Source | poly (ADP-ribose) polymerase family, member 9 | 24171 | -0.559 | -0.6933 | Yes |
| 121 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 24212 | -0.568 | -0.6862 | Yes |
| 122 | ARHGAP27 | ARHGAP27 Entrez, Source | Rho GTPase activating protein 27 | 24315 | -0.591 | -0.6810 | Yes |
| 123 | TRAF1 | TRAF1 Entrez, Source | TNF receptor-associated factor 1 | 24399 | -0.616 | -0.6748 | Yes |
| 124 | RNF43 | RNF43 Entrez, Source | ring finger protein 43 | 24425 | -0.625 | -0.6661 | Yes |
| 125 | DHX58 | 24511 | -0.647 | -0.6595 | Yes | ||
| 126 | DTX3L | DTX3L Entrez, Source | deltex 3-like (Drosophila) | 24573 | -0.670 | -0.6516 | Yes |
| 127 | IFIT2 | IFIT2 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 2 | 24672 | -0.695 | -0.6447 | Yes |
| 128 | REC8 | 24743 | -0.715 | -0.6364 | Yes | ||
| 129 | DUSP15 | DUSP15 Entrez, Source | dual specificity phosphatase 15 | 24847 | -0.752 | -0.6288 | Yes |
| 130 | TRIM25 | TRIM25 Entrez, Source | tripartite motif-containing 25 | 25023 | -0.834 | -0.6228 | Yes |
| 131 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 25230 | -0.948 | -0.6162 | Yes |
| 132 | RAB11FIP1 | RAB11FIP1 Entrez, Source | RAB11 family interacting protein 1 (class I) | 25255 | -0.970 | -0.6022 | Yes |
| 133 | CD274 | CD274 Entrez, Source | CD274 molecule | 25335 | -1.036 | -0.5893 | Yes |
| 134 | SYNPR | SYNPR Entrez, Source | synaptoporin | 25371 | -1.062 | -0.5744 | Yes |
| 135 | RASAL1 | RASAL1 Entrez, Source | RAS protein activator like 1 (GAP1 like) | 25390 | -1.079 | -0.5584 | Yes |
| 136 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 25437 | -1.129 | -0.5429 | Yes |
| 137 | OAS2 | OAS2 Entrez, Source | 2'-5'-oligoadenylate synthetase 2, 69/71kDa | 25558 | -1.284 | -0.5278 | Yes |
| 138 | IFI35 | IFI35 Entrez, Source | interferon-induced protein 35 | 25621 | -1.385 | -0.5089 | Yes |
| 139 | PARP14 | PARP14 Entrez, Source | poly (ADP-ribose) polymerase family, member 14 | 25624 | -1.394 | -0.4875 | Yes |
| 140 | ISG15 | ISG15 Entrez, Source | ISG15 ubiquitin-like modifier | 25699 | -1.594 | -0.4658 | Yes |
| 141 | DDX60 | 25710 | -1.643 | -0.4410 | Yes | ||
| 142 | IRAK2 | IRAK2 Entrez, Source | interleukin-1 receptor-associated kinase 2 | 25730 | -1.689 | -0.4157 | Yes |
| 143 | IRF7 | IRF7 Entrez, Source | interferon regulatory factor 7 | 25794 | -2.009 | -0.3873 | Yes |
| 144 | RSAD2 | RSAD2 Entrez, Source | radical S-adenosyl methionine domain containing 2 | 25795 | -2.013 | -0.3563 | Yes |
| 145 | IFIH1 | IFIH1 Entrez, Source | interferon induced with helicase C domain 1 | 25804 | -2.052 | -0.3250 | Yes |
| 146 | XAF1 | 25822 | -2.156 | -0.2925 | Yes | ||
| 147 | USP18 | USP18 Entrez, Source | ubiquitin specific peptidase 18 | 25830 | -2.231 | -0.2584 | Yes |
| 148 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 25831 | -2.278 | -0.2234 | Yes |
| 149 | RTP4 | RTP4 Entrez, Source | receptor transporter protein 4 | 25833 | -2.300 | -0.1880 | Yes |
| 150 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 25844 | -2.414 | -0.1513 | Yes |
| 151 | NLRC5 | 25867 | -2.711 | -0.1104 | Yes | ||
| 152 | SAMD9L | SAMD9L Entrez, Source | sterile alpha motif domain containing 9-like | 25888 | -3.178 | -0.0623 | Yes |
| 153 | IFIT1 | IFIT1 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 1 | 25894 | -4.073 | 0.0002 | Yes |