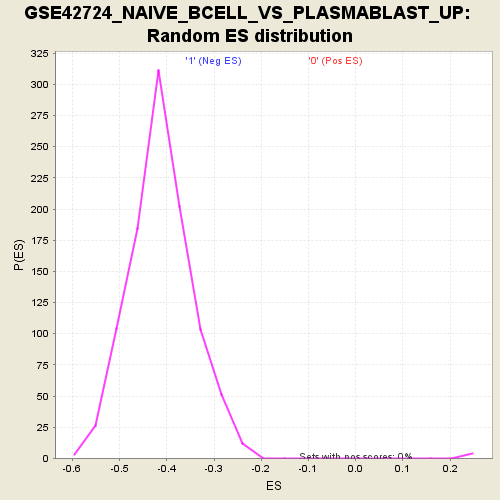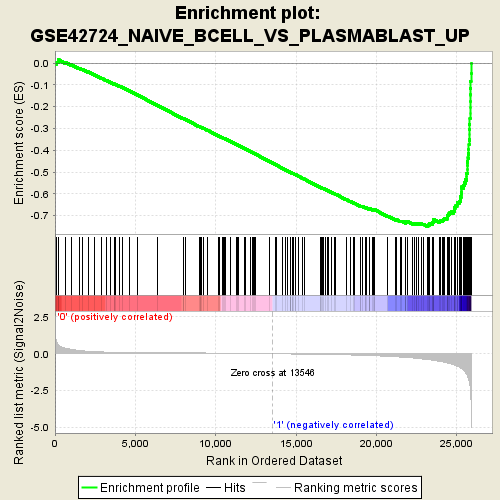
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 1 |
| GeneSet | GSE42724_NAIVE_BCELL_VS_PLASMABLAST_UP |
| Enrichment Score (ES) | -0.7489225 |
| Normalized Enrichment Score (NES) | -1.8175259 |
| Nominal p-value | 0.0 |
| FDR q-value | 6.322941E-4 |
| FWER p-Value | 0.025 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SIGLEC1 | SIGLEC1 Entrez, Source | sialic acid binding Ig-like lectin 1, sialoadhesin | 114 | 0.750 | 0.0046 | No |
| 2 | LAG3 | LAG3 Entrez, Source | lymphocyte-activation gene 3 | 197 | 0.620 | 0.0088 | No |
| 3 | GBP5 | GBP5 Entrez, Source | guanylate binding protein 5 | 202 | 0.614 | 0.0160 | No |
| 4 | TRIM21 | TRIM21 Entrez, Source | tripartite motif-containing 21 | 618 | 0.389 | 0.0045 | No |
| 5 | IFITM1 | IFITM1 Entrez, Source | interferon induced transmembrane protein 1 (9-27) | 995 | 0.299 | -0.0065 | No |
| 6 | OTOF | OTOF Entrez, Source | otoferlin | 1512 | 0.230 | -0.0238 | No |
| 7 | MVP | MVP Entrez, Source | major vault protein | 1693 | 0.205 | -0.0283 | No |
| 8 | BRIP1 | BRIP1 Entrez, Source | BRCA1 interacting protein C-terminal helicase 1 | 2054 | 0.171 | -0.0402 | No |
| 9 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 2436 | 0.157 | -0.0532 | No |
| 10 | CFB | CFB Entrez, Source | complement factor B | 2894 | 0.128 | -0.0694 | No |
| 11 | STARD4 | STARD4 Entrez, Source | START domain containing 4, sterol regulated | 3169 | 0.112 | -0.0787 | No |
| 12 | STAT2 | STAT2 Entrez, Source | signal transducer and activator of transcription 2, 113kDa | 3441 | 0.099 | -0.0880 | No |
| 13 | SPATS2L | 3682 | 0.090 | -0.0963 | No | ||
| 14 | HSH2D | HSH2D Entrez, Source | hematopoietic SH2 domain containing | 3695 | 0.089 | -0.0957 | No |
| 15 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 3743 | 0.087 | -0.0965 | No |
| 16 | ZBP1 | ZBP1 Entrez, Source | Z-DNA binding protein 1 | 4016 | 0.085 | -0.1060 | No |
| 17 | MASTL | MASTL Entrez, Source | microtubule associated serine/threonine kinase-like | 4210 | 0.078 | -0.1126 | No |
| 18 | TIMD4 | TIMD4 Entrez, Source | T-cell immunoglobulin and mucin domain containing 4 | 4614 | 0.074 | -0.1273 | No |
| 19 | HESX1 | HESX1 Entrez, Source | homeobox, ES cell expressed 1 | 5122 | 0.074 | -0.1462 | No |
| 20 | PRR9 | PRR9 Entrez, Source | proline rich 9 | 6355 | 0.074 | -0.1931 | No |
| 21 | NT5C3 | NT5C3 Entrez, Source | 5'-nucleotidase, cytosolic III | 7985 | 0.074 | -0.2556 | No |
| 22 | E2F8 | E2F8 Entrez, Source | E2F transcription factor 8 | 8004 | 0.073 | -0.2554 | No |
| 23 | CLEC4D | CLEC4D Entrez, Source | C-type lectin domain family 4, member D | 8126 | 0.072 | -0.2592 | No |
| 24 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 8961 | 0.059 | -0.2909 | No |
| 25 | MCM5 | MCM5 Entrez, Source | MCM5 minichromosome maintenance deficient 5, cell division cycle 46 (S. cerevisiae) | 9075 | 0.057 | -0.2946 | No |
| 26 | SP110 | SP110 Entrez, Source | SP110 nuclear body protein | 9080 | 0.057 | -0.2941 | No |
| 27 | TREX1 | TREX1 Entrez, Source | three prime repair exonuclease 1 | 9102 | 0.056 | -0.2942 | No |
| 28 | OAS3 | OAS3 Entrez, Source | 2'-5'-oligoadenylate synthetase 3, 100kDa | 9242 | 0.055 | -0.2990 | No |
| 29 | PPM1K | PPM1K Entrez, Source | protein phosphatase 1K (PP2C domain containing) | 9477 | 0.051 | -0.3075 | No |
| 30 | TDRD7 | TDRD7 Entrez, Source | tudor domain containing 7 | 10188 | 0.041 | -0.3346 | No |
| 31 | TNFSF10 | TNFSF10 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 10 | 10244 | 0.040 | -0.3362 | No |
| 32 | MRPL40 | MRPL40 Entrez, Source | mitochondrial ribosomal protein L40 | 10410 | 0.038 | -0.3422 | No |
| 33 | SSB | SSB Entrez, Source | Sjogren syndrome antigen B (autoantigen La) | 10503 | 0.037 | -0.3453 | No |
| 34 | UBAP1 | UBAP1 Entrez, Source | ubiquitin associated protein 1 | 10552 | 0.036 | -0.3468 | No |
| 35 | GBP1 | GBP1 Entrez, Source | guanylate binding protein 1, interferon-inducible, 67kDa | 10631 | 0.035 | -0.3494 | No |
| 36 | HUS1 | HUS1 Entrez, Source | HUS1 checkpoint homolog (S. pombe) | 10921 | 0.031 | -0.3602 | No |
| 37 | RAB8A | RAB8A Entrez, Source | RAB8A, member RAS oncogene family | 11317 | 0.027 | -0.3753 | No |
| 38 | CNP | CNP Entrez, Source | 2',3'-cyclic nucleotide 3' phosphodiesterase | 11335 | 0.027 | -0.3756 | No |
| 39 | LAP3 | LAP3 Entrez, Source | leucine aminopeptidase 3 | 11434 | 0.025 | -0.3791 | No |
| 40 | ST7 | ST7 Entrez, Source | suppression of tumorigenicity 7 | 11768 | 0.022 | -0.3918 | No |
| 41 | SRGAP2 | SRGAP2 Entrez, Source | SLIT-ROBO Rho GTPase activating protein 2 | 11880 | 0.020 | -0.3959 | No |
| 42 | PATL1 | 12138 | 0.017 | -0.4056 | No | ||
| 43 | NUB1 | NUB1 Entrez, Source | negative regulator of ubiquitin-like proteins 1 | 12297 | 0.015 | -0.4116 | No |
| 44 | PNPT1 | PNPT1 Entrez, Source | polyribonucleotide nucleotidyltransferase 1 | 12329 | 0.015 | -0.4126 | No |
| 45 | FAM46A | FAM46A Entrez, Source | family with sequence similarity 46, member A | 12429 | 0.013 | -0.4163 | No |
| 46 | RIPK1 | RIPK1 Entrez, Source | receptor (TNFRSF)-interacting serine-threonine kinase 1 | 12485 | 0.013 | -0.4183 | No |
| 47 | MAD2L1BP | MAD2L1BP Entrez, Source | MAD2L1 binding protein | 13326 | 0.003 | -0.4509 | No |
| 48 | PHACTR4 | PHACTR4 Entrez, Source | phosphatase and actin regulator 4 | 13746 | -0.003 | -0.4672 | No |
| 49 | COQ10B | COQ10B Entrez, Source | coenzyme Q10 homolog B (S. cerevisiae) | 13783 | -0.003 | -0.4685 | No |
| 50 | NEXN | NEXN Entrez, Source | nexilin (F actin binding protein) | 14171 | -0.007 | -0.4835 | No |
| 51 | FBXO6 | FBXO6 Entrez, Source | F-box protein 6 | 14359 | -0.010 | -0.4906 | No |
| 52 | NADK | NADK Entrez, Source | NAD kinase | 14442 | -0.011 | -0.4937 | No |
| 53 | TMEM140 | TMEM140 Entrez, Source | transmembrane protein 140 | 14623 | -0.014 | -0.5005 | No |
| 54 | SCARB2 | SCARB2 Entrez, Source | scavenger receptor class B, member 2 | 14644 | -0.014 | -0.5011 | No |
| 55 | KIRREL | KIRREL Entrez, Source | kin of IRRE like (Drosophila) | 14650 | -0.014 | -0.5011 | No |
| 56 | XPO6 | XPO6 Entrez, Source | exportin 6 | 14770 | -0.016 | -0.5056 | No |
| 57 | TYMP | 14791 | -0.016 | -0.5061 | No | ||
| 58 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 14830 | -0.017 | -0.5074 | No |
| 59 | PPPDE2 | 14941 | -0.019 | -0.5115 | No | ||
| 60 | NKIRAS2 | NKIRAS2 Entrez, Source | NFKB inhibitor interacting Ras-like 2 | 14964 | -0.019 | -0.5121 | No |
| 61 | PSMA4 | PSMA4 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 4 | 15175 | -0.022 | -0.5200 | No |
| 62 | EPDR1 | EPDR1 Entrez, Source | ependymin related protein 1 (zebrafish) | 15375 | -0.024 | -0.5274 | No |
| 63 | FRMD3 | FRMD3 Entrez, Source | FERM domain containing 3 | 15496 | -0.026 | -0.5318 | No |
| 64 | BATF2 | BATF2 Entrez, Source | basic leucine zipper transcription factor, ATF-like 2 | 16492 | -0.039 | -0.5700 | No |
| 65 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 16525 | -0.039 | -0.5708 | No |
| 66 | FTSJD2 | 16599 | -0.041 | -0.5731 | No | ||
| 67 | SQSTM1 | SQSTM1 Entrez, Source | sequestosome 1 | 16666 | -0.042 | -0.5752 | No |
| 68 | CD2AP | CD2AP Entrez, Source | CD2-associated protein | 16696 | -0.043 | -0.5758 | No |
| 69 | FBXO32 | FBXO32 Entrez, Source | F-box protein 32 | 16806 | -0.045 | -0.5795 | No |
| 70 | CLEC1A | CLEC1A Entrez, Source | C-type lectin domain family 1, member A | 16965 | -0.048 | -0.5850 | No |
| 71 | CSRNP1 | 16989 | -0.048 | -0.5854 | No | ||
| 72 | PSME1 | PSME1 Entrez, Source | proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) | 17203 | -0.052 | -0.5930 | No |
| 73 | NAPA | NAPA Entrez, Source | N-ethylmaleimide-sensitive factor attachment protein, alpha | 17380 | -0.056 | -0.5992 | No |
| 74 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 17473 | -0.058 | -0.6020 | No |
| 75 | MOV10 | MOV10 Entrez, Source | Mov10, Moloney leukemia virus 10, homolog (mouse) | 18149 | -0.075 | -0.6274 | No |
| 76 | UBA7 | 18403 | -0.082 | -0.6362 | No | ||
| 77 | PCGF5 | PCGF5 Entrez, Source | polycomb group ring finger 5 | 18570 | -0.087 | -0.6416 | No |
| 78 | UBE2L6 | UBE2L6 Entrez, Source | ubiquitin-conjugating enzyme E2L 6 | 18633 | -0.089 | -0.6430 | No |
| 79 | TRAFD1 | TRAFD1 Entrez, Source | TRAF-type zinc finger domain containing 1 | 19019 | -0.101 | -0.6567 | No |
| 80 | SLC25A28 | SLC25A28 Entrez, Source | solute carrier family 25, member 28 | 19038 | -0.102 | -0.6562 | No |
| 81 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 19108 | -0.104 | -0.6576 | No |
| 82 | DDHD1 | DDHD1 Entrez, Source | DDHD domain containing 1 | 19301 | -0.111 | -0.6638 | No |
| 83 | SHISA5 | 19322 | -0.112 | -0.6632 | No | ||
| 84 | SP140 | SP140 Entrez, Source | SP140 nuclear body protein | 19352 | -0.114 | -0.6630 | No |
| 85 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 19540 | -0.121 | -0.6688 | No |
| 86 | NRIP1 | NRIP1 Entrez, Source | nuclear receptor interacting protein 1 | 19549 | -0.121 | -0.6676 | No |
| 87 | STOM | STOM Entrez, Source | stomatin | 19559 | -0.122 | -0.6665 | No |
| 88 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 19768 | -0.131 | -0.6731 | No |
| 89 | APOL6 | APOL6 Entrez, Source | apolipoprotein L, 6 | 19838 | -0.133 | -0.6741 | No |
| 90 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 19862 | -0.134 | -0.6734 | No |
| 91 | ZNFX1 | ZNFX1 Entrez, Source | zinc finger, NFX1-type containing 1 | 19866 | -0.134 | -0.6719 | No |
| 92 | PML | PML Entrez, Source | promyelocytic leukemia | 20690 | -0.171 | -0.7018 | No |
| 93 | MYD88 | MYD88 Entrez, Source | myeloid differentiation primary response gene (88) | 21196 | -0.201 | -0.7191 | No |
| 94 | B3GNT2 | B3GNT2 Entrez, Source | UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 2 | 21219 | -0.203 | -0.7175 | No |
| 95 | IL15RA | IL15RA Entrez, Source | interleukin 15 receptor, alpha | 21510 | -0.224 | -0.7261 | No |
| 96 | EPHB2 | EPHB2 Entrez, Source | EPH receptor B2 | 21579 | -0.230 | -0.7260 | No |
| 97 | HERC6 | HERC6 Entrez, Source | hect domain and RLD 6 | 21820 | -0.248 | -0.7323 | No |
| 98 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 21833 | -0.249 | -0.7298 | No |
| 99 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 21835 | -0.250 | -0.7268 | No |
| 100 | EIF2AK2 | EIF2AK2 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 2 | 21911 | -0.256 | -0.7267 | No |
| 101 | FCHSD1 | FCHSD1 Entrez, Source | FCH and double SH3 domains 1 | 22223 | -0.281 | -0.7354 | No |
| 102 | ITIH4 | ITIH4 Entrez, Source | inter-alpha (globulin) inhibitor H4 (plasma Kallikrein-sensitive glycoprotein) | 22352 | -0.293 | -0.7368 | No |
| 103 | IRF9 | 22485 | -0.307 | -0.7383 | No | ||
| 104 | GMPR | GMPR Entrez, Source | guanosine monophosphate reductase | 22516 | -0.310 | -0.7357 | No |
| 105 | LY6E | LY6E Entrez, Source | lymphocyte antigen 6 complex, locus E | 22607 | -0.320 | -0.7354 | No |
| 106 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 22778 | -0.338 | -0.7379 | No |
| 107 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 22902 | -0.355 | -0.7385 | No |
| 108 | CLDN1 | CLDN1 Entrez, Source | claudin 1 | 23172 | -0.387 | -0.7443 | Yes |
| 109 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 23259 | -0.399 | -0.7428 | Yes |
| 110 | CMPK2 | 23278 | -0.403 | -0.7387 | Yes | ||
| 111 | AXL | AXL Entrez, Source | AXL receptor tyrosine kinase | 23294 | -0.405 | -0.7344 | Yes |
| 112 | TRIM5 | TRIM5 Entrez, Source | tripartite motif-containing 5 | 23487 | -0.433 | -0.7367 | Yes |
| 113 | FUT4 | FUT4 Entrez, Source | fucosyltransferase 4 (alpha (1,3) fucosyltransferase, myeloid-specific) | 23500 | -0.434 | -0.7320 | Yes |
| 114 | EPSTI1 | EPSTI1 Entrez, Source | epithelial stromal interaction 1 (breast) | 23538 | -0.439 | -0.7282 | Yes |
| 115 | PARP12 | PARP12 Entrez, Source | poly (ADP-ribose) polymerase family, member 12 | 23548 | -0.441 | -0.7232 | Yes |
| 116 | KCTD14 | KCTD14 Entrez, Source | potassium channel tetramerisation domain containing 14 | 23568 | -0.444 | -0.7186 | Yes |
| 117 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 23939 | -0.510 | -0.7269 | Yes |
| 118 | ACSM5 | 23961 | -0.514 | -0.7215 | Yes | ||
| 119 | DDX58 | DDX58 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 58 | 24122 | -0.545 | -0.7212 | Yes |
| 120 | PARP9 | PARP9 Entrez, Source | poly (ADP-ribose) polymerase family, member 9 | 24171 | -0.559 | -0.7164 | Yes |
| 121 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 24212 | -0.568 | -0.7111 | Yes |
| 122 | TAPBPL | TAPBPL Entrez, Source | TAP binding protein-like | 24395 | -0.615 | -0.7108 | Yes |
| 123 | RTP1 | RTP1 Entrez, Source | receptor transporter protein 1 | 24434 | -0.627 | -0.7048 | Yes |
| 124 | MTL5 | MTL5 Entrez, Source | metallothionein-like 5, testis-specific (tesmin) | 24436 | -0.627 | -0.6973 | Yes |
| 125 | DHX58 | 24511 | -0.647 | -0.6924 | Yes | ||
| 126 | DTX3L | DTX3L Entrez, Source | deltex 3-like (Drosophila) | 24573 | -0.670 | -0.6868 | Yes |
| 127 | IFIT2 | IFIT2 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 2 | 24672 | -0.695 | -0.6822 | Yes |
| 128 | CHRM1 | CHRM1 Entrez, Source | cholinergic receptor, muscarinic 1 | 24828 | -0.746 | -0.6793 | Yes |
| 129 | CD38 | CD38 Entrez, Source | CD38 molecule | 24844 | -0.751 | -0.6709 | Yes |
| 130 | TUFT1 | TUFT1 Entrez, Source | tuftelin 1 | 24867 | -0.763 | -0.6626 | Yes |
| 131 | SLC22A4 | SLC22A4 Entrez, Source | solute carrier family 22 (organic cation transporter), member 4 | 24933 | -0.788 | -0.6557 | Yes |
| 132 | PDGFRL | PDGFRL Entrez, Source | platelet-derived growth factor receptor-like | 25017 | -0.831 | -0.6489 | Yes |
| 133 | TRIM25 | TRIM25 Entrez, Source | tripartite motif-containing 25 | 25023 | -0.834 | -0.6391 | Yes |
| 134 | TAP2 | TAP2 Entrez, Source | transporter 2, ATP-binding cassette, sub-family B (MDR/TAP) | 25145 | -0.894 | -0.6331 | Yes |
| 135 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 25230 | -0.948 | -0.6250 | Yes |
| 136 | VSIG10L | 25240 | -0.957 | -0.6139 | Yes | ||
| 137 | SERPING1 | SERPING1 Entrez, Source | serpin peptidase inhibitor, clade G (C1 inhibitor), member 1, (angioedema, hereditary) | 25269 | -0.979 | -0.6032 | Yes |
| 138 | DACT2 | DACT2 Entrez, Source | dapper, antagonist of beta-catenin, homolog 2 (Xenopus laevis) | 25275 | -0.983 | -0.5916 | Yes |
| 139 | TLR3 | TLR3 Entrez, Source | toll-like receptor 3 | 25298 | -1.003 | -0.5805 | Yes |
| 140 | HIST1H2AC | HIST1H2AC Entrez, Source | histone cluster 1, H2ac | 25301 | -1.006 | -0.5685 | Yes |
| 141 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 25437 | -1.129 | -0.5602 | Yes |
| 142 | RBM11 | RBM11 Entrez, Source | RNA binding motif protein 11 | 25483 | -1.182 | -0.5478 | Yes |
| 143 | OAS2 | OAS2 Entrez, Source | 2'-5'-oligoadenylate synthetase 2, 69/71kDa | 25558 | -1.284 | -0.5353 | Yes |
| 144 | IFI35 | IFI35 Entrez, Source | interferon-induced protein 35 | 25621 | -1.385 | -0.5211 | Yes |
| 145 | PARP14 | PARP14 Entrez, Source | poly (ADP-ribose) polymerase family, member 14 | 25624 | -1.394 | -0.5044 | Yes |
| 146 | GBP4 | GBP4 Entrez, Source | guanylate binding protein 4 | 25655 | -1.448 | -0.4882 | Yes |
| 147 | AIM2 | AIM2 Entrez, Source | absent in melanoma 2 | 25658 | -1.467 | -0.4707 | Yes |
| 148 | PARP10 | PARP10 Entrez, Source | poly (ADP-ribose) polymerase family, member 10 | 25663 | -1.472 | -0.4532 | Yes |
| 149 | ISG15 | ISG15 Entrez, Source | ISG15 ubiquitin-like modifier | 25699 | -1.594 | -0.4355 | Yes |
| 150 | DDX60 | 25710 | -1.643 | -0.4162 | Yes | ||
| 151 | CASP12 | CASP12 Entrez, Source | caspase 12 | 25754 | -1.792 | -0.3963 | Yes |
| 152 | LGALS3BP | LGALS3BP Entrez, Source | lectin, galactoside-binding, soluble, 3 binding protein | 25761 | -1.839 | -0.3745 | Yes |
| 153 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 25769 | -1.855 | -0.3525 | Yes |
| 154 | IRF7 | IRF7 Entrez, Source | interferon regulatory factor 7 | 25794 | -2.009 | -0.3294 | Yes |
| 155 | RSAD2 | RSAD2 Entrez, Source | radical S-adenosyl methionine domain containing 2 | 25795 | -2.013 | -0.3053 | Yes |
| 156 | IFIH1 | IFIH1 Entrez, Source | interferon induced with helicase C domain 1 | 25804 | -2.052 | -0.2810 | Yes |
| 157 | XAF1 | 25822 | -2.156 | -0.2558 | Yes | ||
| 158 | USP18 | USP18 Entrez, Source | ubiquitin specific peptidase 18 | 25830 | -2.231 | -0.2293 | Yes |
| 159 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 25831 | -2.278 | -0.2020 | Yes |
| 160 | RTP4 | RTP4 Entrez, Source | receptor transporter protein 4 | 25833 | -2.300 | -0.1744 | Yes |
| 161 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 25838 | -2.365 | -0.1462 | Yes |
| 162 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 25844 | -2.414 | -0.1175 | Yes |
| 163 | NLRC5 | 25867 | -2.711 | -0.0858 | Yes | ||
| 164 | SAMD9L | SAMD9L Entrez, Source | sterile alpha motif domain containing 9-like | 25888 | -3.178 | -0.0485 | Yes |
| 165 | IFIT1 | IFIT1 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 1 | 25894 | -4.073 | 0.0002 | Yes |