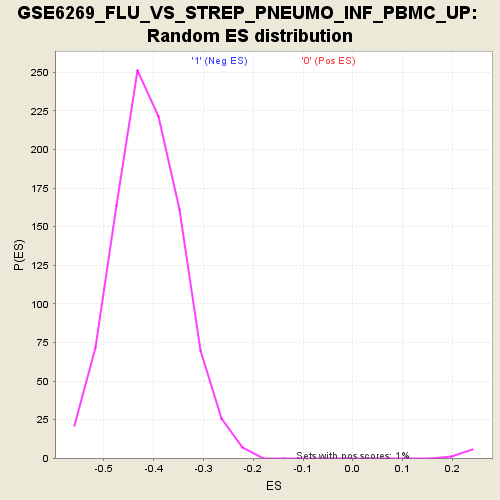
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 1 |
| GeneSet | GSE6269_FLU_VS_STREP_PNEUMO_INF_PBMC_UP |
| Enrichment Score (ES) | -0.7469971 |
| Normalized Enrichment Score (NES) | -1.8187104 |
| Nominal p-value | 0.0 |
| FDR q-value | 5.969257E-4 |
| FWER p-Value | 0.023 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SIGLEC1 | SIGLEC1 Entrez, Source | sialic acid binding Ig-like lectin 1, sialoadhesin | 114 | 0.750 | 0.0120 | No |
| 2 | IFITM1 | IFITM1 Entrez, Source | interferon induced transmembrane protein 1 (9-27) | 995 | 0.299 | -0.0156 | No |
| 3 | ADA | ADA Entrez, Source | adenosine deaminase | 1403 | 0.241 | -0.0261 | No |
| 4 | OTOF | OTOF Entrez, Source | otoferlin | 1512 | 0.230 | -0.0252 | No |
| 5 | ST3GAL5 | ST3GAL5 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 5 | 2050 | 0.172 | -0.0423 | No |
| 6 | CCDC99 | CCDC99 Entrez, Source | coiled-coil domain containing 99 | 2125 | 0.165 | -0.0416 | No |
| 7 | CXCL11 | CXCL11 Entrez, Source | chemokine (C-X-C motif) ligand 11 | 2436 | 0.157 | -0.0502 | No |
| 8 | FANCL | FANCL Entrez, Source | Fanconi anemia, complementation group L | 2469 | 0.153 | -0.0480 | No |
| 9 | GMIP | GMIP Entrez, Source | GEM interacting protein | 3527 | 0.096 | -0.0870 | No |
| 10 | HELLS | HELLS Entrez, Source | helicase, lymphoid-specific | 3542 | 0.095 | -0.0854 | No |
| 11 | SPATS2L | 3682 | 0.090 | -0.0889 | No | ||
| 12 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 3743 | 0.087 | -0.0893 | No |
| 13 | IL2RG | IL2RG Entrez, Source | interleukin 2 receptor, gamma (severe combined immunodeficiency) | 3761 | 0.087 | -0.0880 | No |
| 14 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 3988 | 0.086 | -0.0949 | No |
| 15 | ESF1 | 3994 | 0.086 | -0.0932 | No | ||
| 16 | ZBP1 | ZBP1 Entrez, Source | Z-DNA binding protein 1 | 4016 | 0.085 | -0.0922 | No |
| 17 | TRAPPC4 | TRAPPC4 Entrez, Source | trafficking protein particle complex 4 | 7986 | 0.073 | -0.2447 | No |
| 18 | PPIG | PPIG Entrez, Source | peptidylprolyl isomerase G (cyclophilin G) | 8142 | 0.071 | -0.2492 | No |
| 19 | PARP11 | PARP11 Entrez, Source | poly (ADP-ribose) polymerase family, member 11 | 8252 | 0.069 | -0.2519 | No |
| 20 | COX8A | COX8A Entrez, Source | cytochrome c oxidase subunit 8A (ubiquitous) | 8278 | 0.069 | -0.2513 | No |
| 21 | ZWILCH | ZWILCH Entrez, Source | Zwilch, kinetochore associated, homolog (Drosophila) | 8495 | 0.064 | -0.2583 | No |
| 22 | WDHD1 | WDHD1 Entrez, Source | WD repeat and HMG-box DNA binding protein 1 | 8662 | 0.061 | -0.2634 | No |
| 23 | PRR5 | PRR5 Entrez, Source | proline rich 5 (renal) | 8671 | 0.061 | -0.2624 | No |
| 24 | CEP290 | CEP290 Entrez, Source | centrosomal protein 290kDa | 8925 | 0.059 | -0.2709 | No |
| 25 | SP110 | SP110 Entrez, Source | SP110 nuclear body protein | 9080 | 0.057 | -0.2757 | No |
| 26 | MRPL42 | MRPL42 Entrez, Source | mitochondrial ribosomal protein L42 | 9124 | 0.056 | -0.2761 | No |
| 27 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 9173 | 0.055 | -0.2767 | No |
| 28 | OAS3 | OAS3 Entrez, Source | 2'-5'-oligoadenylate synthetase 3, 100kDa | 9242 | 0.055 | -0.2782 | No |
| 29 | PARP2 | PARP2 Entrez, Source | poly (ADP-ribose) polymerase family, member 2 | 9342 | 0.053 | -0.2809 | No |
| 30 | STIL | STIL Entrez, Source | SCL/TAL1 interrupting locus | 9584 | 0.049 | -0.2892 | No |
| 31 | RHBDD3 | RHBDD3 Entrez, Source | rhomboid domain containing 3 | 9609 | 0.049 | -0.2890 | No |
| 32 | NFATC2IP | NFATC2IP Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 2 interacting protein | 9722 | 0.047 | -0.2923 | No |
| 33 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 9818 | 0.046 | -0.2950 | No |
| 34 | COIL | COIL Entrez, Source | coilin | 9952 | 0.044 | -0.2992 | No |
| 35 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 10152 | 0.041 | -0.3060 | No |
| 36 | NUP62 | NUP62 Entrez, Source | nucleoporin 62kDa | 10464 | 0.037 | -0.3173 | No |
| 37 | AASDHPPT | AASDHPPT Entrez, Source | aminoadipate-semialdehyde dehydrogenase-phosphopantetheinyl transferase | 10593 | 0.035 | -0.3215 | No |
| 38 | PRPF40A | PRPF40A Entrez, Source | PRP40 pre-mRNA processing factor 40 homolog A (yeast) | 10612 | 0.035 | -0.3214 | No |
| 39 | NAA15 | 10629 | 0.035 | -0.3213 | No | ||
| 40 | GBP1 | GBP1 Entrez, Source | guanylate binding protein 1, interferon-inducible, 67kDa | 10631 | 0.035 | -0.3205 | No |
| 41 | RFWD3 | RFWD3 Entrez, Source | ring finger and WD repeat domain 3 | 10819 | 0.033 | -0.3271 | No |
| 42 | CHMP5 | CHMP5 Entrez, Source | chromatin modifying protein 5 | 10912 | 0.031 | -0.3300 | No |
| 43 | TNIK | TNIK Entrez, Source | TRAF2 and NCK interacting kinase | 11151 | 0.029 | -0.3386 | No |
| 44 | THOC2 | THOC2 Entrez, Source | THO complex 2 | 11222 | 0.028 | -0.3407 | No |
| 45 | PPPDE1 | 11244 | 0.028 | -0.3409 | No | ||
| 46 | EIF5B | EIF5B Entrez, Source | eukaryotic translation initiation factor 5B | 11284 | 0.027 | -0.3418 | No |
| 47 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 11315 | 0.027 | -0.3424 | No |
| 48 | NAA35 | 11374 | 0.026 | -0.3441 | No | ||
| 49 | MKLN1 | MKLN1 Entrez, Source | muskelin 1, intracellular mediator containing kelch motifs | 11429 | 0.025 | -0.3456 | No |
| 50 | RBM25 | RBM25 Entrez, Source | RNA binding motif protein 25 | 11703 | 0.022 | -0.3557 | No |
| 51 | IDH2 | IDH2 Entrez, Source | isocitrate dehydrogenase 2 (NADP+), mitochondrial | 11732 | 0.022 | -0.3563 | No |
| 52 | DHX29 | DHX29 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 29 | 11770 | 0.022 | -0.3573 | No |
| 53 | MED1 | 11875 | 0.020 | -0.3609 | No | ||
| 54 | GART | GART Entrez, Source | phosphoribosylglycinamide formyltransferase, phosphoribosylglycinamide synthetase, phosphoribosylaminoimidazole synthetase | 11889 | 0.020 | -0.3609 | No |
| 55 | DCAF15 | 11908 | 0.020 | -0.3612 | No | ||
| 56 | MED24 | 11910 | 0.020 | -0.3608 | No | ||
| 57 | CHD4 | CHD4 Entrez, Source | chromodomain helicase DNA binding protein 4 | 11996 | 0.019 | -0.3637 | No |
| 58 | CALR | CALR Entrez, Source | calreticulin | 12016 | 0.019 | -0.3640 | No |
| 59 | SETX | SETX Entrez, Source | senataxin | 12146 | 0.017 | -0.3686 | No |
| 60 | CKS2 | CKS2 Entrez, Source | CDC28 protein kinase regulatory subunit 2 | 12217 | 0.016 | -0.3710 | No |
| 61 | DDX39B | 12240 | 0.016 | -0.3715 | No | ||
| 62 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 12263 | 0.016 | -0.3720 | No |
| 63 | SCAF11 | 12385 | 0.014 | -0.3764 | No | ||
| 64 | FAM70A | FAM70A Entrez, Source | family with sequence similarity 70, member A | 12481 | 0.013 | -0.3798 | No |
| 65 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 12582 | 0.012 | -0.3835 | No |
| 66 | PABPN1 | PABPN1 Entrez, Source | poly(A) binding protein, nuclear 1 | 12852 | 0.009 | -0.3937 | No |
| 67 | UBXN4 | 12891 | 0.008 | -0.3950 | No | ||
| 68 | ACTN4 | ACTN4 Entrez, Source | actinin, alpha 4 | 13154 | 0.005 | -0.4051 | No |
| 69 | BPTF | 13209 | 0.004 | -0.4071 | No | ||
| 70 | N4BP1 | N4BP1 Entrez, Source | - | 13482 | 0.001 | -0.4176 | No |
| 71 | PTPLAD1 | PTPLAD1 Entrez, Source | protein tyrosine phosphatase-like A domain containing 1 | 13552 | -0.000 | -0.4203 | No |
| 72 | PHACTR2 | PHACTR2 Entrez, Source | phosphatase and actin regulator 2 | 13567 | -0.000 | -0.4209 | No |
| 73 | RORA | RORA Entrez, Source | RAR-related orphan receptor A | 13692 | -0.002 | -0.4256 | No |
| 74 | GLYR1 | 13743 | -0.003 | -0.4275 | No | ||
| 75 | SART3 | SART3 Entrez, Source | squamous cell carcinoma antigen recognised by T cells 3 | 13887 | -0.004 | -0.4330 | No |
| 76 | ERGIC2 | ERGIC2 Entrez, Source | ERGIC and golgi 2 | 13929 | -0.005 | -0.4345 | No |
| 77 | SMG7 | SMG7 Entrez, Source | Smg-7 homolog, nonsense mediated mRNA decay factor (C. elegans) | 14127 | -0.007 | -0.4420 | No |
| 78 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 14130 | -0.007 | -0.4419 | No |
| 79 | PURA | PURA Entrez, Source | purine-rich element binding protein A | 14253 | -0.009 | -0.4465 | No |
| 80 | TRIM14 | TRIM14 Entrez, Source | tripartite motif-containing 14 | 14280 | -0.009 | -0.4473 | No |
| 81 | GLG1 | GLG1 Entrez, Source | golgi apparatus protein 1 | 14326 | -0.010 | -0.4488 | No |
| 82 | ABL1 | ABL1 Entrez, Source | v-abl Abelson murine leukemia viral oncogene homolog 1 | 14354 | -0.010 | -0.4496 | No |
| 83 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 14481 | -0.012 | -0.4543 | No |
| 84 | PPID | PPID Entrez, Source | peptidylprolyl isomerase D (cyclophilin D) | 14638 | -0.014 | -0.4600 | No |
| 85 | SON | SON Entrez, Source | SON DNA binding protein | 14648 | -0.014 | -0.4601 | No |
| 86 | HEG1 | HEG1 Entrez, Source | HEG homolog 1 (zebrafish) | 14675 | -0.014 | -0.4607 | No |
| 87 | ADAR | ADAR Entrez, Source | adenosine deaminase, RNA-specific | 14749 | -0.016 | -0.4632 | No |
| 88 | TYMP | 14791 | -0.016 | -0.4645 | No | ||
| 89 | TAF1B | TAF1B Entrez, Source | TATA box binding protein (TBP)-associated factor, RNA polymerase I, B, 63kDa | 14853 | -0.017 | -0.4665 | No |
| 90 | MARK2 | MARK2 Entrez, Source | MAP/microtubule affinity-regulating kinase 2 | 15012 | -0.020 | -0.4722 | No |
| 91 | MLEC | 15030 | -0.020 | -0.4724 | No | ||
| 92 | SMG1 | SMG1 Entrez, Source | - | 15090 | -0.021 | -0.4742 | No |
| 93 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 15118 | -0.021 | -0.4748 | No |
| 94 | DDX24 | DDX24 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 24 | 15150 | -0.022 | -0.4755 | No |
| 95 | AP3D1 | AP3D1 Entrez, Source | adaptor-related protein complex 3, delta 1 subunit | 15229 | -0.023 | -0.4780 | No |
| 96 | HMGXB3 | 15252 | -0.023 | -0.4784 | No | ||
| 97 | PHF11 | PHF11 Entrez, Source | PHD finger protein 11 | 15262 | -0.024 | -0.4782 | No |
| 98 | GPATCH8 | 15675 | -0.029 | -0.4936 | No | ||
| 99 | IL12RB1 | IL12RB1 Entrez, Source | interleukin 12 receptor, beta 1 | 15965 | -0.034 | -0.5041 | No |
| 100 | SBF1 | SBF1 Entrez, Source | SET binding factor 1 | 15967 | -0.034 | -0.5034 | No |
| 101 | C2CD3 | 16060 | -0.035 | -0.5062 | No | ||
| 102 | PDHX | PDHX Entrez, Source | pyruvate dehydrogenase complex, component X | 16336 | -0.036 | -0.5160 | No |
| 103 | TMEM223 | 16358 | -0.036 | -0.5161 | No | ||
| 104 | RPL28 | RPL28 Entrez, Source | ribosomal protein L28 | 16548 | -0.040 | -0.5225 | No |
| 105 | FTSJD2 | 16599 | -0.041 | -0.5236 | No | ||
| 106 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 16617 | -0.041 | -0.5233 | No |
| 107 | CD2AP | CD2AP Entrez, Source | CD2-associated protein | 16696 | -0.043 | -0.5254 | No |
| 108 | SLC25A11 | SLC25A11 Entrez, Source | solute carrier family 25 (mitochondrial carrier; oxoglutarate carrier), member 11 | 16830 | -0.045 | -0.5296 | No |
| 109 | AKAP1 | AKAP1 Entrez, Source | A kinase (PRKA) anchor protein 1 | 16890 | -0.046 | -0.5309 | No |
| 110 | JUP | JUP Entrez, Source | junction plakoglobin | 17084 | -0.050 | -0.5373 | No |
| 111 | YPEL1 | YPEL1 Entrez, Source | yippee-like 1 (Drosophila) | 17353 | -0.056 | -0.5464 | No |
| 112 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 17473 | -0.058 | -0.5498 | No |
| 113 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 17701 | -0.063 | -0.5572 | No |
| 114 | MED15 | 17944 | -0.069 | -0.5651 | No | ||
| 115 | IFITM2 | IFITM2 Entrez, Source | interferon induced transmembrane protein 2 (1-8D) | 18119 | -0.074 | -0.5702 | No |
| 116 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 18571 | -0.087 | -0.5858 | No |
| 117 | UBE2L6 | UBE2L6 Entrez, Source | ubiquitin-conjugating enzyme E2L 6 | 18633 | -0.089 | -0.5862 | No |
| 118 | IFITM3 | IFITM3 Entrez, Source | interferon induced transmembrane protein 3 (1-8U) | 19157 | -0.106 | -0.6042 | No |
| 119 | PML | PML Entrez, Source | promyelocytic leukemia | 20690 | -0.171 | -0.6600 | No |
| 120 | HERC6 | HERC6 Entrez, Source | hect domain and RLD 6 | 21820 | -0.248 | -0.6983 | No |
| 121 | EIF2AK2 | EIF2AK2 Entrez, Source | eukaryotic translation initiation factor 2-alpha kinase 2 | 21911 | -0.256 | -0.6962 | No |
| 122 | IRF9 | 22485 | -0.307 | -0.7117 | No | ||
| 123 | LY6E | LY6E Entrez, Source | lymphocyte antigen 6 complex, locus E | 22607 | -0.320 | -0.7094 | No |
| 124 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 23259 | -0.399 | -0.7259 | No |
| 125 | PARP12 | PARP12 Entrez, Source | poly (ADP-ribose) polymerase family, member 12 | 23548 | -0.441 | -0.7274 | No |
| 126 | CHST12 | CHST12 Entrez, Source | carbohydrate (chondroitin 4) sulfotransferase 12 | 23563 | -0.444 | -0.7182 | No |
| 127 | KCTD14 | KCTD14 Entrez, Source | potassium channel tetramerisation domain containing 14 | 23568 | -0.444 | -0.7086 | No |
| 128 | PSME2 | PSME2 Entrez, Source | proteasome (prosome, macropain) activator subunit 2 (PA28 beta) | 23724 | -0.470 | -0.7043 | No |
| 129 | REC8 | 24743 | -0.715 | -0.7281 | Yes | ||
| 130 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 25230 | -0.948 | -0.7262 | Yes |
| 131 | SERPING1 | SERPING1 Entrez, Source | serpin peptidase inhibitor, clade G (C1 inhibitor), member 1, (angioedema, hereditary) | 25269 | -0.979 | -0.7062 | Yes |
| 132 | TRANK1 | 25274 | -0.982 | -0.6847 | Yes | ||
| 133 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 25437 | -1.129 | -0.6662 | Yes |
| 134 | OAS2 | OAS2 Entrez, Source | 2'-5'-oligoadenylate synthetase 2, 69/71kDa | 25558 | -1.284 | -0.6427 | Yes |
| 135 | IFI35 | IFI35 Entrez, Source | interferon-induced protein 35 | 25621 | -1.385 | -0.6147 | Yes |
| 136 | ISG15 | ISG15 Entrez, Source | ISG15 ubiquitin-like modifier | 25699 | -1.594 | -0.5827 | Yes |
| 137 | DDX60 | 25710 | -1.643 | -0.5470 | Yes | ||
| 138 | LGALS3BP | LGALS3BP Entrez, Source | lectin, galactoside-binding, soluble, 3 binding protein | 25761 | -1.839 | -0.5085 | Yes |
| 139 | PSMB9 | PSMB9 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) | 25769 | -1.855 | -0.4681 | Yes |
| 140 | IRF7 | IRF7 Entrez, Source | interferon regulatory factor 7 | 25794 | -2.009 | -0.4249 | Yes |
| 141 | RSAD2 | RSAD2 Entrez, Source | radical S-adenosyl methionine domain containing 2 | 25795 | -2.013 | -0.3807 | Yes |
| 142 | IFIH1 | IFIH1 Entrez, Source | interferon induced with helicase C domain 1 | 25804 | -2.052 | -0.3359 | Yes |
| 143 | XAF1 | 25822 | -2.156 | -0.2892 | Yes | ||
| 144 | USP18 | USP18 Entrez, Source | ubiquitin specific peptidase 18 | 25830 | -2.231 | -0.2405 | Yes |
| 145 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 25831 | -2.278 | -0.1905 | Yes |
| 146 | RTP4 | RTP4 Entrez, Source | receptor transporter protein 4 | 25833 | -2.300 | -0.1400 | Yes |
| 147 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 25844 | -2.414 | -0.0874 | Yes |
| 148 | IFIT1 | IFIT1 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 1 | 25894 | -4.073 | 0.0002 | Yes |