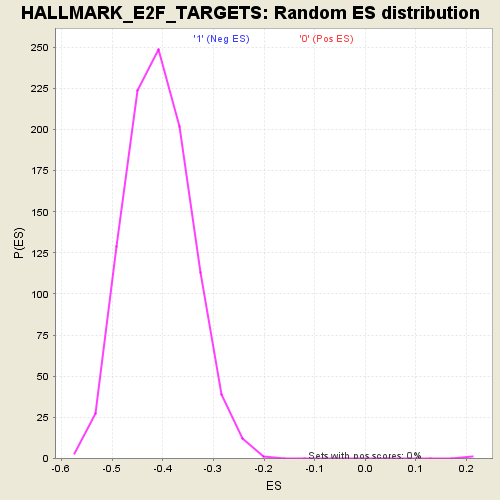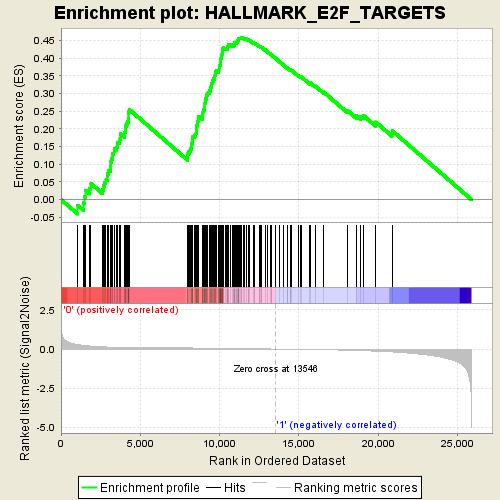
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 0 |
| GeneSet | HALLMARK_E2F_TARGETS |
| Enrichment Score (ES) | 0.45815733 |
| Normalized Enrichment Score (NES) | 1.9776442 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.1352297 |
| FWER p-Value | 0.476 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RAD1 | RAD1 Entrez, Source | RAD1 homolog (S. pombe) | 1045 | 0.290 | -0.0155 | Yes |
| 2 | DCLRE1B | DCLRE1B Entrez, Source | DNA cross-link repair 1B (PSO2 homolog, S. cerevisiae) | 1421 | 0.239 | -0.0093 | Yes |
| 3 | ASF1B | ASF1B Entrez, Source | ASF1 anti-silencing function 1 homolog B (S. cerevisiae) | 1459 | 0.234 | 0.0096 | Yes |
| 4 | GINS3 | GINS3 Entrez, Source | GINS complex subunit 3 (Psf3 homolog) | 1544 | 0.224 | 0.0258 | Yes |
| 5 | GINS1 | GINS1 Entrez, Source | GINS complex subunit 1 (Psf1 homolog) | 1800 | 0.195 | 0.0328 | Yes |
| 6 | RPA3 | RPA3 Entrez, Source | replication protein A3, 14kDa | 1886 | 0.184 | 0.0454 | Yes |
| 7 | RAD51AP1 | RAD51AP1 Entrez, Source | RAD51 associated protein 1 | 2626 | 0.146 | 0.0294 | Yes |
| 8 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 2702 | 0.141 | 0.0388 | Yes |
| 9 | DSCC1 | 2754 | 0.137 | 0.0487 | Yes | ||
| 10 | RAD50 | RAD50 Entrez, Source | RAD50 homolog (S. cerevisiae) | 2833 | 0.133 | 0.0571 | Yes |
| 11 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 2928 | 0.126 | 0.0645 | Yes |
| 12 | CKS1B | CKS1B Entrez, Source | CDC28 protein kinase regulatory subunit 1B | 2942 | 0.125 | 0.0749 | Yes |
| 13 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 3012 | 0.121 | 0.0827 | Yes |
| 14 | MMS22L | 3118 | 0.115 | 0.0885 | Yes | ||
| 15 | TIPIN | TIPIN Entrez, Source | TIMELESS interacting protein | 3139 | 0.114 | 0.0976 | Yes |
| 16 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 3146 | 0.113 | 0.1072 | Yes |
| 17 | SPC24 | 3195 | 0.111 | 0.1150 | Yes | ||
| 18 | NUP107 | NUP107 Entrez, Source | nucleoporin 107kDa | 3247 | 0.108 | 0.1224 | Yes |
| 19 | PLK4 | PLK4 Entrez, Source | polo-like kinase 4 (Drosophila) | 3251 | 0.108 | 0.1316 | Yes |
| 20 | WDR90 | WDR90 Entrez, Source | WD repeat domain 90 | 3339 | 0.103 | 0.1372 | Yes |
| 21 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 3361 | 0.102 | 0.1453 | Yes |
| 22 | LBR | LBR Entrez, Source | lamin B receptor | 3497 | 0.098 | 0.1485 | Yes |
| 23 | HELLS | HELLS Entrez, Source | helicase, lymphoid-specific | 3542 | 0.095 | 0.1550 | Yes |
| 24 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 3571 | 0.094 | 0.1621 | Yes |
| 25 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 3691 | 0.090 | 0.1653 | Yes |
| 26 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 3717 | 0.088 | 0.1720 | Yes |
| 27 | AURKA | AURKA Entrez, Source | aurora kinase A | 3750 | 0.087 | 0.1783 | Yes |
| 28 | STAG1 | STAG1 Entrez, Source | stromal antigen 1 | 3751 | 0.087 | 0.1859 | Yes |
| 29 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 3988 | 0.086 | 0.1841 | Yes |
| 30 | MTHFD2 | MTHFD2 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 2, methenyltetrahydrofolate cyclohydrolase | 4027 | 0.084 | 0.1900 | Yes |
| 31 | PRIM2 | 4066 | 0.083 | 0.1957 | Yes | ||
| 32 | MAD2L1 | MAD2L1 Entrez, Source | MAD2 mitotic arrest deficient-like 1 (yeast) | 4081 | 0.083 | 0.2024 | Yes |
| 33 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 4087 | 0.082 | 0.2093 | Yes |
| 34 | ORC2 | 4143 | 0.080 | 0.2142 | Yes | ||
| 35 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 4179 | 0.079 | 0.2197 | Yes |
| 36 | ASF1A | ASF1A Entrez, Source | ASF1 anti-silencing function 1 homolog A (S. cerevisiae) | 4233 | 0.077 | 0.2243 | Yes |
| 37 | SMC6 | SMC6 Entrez, Source | structural maintenance of chromosomes 6 | 4260 | 0.077 | 0.2300 | Yes |
| 38 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 4267 | 0.076 | 0.2364 | Yes |
| 39 | NBN | NBN Entrez, Source | nibrin | 4268 | 0.076 | 0.2430 | Yes |
| 40 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 4276 | 0.076 | 0.2493 | Yes |
| 41 | DCTPP1 | 4335 | 0.074 | 0.2535 | Yes | ||
| 42 | SRSF1 | 7992 | 0.073 | 0.1177 | Yes | ||
| 43 | SLBP | SLBP Entrez, Source | stem-loop (histone) binding protein | 7995 | 0.073 | 0.1240 | Yes |
| 44 | E2F8 | E2F8 Entrez, Source | E2F transcription factor 8 | 8004 | 0.073 | 0.1300 | Yes |
| 45 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 8040 | 0.072 | 0.1349 | Yes |
| 46 | LYAR | LYAR Entrez, Source | - | 8134 | 0.072 | 0.1375 | Yes |
| 47 | GINS4 | GINS4 Entrez, Source | GINS complex subunit 4 (Sld5 homolog) | 8144 | 0.071 | 0.1433 | Yes |
| 48 | TMPO | TMPO Entrez, Source | thymopoietin | 8207 | 0.070 | 0.1470 | Yes |
| 49 | NUDT21 | NUDT21 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 21 | 8235 | 0.069 | 0.1520 | Yes |
| 50 | MELK | MELK Entrez, Source | maternal embryonic leucine zipper kinase | 8236 | 0.069 | 0.1580 | Yes |
| 51 | RQCD1 | RQCD1 Entrez, Source | RCD1 required for cell differentiation1 homolog (S. pombe) | 8264 | 0.069 | 0.1629 | Yes |
| 52 | PSMC3IP | PSMC3IP Entrez, Source | PSMC3 interacting protein | 8290 | 0.068 | 0.1679 | Yes |
| 53 | POLE4 | POLE4 Entrez, Source | polymerase (DNA-directed), epsilon 4 (p12 subunit) | 8300 | 0.068 | 0.1735 | Yes |
| 54 | DLGAP5 | 8319 | 0.068 | 0.1787 | Yes | ||
| 55 | EXOSC8 | EXOSC8 Entrez, Source | exosome component 8 | 8408 | 0.066 | 0.1810 | Yes |
| 56 | NAA38 | 8477 | 0.065 | 0.1840 | Yes | ||
| 57 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 8511 | 0.064 | 0.1882 | Yes |
| 58 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 8541 | 0.063 | 0.1926 | Yes |
| 59 | CDK1 | 8548 | 0.063 | 0.1978 | Yes | ||
| 60 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 8553 | 0.063 | 0.2031 | Yes |
| 61 | SPC25 | 8559 | 0.063 | 0.2084 | Yes | ||
| 62 | MCM4 | MCM4 Entrez, Source | MCM4 minichromosome maintenance deficient 4 (S. cerevisiae) | 8582 | 0.062 | 0.2129 | Yes |
| 63 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 8593 | 0.062 | 0.2179 | Yes |
| 64 | EZH2 | EZH2 Entrez, Source | enhancer of zeste homolog 2 (Drosophila) | 8626 | 0.061 | 0.2220 | Yes |
| 65 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 8646 | 0.061 | 0.2266 | Yes |
| 66 | RNASEH2A | RNASEH2A Entrez, Source | ribonuclease H2, subunit A | 8648 | 0.061 | 0.2318 | Yes |
| 67 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 8688 | 0.060 | 0.2356 | Yes |
| 68 | KIF18B | 8919 | 0.060 | 0.2318 | Yes | ||
| 69 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 8928 | 0.059 | 0.2366 | Yes |
| 70 | CSE1L | CSE1L Entrez, Source | CSE1 chromosome segregation 1-like (yeast) | 8948 | 0.059 | 0.2410 | Yes |
| 71 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 8951 | 0.059 | 0.2460 | Yes |
| 72 | PNN | PNN Entrez, Source | pinin, desmosome associated protein | 8960 | 0.059 | 0.2508 | Yes |
| 73 | SMC3 | SMC3 Entrez, Source | structural maintenance of chromosomes 3 | 8964 | 0.059 | 0.2558 | Yes |
| 74 | RAD21 | RAD21 Entrez, Source | RAD21 homolog (S. pombe) | 9065 | 0.057 | 0.2569 | Yes |
| 75 | ZW10 | ZW10 Entrez, Source | ZW10, kinetochore associated, homolog (Drosophila) | 9070 | 0.057 | 0.2617 | Yes |
| 76 | MCM5 | MCM5 Entrez, Source | MCM5 minichromosome maintenance deficient 5, cell division cycle 46 (S. cerevisiae) | 9075 | 0.057 | 0.2665 | Yes |
| 77 | DCK | DCK Entrez, Source | deoxycytidine kinase | 9076 | 0.057 | 0.2714 | Yes |
| 78 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 9097 | 0.057 | 0.2756 | Yes |
| 79 | RANBP1 | RANBP1 Entrez, Source | RAN binding protein 1 | 9100 | 0.057 | 0.2804 | Yes |
| 80 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 9113 | 0.056 | 0.2848 | Yes |
| 81 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 9173 | 0.055 | 0.2873 | Yes |
| 82 | POLE | POLE Entrez, Source | polymerase (DNA directed), epsilon | 9191 | 0.055 | 0.2914 | Yes |
| 83 | TACC3 | TACC3 Entrez, Source | transforming, acidic coiled-coil containing protein 3 | 9196 | 0.055 | 0.2960 | Yes |
| 84 | RACGAP1 | RACGAP1 Entrez, Source | Rac GTPase activating protein 1 | 9253 | 0.055 | 0.2986 | Yes |
| 85 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9270 | 0.054 | 0.3026 | Yes |
| 86 | PAICS | PAICS Entrez, Source | phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase | 9343 | 0.053 | 0.3044 | Yes |
| 87 | MLH1 | MLH1 Entrez, Source | mutL homolog 1, colon cancer, nonpolyposis type 2 (E. coli) | 9347 | 0.053 | 0.3089 | Yes |
| 88 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 9427 | 0.052 | 0.3103 | Yes |
| 89 | SSRP1 | SSRP1 Entrez, Source | structure specific recognition protein 1 | 9429 | 0.052 | 0.3147 | Yes |
| 90 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 9453 | 0.051 | 0.3183 | Yes |
| 91 | DONSON | DONSON Entrez, Source | downstream neighbor of SON | 9489 | 0.051 | 0.3213 | Yes |
| 92 | PSIP1 | PSIP1 Entrez, Source | PC4 and SFRS1 interacting protein 1 | 9503 | 0.050 | 0.3252 | Yes |
| 93 | MSH2 | MSH2 Entrez, Source | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | 9516 | 0.050 | 0.3291 | Yes |
| 94 | CENPE | CENPE Entrez, Source | centromere protein E, 312kDa | 9548 | 0.050 | 0.3322 | Yes |
| 95 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 9554 | 0.050 | 0.3363 | Yes |
| 96 | SMC1A | SMC1A Entrez, Source | structural maintenance of chromosomes 1A | 9605 | 0.049 | 0.3386 | Yes |
| 97 | MCM3 | MCM3 Entrez, Source | MCM3 minichromosome maintenance deficient 3 (S. cerevisiae) | 9611 | 0.049 | 0.3427 | Yes |
| 98 | POLD2 | POLD2 Entrez, Source | polymerase (DNA directed), delta 2, regulatory subunit 50kDa | 9681 | 0.048 | 0.3441 | Yes |
| 99 | POLD3 | POLD3 Entrez, Source | polymerase (DNA-directed), delta 3, accessory subunit | 9698 | 0.048 | 0.3477 | Yes |
| 100 | HMGB3 | HMGB3 Entrez, Source | high-mobility group box 3 | 9708 | 0.047 | 0.3514 | Yes |
| 101 | AURKB | AURKB Entrez, Source | aurora kinase B | 9747 | 0.047 | 0.3540 | Yes |
| 102 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 9757 | 0.047 | 0.3577 | Yes |
| 103 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 9766 | 0.047 | 0.3615 | Yes |
| 104 | SNRPB | SNRPB Entrez, Source | small nuclear ribonucleoprotein polypeptides B and B1 | 9810 | 0.046 | 0.3638 | Yes |
| 105 | PHF5A | PHF5A Entrez, Source | PHD finger protein 5A | 9937 | 0.045 | 0.3628 | Yes |
| 106 | DEK | DEK Entrez, Source | DEK oncogene (DNA binding) | 9948 | 0.044 | 0.3662 | Yes |
| 107 | SRSF2 | 9966 | 0.044 | 0.3694 | Yes | ||
| 108 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 9968 | 0.044 | 0.3732 | Yes |
| 109 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 9993 | 0.044 | 0.3760 | Yes |
| 110 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 10017 | 0.043 | 0.3789 | Yes |
| 111 | PPM1D | PPM1D Entrez, Source | protein phosphatase 1D magnesium-dependent, delta isoform | 10052 | 0.043 | 0.3813 | Yes |
| 112 | UBR7 | 10059 | 0.043 | 0.3848 | Yes | ||
| 113 | RAN | RAN Entrez, Source | RAN, member RAS oncogene family | 10062 | 0.043 | 0.3884 | Yes |
| 114 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 10082 | 0.042 | 0.3913 | Yes |
| 115 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 10084 | 0.042 | 0.3950 | Yes |
| 116 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10086 | 0.042 | 0.3986 | Yes |
| 117 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 10107 | 0.042 | 0.4015 | Yes |
| 118 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 10123 | 0.042 | 0.4045 | Yes |
| 119 | PDS5B | 10137 | 0.042 | 0.4076 | Yes | ||
| 120 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 10152 | 0.041 | 0.4107 | Yes |
| 121 | CBX5 | CBX5 Entrez, Source | chromobox homolog 5 (HP1 alpha homolog, Drosophila) | 10161 | 0.041 | 0.4140 | Yes |
| 122 | LUC7L3 | 10186 | 0.041 | 0.4166 | Yes | ||
| 123 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 10197 | 0.041 | 0.4198 | Yes |
| 124 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 10207 | 0.041 | 0.4230 | Yes |
| 125 | CDCA3 | CDCA3 Entrez, Source | cell division cycle associated 3 | 10212 | 0.041 | 0.4263 | Yes |
| 126 | IPO7 | IPO7 Entrez, Source | importin 7 | 10227 | 0.040 | 0.4293 | Yes |
| 127 | MYBL2 | MYBL2 Entrez, Source | v-myb myeloblastosis viral oncogene homolog (avian)-like 2 | 10368 | 0.039 | 0.4272 | Yes |
| 128 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 10437 | 0.037 | 0.4278 | Yes |
| 129 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 10509 | 0.036 | 0.4282 | Yes |
| 130 | NOP56 | 10512 | 0.036 | 0.4313 | Yes | ||
| 131 | SUV39H1 | SUV39H1 Entrez, Source | suppressor of variegation 3-9 homolog 1 (Drosophila) | 10524 | 0.036 | 0.4340 | Yes |
| 132 | BRMS1L | BRMS1L Entrez, Source | breast cancer metastasis-suppressor 1-like | 10562 | 0.036 | 0.4357 | Yes |
| 133 | ORC6 | 10564 | 0.036 | 0.4387 | Yes | ||
| 134 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 10680 | 0.034 | 0.4372 | Yes |
| 135 | NME1 | NME1 Entrez, Source | non-metastatic cells 1, protein (NM23A) expressed in | 10797 | 0.033 | 0.4356 | Yes |
| 136 | PAN2 | 10865 | 0.032 | 0.4358 | Yes | ||
| 137 | NCAPD2 | NCAPD2 Entrez, Source | non-SMC condensin I complex, subunit D2 | 10892 | 0.032 | 0.4375 | Yes |
| 138 | HUS1 | HUS1 Entrez, Source | HUS1 checkpoint homolog (S. pombe) | 10921 | 0.031 | 0.4391 | Yes |
| 139 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 10939 | 0.031 | 0.4412 | Yes |
| 140 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 10950 | 0.031 | 0.4434 | Yes |
| 141 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 11033 | 0.030 | 0.4429 | Yes |
| 142 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 11053 | 0.030 | 0.4447 | Yes |
| 143 | PA2G4 | PA2G4 Entrez, Source | proliferation-associated 2G4, 38kDa | 11100 | 0.029 | 0.4455 | Yes |
| 144 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 11118 | 0.029 | 0.4473 | Yes |
| 145 | ATAD2 | ATAD2 Entrez, Source | ATPase family, AAA domain containing 2 | 11145 | 0.029 | 0.4488 | Yes |
| 146 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 11167 | 0.029 | 0.4505 | Yes |
| 147 | TRA2B | 11181 | 0.028 | 0.4524 | Yes | ||
| 148 | RRM2 | RRM2 Entrez, Source | ribonucleotide reductase M2 polypeptide | 11196 | 0.028 | 0.4544 | Yes |
| 149 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 11214 | 0.028 | 0.4561 | Yes |
| 150 | PMS2 | PMS2 Entrez, Source | PMS2 postmeiotic segregation increased 2 (S. cerevisiae) | 11273 | 0.027 | 0.4562 | Yes |
| 151 | RFC2 | RFC2 Entrez, Source | replication factor C (activator 1) 2, 40kDa | 11328 | 0.027 | 0.4564 | Yes |
| 152 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 11375 | 0.026 | 0.4569 | Yes |
| 153 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 11402 | 0.026 | 0.4582 | Yes |
| 154 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 11514 | 0.024 | 0.4560 | No |
| 155 | NUP205 | NUP205 Entrez, Source | nucleoporin 205kDa | 11583 | 0.024 | 0.4554 | No |
| 156 | ESPL1 | ESPL1 Entrez, Source | extra spindle poles like 1 (S. cerevisiae) | 11672 | 0.023 | 0.4539 | No |
| 157 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 11795 | 0.021 | 0.4510 | No |
| 158 | PPP1R8 | PPP1R8 Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 8 | 11831 | 0.021 | 0.4515 | No |
| 159 | TK1 | TK1 Entrez, Source | thymidine kinase 1, soluble | 11893 | 0.020 | 0.4509 | No |
| 160 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 12135 | 0.017 | 0.4430 | No |
| 161 | CKS2 | CKS2 Entrez, Source | CDC28 protein kinase regulatory subunit 2 | 12217 | 0.016 | 0.4412 | No |
| 162 | CENPM | CENPM Entrez, Source | centromere protein M | 12220 | 0.016 | 0.4425 | No |
| 163 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 12492 | 0.013 | 0.4331 | No |
| 164 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 12512 | 0.012 | 0.4334 | No |
| 165 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 12568 | 0.012 | 0.4323 | No |
| 166 | SHMT1 | SHMT1 Entrez, Source | serine hydroxymethyltransferase 1 (soluble) | 12642 | 0.011 | 0.4304 | No |
| 167 | UBE2T | UBE2T Entrez, Source | ubiquitin-conjugating enzyme E2T (putative) | 12668 | 0.011 | 0.4303 | No |
| 168 | RPA1 | RPA1 Entrez, Source | replication protein A1, 70kDa | 12870 | 0.008 | 0.4232 | No |
| 169 | HNRNPD | 13015 | 0.006 | 0.4182 | No | ||
| 170 | POP7 | POP7 Entrez, Source | processing of precursor 7, ribonuclease P subunit (S. cerevisiae) | 13224 | 0.004 | 0.4104 | No |
| 171 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 13293 | 0.003 | 0.4081 | No |
| 172 | CTPS | CTPS Entrez, Source | CTP synthase | 13296 | 0.003 | 0.4082 | No |
| 173 | TBRG4 | TBRG4 Entrez, Source | transforming growth factor beta regulator 4 | 13556 | -0.000 | 0.3982 | No |
| 174 | SPAG5 | SPAG5 Entrez, Source | sperm associated antigen 5 | 13814 | -0.003 | 0.3885 | No |
| 175 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 14049 | -0.006 | 0.3799 | No |
| 176 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 14264 | -0.009 | 0.3723 | No |
| 177 | CIT | CIT Entrez, Source | citron (rho-interacting, serine/threonine kinase 21) | 14306 | -0.009 | 0.3715 | No |
| 178 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 14461 | -0.012 | 0.3665 | No |
| 179 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 14481 | -0.012 | 0.3668 | No |
| 180 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 14529 | -0.012 | 0.3661 | No |
| 181 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 14958 | -0.019 | 0.3510 | No |
| 182 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 15118 | -0.021 | 0.3467 | No |
| 183 | MKI67 | MKI67 Entrez, Source | antigen identified by monoclonal antibody Ki-67 | 15149 | -0.022 | 0.3474 | No |
| 184 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 15707 | -0.029 | 0.3283 | No |
| 185 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 15717 | -0.029 | 0.3305 | No |
| 186 | EED | EED Entrez, Source | embryonic ectoderm development | 16038 | -0.035 | 0.3211 | No |
| 187 | TCF19 | TCF19 Entrez, Source | transcription factor 19 (SC1) | 16559 | -0.040 | 0.3043 | No |
| 188 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 18048 | -0.072 | 0.2527 | No |
| 189 | CCNE1 | CCNE1 Entrez, Source | cyclin E1 | 18646 | -0.089 | 0.2372 | No |
| 190 | UBE2S | UBE2S Entrez, Source | ubiquitin-conjugating enzyme E2S | 18915 | -0.097 | 0.2352 | No |
| 191 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 19069 | -0.103 | 0.2382 | No |
| 192 | PRKDC | PRKDC Entrez, Source | protein kinase, DNA-activated, catalytic polypeptide | 19853 | -0.134 | 0.2193 | No |
| 193 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 20884 | -0.182 | 0.1951 | No |