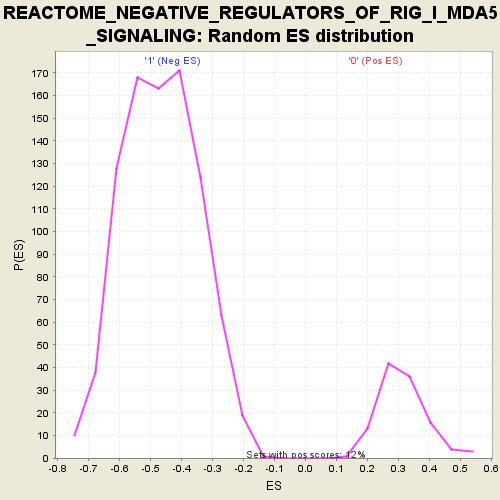
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 1 |
| GeneSet | REACTOME_NEGATIVE_REGULATORS_OF_RIG_I_MDA5_SIGNALING |
| Enrichment Score (ES) | -0.831287 |
| Normalized Enrichment Score (NES) | -1.7885009 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0018629797 |
| FWER p-Value | 0.082 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | UBE2K | 8561 | 0.063 | -0.3252 | No | ||
| 2 | IRF3 | IRF3 Entrez, Source | interferon regulatory factor 3 | 9775 | 0.046 | -0.3678 | No |
| 3 | PCBP2 | PCBP2 Entrez, Source | poly(rC) binding protein 2 | 10075 | 0.043 | -0.3755 | No |
| 4 | ATG5 | ATG5 Entrez, Source | ATG5 autophagy related 5 homolog (S. cerevisiae) | 11021 | 0.030 | -0.4093 | No |
| 5 | ATG12 | ATG12 Entrez, Source | ATG12 autophagy related 12 homolog (S. cerevisiae) | 11061 | 0.030 | -0.4081 | No |
| 6 | UBE2D1 | UBE2D1 Entrez, Source | ubiquitin-conjugating enzyme E2D 1 (UBC4/5 homolog, yeast) | 11295 | 0.027 | -0.4146 | No |
| 7 | TAX1BP1 | TAX1BP1 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 1 | 11914 | 0.020 | -0.4367 | No |
| 8 | UBA52 | UBA52 Entrez, Source | ubiquitin A-52 residue ribosomal protein fusion product 1 | 12210 | 0.016 | -0.4466 | No |
| 9 | UBE2D3 | UBE2D3 Entrez, Source | ubiquitin-conjugating enzyme E2D 3 (UBC4/5 homolog, yeast) | 12554 | 0.012 | -0.4588 | No |
| 10 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 13647 | -0.001 | -0.5009 | No |
| 11 | OTUD5 | OTUD5 Entrez, Source | OTU domain containing 5 | 13971 | -0.005 | -0.5129 | No |
| 12 | PIN1 | PIN1 Entrez, Source | protein (peptidylprolyl cis/trans isomerase) NIMA-interacting 1 | 14885 | -0.018 | -0.5466 | No |
| 13 | RPS27A | RPS27A Entrez, Source | ribosomal protein S27a | 15361 | -0.024 | -0.5627 | No |
| 14 | TBK1 | TBK1 Entrez, Source | TANK-binding kinase 1 | 15945 | -0.033 | -0.5822 | No |
| 15 | TRAF3 | TRAF3 Entrez, Source | TNF receptor-associated factor 3 | 16418 | -0.038 | -0.5970 | No |
| 16 | TNFAIP3 | TNFAIP3 Entrez, Source | tumor necrosis factor, alpha-induced protein 3 | 17675 | -0.062 | -0.6398 | No |
| 17 | NLRX1 | 17872 | -0.067 | -0.6413 | No | ||
| 18 | MAVS | 18324 | -0.079 | -0.6514 | No | ||
| 19 | UBA7 | 18403 | -0.082 | -0.6470 | No | ||
| 20 | UBE2L6 | UBE2L6 Entrez, Source | ubiquitin-conjugating enzyme E2L 6 | 18633 | -0.089 | -0.6477 | No |
| 21 | CYLD | CYLD Entrez, Source | cylindromatosis (turban tumor syndrome) | 19987 | -0.138 | -0.6874 | No |
| 22 | IKBKE | IKBKE Entrez, Source | inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase epsilon | 20727 | -0.174 | -0.7001 | No |
| 23 | DDX58 | DDX58 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 58 | 24122 | -0.545 | -0.7814 | Yes |
| 24 | TRIM25 | TRIM25 Entrez, Source | tripartite motif-containing 25 | 25023 | -0.834 | -0.7400 | Yes |
| 25 | RNF135 | RNF135 Entrez, Source | ring finger protein 135 | 25158 | -0.899 | -0.6630 | Yes |
| 26 | RNF125 | RNF125 Entrez, Source | ring finger protein 125 | 25511 | -1.209 | -0.5662 | Yes |
| 27 | ISG15 | ISG15 Entrez, Source | ISG15 ubiquitin-like modifier | 25699 | -1.594 | -0.4277 | Yes |
| 28 | IFIH1 | IFIH1 Entrez, Source | interferon induced with helicase C domain 1 | 25804 | -2.052 | -0.2442 | Yes |
| 29 | NLRC5 | 25867 | -2.711 | 0.0012 | Yes |