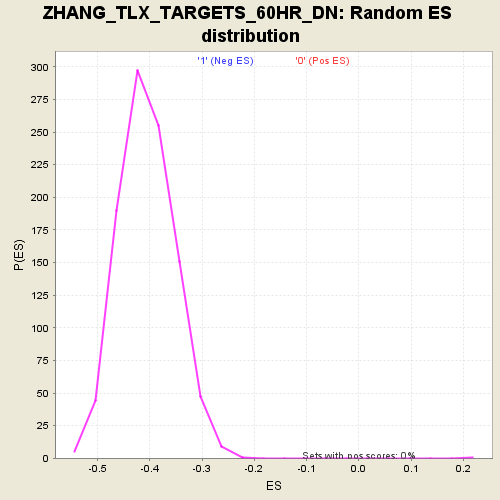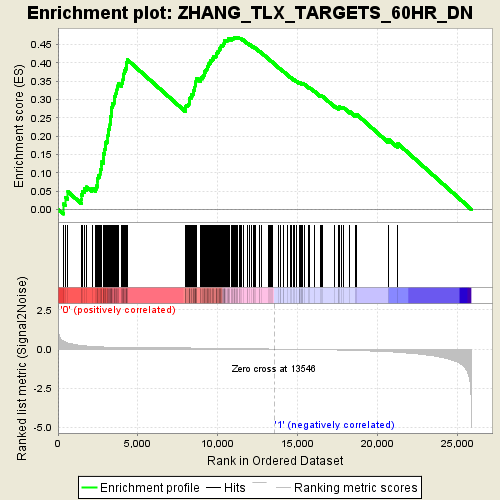
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 0 |
| GeneSet | ZHANG_TLX_TARGETS_60HR_DN |
| Enrichment Score (ES) | 0.46967688 |
| Normalized Enrichment Score (NES) | 1.9857862 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.18165211 |
| FWER p-Value | 0.444 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LIPT2 | 326 | 0.511 | 0.0148 | Yes | ||
| 2 | GSG2 | GSG2 Entrez, Source | germ cell associated 2 (haspin) | 459 | 0.442 | 0.0334 | Yes |
| 3 | GJC1 | GJC1 Entrez, Source | gap junction protein, chi 1, 31.9kDa (connexin 31.9) | 617 | 0.389 | 0.0482 | Yes |
| 4 | ASF1B | ASF1B Entrez, Source | ASF1 anti-silencing function 1 homolog B (S. cerevisiae) | 1459 | 0.234 | 0.0280 | Yes |
| 5 | LAMP3 | LAMP3 Entrez, Source | lysosomal-associated membrane protein 3 | 1475 | 0.233 | 0.0399 | Yes |
| 6 | LRR1 | 1534 | 0.225 | 0.0497 | Yes | ||
| 7 | CENPK | CENPK Entrez, Source | centromere protein K | 1652 | 0.210 | 0.0565 | Yes |
| 8 | GINS1 | GINS1 Entrez, Source | GINS complex subunit 1 (Psf1 homolog) | 1800 | 0.195 | 0.0612 | Yes |
| 9 | CCDC99 | CCDC99 Entrez, Source | coiled-coil domain containing 99 | 2125 | 0.165 | 0.0574 | Yes |
| 10 | SKP2 | SKP2 Entrez, Source | S-phase kinase-associated protein 2 (p45) | 2349 | 0.163 | 0.0575 | Yes |
| 11 | SGOL1 | SGOL1 Entrez, Source | shugoshin-like 1 (S. pombe) | 2403 | 0.160 | 0.0640 | Yes |
| 12 | ETAA1 | ETAA1 Entrez, Source | Ewing's tumor-associated antigen 1 | 2481 | 0.152 | 0.0692 | Yes |
| 13 | FAM111A | FAM111A Entrez, Source | family with sequence similarity 111, member A | 2484 | 0.152 | 0.0773 | Yes |
| 14 | ERCC6L | 2491 | 0.151 | 0.0852 | Yes | ||
| 15 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 2505 | 0.150 | 0.0928 | Yes |
| 16 | RAD51AP1 | RAD51AP1 Entrez, Source | RAD51 associated protein 1 | 2626 | 0.146 | 0.0960 | Yes |
| 17 | POLE2 | POLE2 Entrez, Source | polymerase (DNA directed), epsilon 2 (p59 subunit) | 2630 | 0.146 | 0.1037 | Yes |
| 18 | SASS6 | SASS6 Entrez, Source | spindle assembly 6 homolog (C. elegans) | 2674 | 0.143 | 0.1098 | Yes |
| 19 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 2702 | 0.141 | 0.1163 | Yes |
| 20 | CD93 | CD93 Entrez, Source | CD93 molecule | 2706 | 0.141 | 0.1238 | Yes |
| 21 | DACT1 | DACT1 Entrez, Source | dapper, antagonist of beta-catenin, homolog 1 (Xenopus laevis) | 2747 | 0.137 | 0.1296 | Yes |
| 22 | DDX20 | DDX20 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 20 | 2836 | 0.133 | 0.1333 | Yes |
| 23 | EME1 | EME1 Entrez, Source | essential meiotic endonuclease 1 homolog 1 (S. pombe) | 2841 | 0.132 | 0.1403 | Yes |
| 24 | SYCE2 | SYCE2 Entrez, Source | synaptonemal complex central element protein 2 | 2860 | 0.131 | 0.1466 | Yes |
| 25 | CDCA5 | CDCA5 Entrez, Source | cell division cycle associated 5 | 2865 | 0.130 | 0.1535 | Yes |
| 26 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 2928 | 0.126 | 0.1579 | Yes |
| 27 | FIGNL1 | FIGNL1 Entrez, Source | fidgetin-like 1 | 2930 | 0.126 | 0.1646 | Yes |
| 28 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 2971 | 0.124 | 0.1697 | Yes |
| 29 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 2983 | 0.123 | 0.1759 | Yes |
| 30 | POLA1 | POLA1 Entrez, Source | polymerase (DNA directed), alpha 1 | 2993 | 0.123 | 0.1821 | Yes |
| 31 | TTK | TTK Entrez, Source | TTK protein kinase | 3042 | 0.119 | 0.1867 | Yes |
| 32 | MMS22L | 3118 | 0.115 | 0.1899 | Yes | ||
| 33 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 3120 | 0.114 | 0.1960 | Yes |
| 34 | DHFR | DHFR Entrez, Source | dihydrofolate reductase | 3124 | 0.114 | 0.2020 | Yes |
| 35 | TIPIN | TIPIN Entrez, Source | TIMELESS interacting protein | 3139 | 0.114 | 0.2076 | Yes |
| 36 | POLR3K | POLR3K Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide K, 12.3 kDa | 3162 | 0.112 | 0.2128 | Yes |
| 37 | SKA1 | 3180 | 0.111 | 0.2181 | Yes | ||
| 38 | SHCBP1 | SHCBP1 Entrez, Source | SHC SH2-domain binding protein 1 | 3202 | 0.110 | 0.2232 | Yes |
| 39 | NUP107 | NUP107 Entrez, Source | nucleoporin 107kDa | 3247 | 0.108 | 0.2273 | Yes |
| 40 | PLK4 | PLK4 Entrez, Source | polo-like kinase 4 (Drosophila) | 3251 | 0.108 | 0.2330 | Yes |
| 41 | BUB1 | BUB1 Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) | 3261 | 0.107 | 0.2384 | Yes |
| 42 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 3268 | 0.107 | 0.2439 | Yes |
| 43 | CDC6 | CDC6 Entrez, Source | CDC6 cell division cycle 6 homolog (S. cerevisiae) | 3285 | 0.106 | 0.2490 | Yes |
| 44 | RFC4 | RFC4 Entrez, Source | replication factor C (activator 1) 4, 37kDa | 3299 | 0.105 | 0.2542 | Yes |
| 45 | WDR90 | WDR90 Entrez, Source | WD repeat domain 90 | 3339 | 0.103 | 0.2582 | Yes |
| 46 | CASP8AP2 | CASP8AP2 Entrez, Source | CASP8 associated protein 2 | 3342 | 0.103 | 0.2637 | Yes |
| 47 | CDC45 | 3350 | 0.103 | 0.2689 | Yes | ||
| 48 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 3361 | 0.102 | 0.2740 | Yes |
| 49 | GMNN | GMNN Entrez, Source | geminin, DNA replication inhibitor | 3370 | 0.102 | 0.2792 | Yes |
| 50 | CENPH | CENPH Entrez, Source | centromere protein H | 3389 | 0.102 | 0.2840 | Yes |
| 51 | BEX1 | BEX1 Entrez, Source | brain expressed, X-linked 1 | 3408 | 0.100 | 0.2887 | Yes |
| 52 | HAUS6 | 3501 | 0.097 | 0.2903 | Yes | ||
| 53 | MCM10 | MCM10 Entrez, Source | MCM10 minichromosome maintenance deficient 10 (S. cerevisiae) | 3508 | 0.097 | 0.2953 | Yes |
| 54 | CEP55 | CEP55 Entrez, Source | centrosomal protein 55kDa | 3514 | 0.097 | 0.3003 | Yes |
| 55 | EXOSC2 | EXOSC2 Entrez, Source | exosome component 2 | 3530 | 0.096 | 0.3049 | Yes |
| 56 | PKDCC | 3545 | 0.095 | 0.3094 | Yes | ||
| 57 | RAD18 | RAD18 Entrez, Source | RAD18 homolog (S. cerevisiae) | 3577 | 0.094 | 0.3133 | Yes |
| 58 | BLM | BLM Entrez, Source | Bloom syndrome | 3626 | 0.092 | 0.3163 | Yes |
| 59 | VRK1 | VRK1 Entrez, Source | vaccinia related kinase 1 | 3639 | 0.091 | 0.3208 | Yes |
| 60 | NCAPG | NCAPG Entrez, Source | non-SMC condensin I complex, subunit G | 3663 | 0.090 | 0.3248 | Yes |
| 61 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 3691 | 0.090 | 0.3285 | Yes |
| 62 | DTL | DTL Entrez, Source | denticleless homolog (Drosophila) | 3696 | 0.089 | 0.3332 | Yes |
| 63 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 3717 | 0.088 | 0.3371 | Yes |
| 64 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 3759 | 0.087 | 0.3402 | Yes |
| 65 | UCHL5 | UCHL5 Entrez, Source | ubiquitin carboxyl-terminal hydrolase L5 | 3781 | 0.086 | 0.3440 | Yes |
| 66 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 3988 | 0.086 | 0.3406 | Yes |
| 67 | CENPN | CENPN Entrez, Source | centromere protein N | 4004 | 0.085 | 0.3446 | Yes |
| 68 | NT5DC2 | NT5DC2 Entrez, Source | 5'-nucleotidase domain containing 2 | 4020 | 0.085 | 0.3486 | Yes |
| 69 | FAM64A | FAM64A Entrez, Source | family with sequence similarity 64, member A | 4062 | 0.083 | 0.3515 | Yes |
| 70 | PRIM2 | 4066 | 0.083 | 0.3559 | Yes | ||
| 71 | EXO1 | EXO1 Entrez, Source | exonuclease 1 | 4079 | 0.083 | 0.3598 | Yes |
| 72 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 4087 | 0.082 | 0.3640 | Yes |
| 73 | SHMT2 | SHMT2 Entrez, Source | serine hydroxymethyltransferase 2 (mitochondrial) | 4093 | 0.082 | 0.3682 | Yes |
| 74 | MRPL50 | MRPL50 Entrez, Source | mitochondrial ribosomal protein L50 | 4130 | 0.081 | 0.3712 | Yes |
| 75 | ORC2 | 4143 | 0.080 | 0.3750 | Yes | ||
| 76 | NCAPG2 | NCAPG2 Entrez, Source | non-SMC condensin II complex, subunit G2 | 4155 | 0.080 | 0.3789 | Yes |
| 77 | ZMYND19 | ZMYND19 Entrez, Source | zinc finger, MYND-type containing 19 | 4209 | 0.078 | 0.3810 | Yes |
| 78 | MASTL | MASTL Entrez, Source | microtubule associated serine/threonine kinase-like | 4210 | 0.078 | 0.3852 | Yes |
| 79 | SMC6 | SMC6 Entrez, Source | structural maintenance of chromosomes 6 | 4260 | 0.077 | 0.3874 | Yes |
| 80 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 4267 | 0.076 | 0.3913 | Yes |
| 81 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 4276 | 0.076 | 0.3951 | Yes |
| 82 | NUCKS1 | NUCKS1 Entrez, Source | nuclear casein kinase and cyclin-dependent kinase substrate 1 | 4278 | 0.076 | 0.3991 | Yes |
| 83 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 4293 | 0.075 | 0.4026 | Yes |
| 84 | RAD51 | RAD51 Entrez, Source | RAD51 homolog (RecA homolog, E. coli) (S. cerevisiae) | 4332 | 0.074 | 0.4052 | Yes |
| 85 | DCTPP1 | 4335 | 0.074 | 0.4091 | Yes | ||
| 86 | ANLN | ANLN Entrez, Source | anillin, actin binding protein | 7973 | 0.074 | 0.2712 | Yes |
| 87 | SRSF1 | 7992 | 0.073 | 0.2744 | Yes | ||
| 88 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 7999 | 0.073 | 0.2781 | Yes |
| 89 | E2F8 | E2F8 Entrez, Source | E2F transcription factor 8 | 8004 | 0.073 | 0.2819 | Yes |
| 90 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 8040 | 0.072 | 0.2844 | Yes |
| 91 | LYAR | LYAR Entrez, Source | - | 8134 | 0.072 | 0.2846 | Yes |
| 92 | RBBP8 | RBBP8 Entrez, Source | retinoblastoma binding protein 8 | 8178 | 0.071 | 0.2867 | Yes |
| 93 | TMPO | TMPO Entrez, Source | thymopoietin | 8207 | 0.070 | 0.2894 | Yes |
| 94 | UHRF1 | UHRF1 Entrez, Source | ubiquitin-like, containing PHD and RING finger domains, 1 | 8213 | 0.070 | 0.2930 | Yes |
| 95 | RBL1 | RBL1 Entrez, Source | retinoblastoma-like 1 (p107) | 8225 | 0.070 | 0.2963 | Yes |
| 96 | MELK | MELK Entrez, Source | maternal embryonic leucine zipper kinase | 8236 | 0.069 | 0.2996 | Yes |
| 97 | LSM2 | LSM2 Entrez, Source | LSM2 homolog, U6 small nuclear RNA associated (S. cerevisiae) | 8239 | 0.069 | 0.3033 | Yes |
| 98 | ECT2 | ECT2 Entrez, Source | epithelial cell transforming sequence 2 oncogene | 8270 | 0.069 | 0.3058 | Yes |
| 99 | LARP7 | LARP7 Entrez, Source | La ribonucleoprotein domain family, member 7 | 8338 | 0.068 | 0.3068 | Yes |
| 100 | SMC2 | SMC2 Entrez, Source | structural maintenance of chromosomes 2 | 8352 | 0.067 | 0.3099 | Yes |
| 101 | RIF1 | RIF1 Entrez, Source | RAP1 interacting factor homolog (yeast) | 8355 | 0.067 | 0.3135 | Yes |
| 102 | ERI1 | 8452 | 0.065 | 0.3132 | Yes | ||
| 103 | FBLN2 | FBLN2 Entrez, Source | fibulin 2 | 8467 | 0.065 | 0.3162 | Yes |
| 104 | TOPBP1 | TOPBP1 Entrez, Source | topoisomerase (DNA) II binding protein 1 | 8478 | 0.065 | 0.3192 | Yes |
| 105 | ZWILCH | ZWILCH Entrez, Source | Zwilch, kinetochore associated, homolog (Drosophila) | 8495 | 0.064 | 0.3221 | Yes |
| 106 | UCK2 | UCK2 Entrez, Source | uridine-cytidine kinase 2 | 8505 | 0.064 | 0.3252 | Yes |
| 107 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 8541 | 0.063 | 0.3272 | Yes |
| 108 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 8553 | 0.063 | 0.3302 | Yes |
| 109 | DBF4 | DBF4 Entrez, Source | DBF4 homolog (S. cerevisiae) | 8568 | 0.063 | 0.3330 | Yes |
| 110 | DNAJC2 | 8580 | 0.062 | 0.3359 | Yes | ||
| 111 | MCM4 | MCM4 Entrez, Source | MCM4 minichromosome maintenance deficient 4 (S. cerevisiae) | 8582 | 0.062 | 0.3392 | Yes |
| 112 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 8593 | 0.062 | 0.3421 | Yes |
| 113 | PBK | PBK Entrez, Source | PDZ binding kinase | 8608 | 0.062 | 0.3449 | Yes |
| 114 | EZH2 | EZH2 Entrez, Source | enhancer of zeste homolog 2 (Drosophila) | 8626 | 0.061 | 0.3475 | Yes |
| 115 | ELOVL2 | ELOVL2 Entrez, Source | elongation of very long chain fatty acids (FEN1/Elo2, SUR4/Elo3, yeast)-like 2 | 8638 | 0.061 | 0.3504 | Yes |
| 116 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 8646 | 0.061 | 0.3534 | Yes |
| 117 | WDHD1 | WDHD1 Entrez, Source | WD repeat and HMG-box DNA binding protein 1 | 8662 | 0.061 | 0.3561 | Yes |
| 118 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 8688 | 0.060 | 0.3584 | Yes |
| 119 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 8912 | 0.060 | 0.3529 | Yes |
| 120 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 8928 | 0.059 | 0.3555 | Yes |
| 121 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 8951 | 0.059 | 0.3578 | Yes |
| 122 | SLC12A2 | SLC12A2 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 2 | 8967 | 0.059 | 0.3604 | Yes |
| 123 | MCM5 | MCM5 Entrez, Source | MCM5 minichromosome maintenance deficient 5, cell division cycle 46 (S. cerevisiae) | 9075 | 0.057 | 0.3593 | Yes |
| 124 | DCK | DCK Entrez, Source | deoxycytidine kinase | 9076 | 0.057 | 0.3623 | Yes |
| 125 | CASQ1 | CASQ1 Entrez, Source | calsequestrin 1 (fast-twitch, skeletal muscle) | 9098 | 0.057 | 0.3645 | Yes |
| 126 | RRM1 | RRM1 Entrez, Source | ribonucleotide reductase M1 polypeptide | 9137 | 0.056 | 0.3661 | Yes |
| 127 | AK4 | 9167 | 0.055 | 0.3679 | Yes | ||
| 128 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 9173 | 0.055 | 0.3707 | Yes |
| 129 | POLE | POLE Entrez, Source | polymerase (DNA directed), epsilon | 9191 | 0.055 | 0.3729 | Yes |
| 130 | TACC3 | TACC3 Entrez, Source | transforming, acidic coiled-coil containing protein 3 | 9196 | 0.055 | 0.3757 | Yes |
| 131 | LTV1 | LTV1 Entrez, Source | LTV1 homolog (S. cerevisiae) | 9252 | 0.055 | 0.3765 | Yes |
| 132 | RACGAP1 | RACGAP1 Entrez, Source | Rac GTPase activating protein 1 | 9253 | 0.055 | 0.3795 | Yes |
| 133 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9270 | 0.054 | 0.3818 | Yes |
| 134 | UTP6 | UTP6 Entrez, Source | UTP6, small subunit (SSU) processome component, homolog (yeast) | 9326 | 0.053 | 0.3825 | Yes |
| 135 | DDX21 | DDX21 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 21 | 9336 | 0.053 | 0.3850 | Yes |
| 136 | IVNS1ABP | IVNS1ABP Entrez, Source | influenza virus NS1A binding protein | 9362 | 0.052 | 0.3868 | Yes |
| 137 | NUP160 | NUP160 Entrez, Source | nucleoporin 160kDa | 9370 | 0.052 | 0.3893 | Yes |
| 138 | FAM54A | FAM54A Entrez, Source | family with sequence similarity 54, member A | 9387 | 0.052 | 0.3915 | Yes |
| 139 | ZBTB12 | ZBTB12 Entrez, Source | zinc finger and BTB domain containing 12 | 9424 | 0.052 | 0.3929 | Yes |
| 140 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 9425 | 0.052 | 0.3957 | Yes |
| 141 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 9453 | 0.051 | 0.3974 | Yes |
| 142 | NUP85 | NUP85 Entrez, Source | nucleoporin 85kDa | 9467 | 0.051 | 0.3996 | Yes |
| 143 | WDR43 | WDR43 Entrez, Source | WD repeat domain 43 | 9517 | 0.050 | 0.4004 | Yes |
| 144 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 9554 | 0.050 | 0.4017 | Yes |
| 145 | DCPS | DCPS Entrez, Source | decapping enzyme, scavenger | 9566 | 0.049 | 0.4039 | Yes |
| 146 | LMNB2 | LMNB2 Entrez, Source | lamin B2 | 9568 | 0.049 | 0.4065 | Yes |
| 147 | MCM3 | MCM3 Entrez, Source | MCM3 minichromosome maintenance deficient 3 (S. cerevisiae) | 9611 | 0.049 | 0.4075 | Yes |
| 148 | KIF11 | KIF11 Entrez, Source | kinesin family member 11 | 9652 | 0.048 | 0.4085 | Yes |
| 149 | POLD2 | POLD2 Entrez, Source | polymerase (DNA directed), delta 2, regulatory subunit 50kDa | 9681 | 0.048 | 0.4100 | Yes |
| 150 | SRSF3 | 9702 | 0.047 | 0.4118 | Yes | ||
| 151 | AURKB | AURKB Entrez, Source | aurora kinase B | 9747 | 0.047 | 0.4126 | Yes |
| 152 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 9757 | 0.047 | 0.4148 | Yes |
| 153 | SRSF7 | 9761 | 0.047 | 0.4172 | Yes | ||
| 154 | SDF2L1 | SDF2L1 Entrez, Source | stromal cell-derived factor 2-like 1 | 9800 | 0.046 | 0.4181 | Yes |
| 155 | RDX | RDX Entrez, Source | radixin | 9856 | 0.046 | 0.4185 | Yes |
| 156 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 9902 | 0.045 | 0.4191 | Yes |
| 157 | NUF2 | 9919 | 0.045 | 0.4209 | Yes | ||
| 158 | FANCI | FANCI Entrez, Source | Fanconi anemia, complementation group I | 9924 | 0.045 | 0.4232 | Yes |
| 159 | HMGXB4 | 9941 | 0.044 | 0.4249 | Yes | ||
| 160 | DEK | DEK Entrez, Source | DEK oncogene (DNA binding) | 9948 | 0.044 | 0.4271 | Yes |
| 161 | POLA2 | POLA2 Entrez, Source | polymerase (DNA directed), alpha 2 (70kD subunit) | 9968 | 0.044 | 0.4287 | Yes |
| 162 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 9993 | 0.044 | 0.4301 | Yes |
| 163 | VARS | VARS Entrez, Source | valyl-tRNA synthetase | 10057 | 0.043 | 0.4300 | Yes |
| 164 | UBR7 | 10059 | 0.043 | 0.4322 | Yes | ||
| 165 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 10084 | 0.042 | 0.4336 | Yes |
| 166 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10086 | 0.042 | 0.4358 | Yes |
| 167 | HMGB2 | HMGB2 Entrez, Source | high-mobility group box 2 | 10107 | 0.042 | 0.4373 | Yes |
| 168 | ORC1 | 10108 | 0.042 | 0.4395 | Yes | ||
| 169 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 10152 | 0.041 | 0.4401 | Yes |
| 170 | CBX5 | CBX5 Entrez, Source | chromobox homolog 5 (HP1 alpha homolog, Drosophila) | 10161 | 0.041 | 0.4420 | Yes |
| 171 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 10197 | 0.041 | 0.4428 | Yes |
| 172 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 10209 | 0.041 | 0.4446 | Yes |
| 173 | DLG4 | DLG4 Entrez, Source | discs, large homolog 4 (Drosophila) | 10224 | 0.040 | 0.4462 | Yes |
| 174 | EPRS | EPRS Entrez, Source | glutamyl-prolyl-tRNA synthetase | 10249 | 0.040 | 0.4474 | Yes |
| 175 | WDR46 | WDR46 Entrez, Source | WD repeat domain 46 | 10293 | 0.040 | 0.4479 | Yes |
| 176 | INCENP | INCENP Entrez, Source | inner centromere protein antigens 135/155kDa | 10322 | 0.039 | 0.4489 | Yes |
| 177 | MYBL2 | MYBL2 Entrez, Source | v-myb myeloblastosis viral oncogene homolog (avian)-like 2 | 10368 | 0.039 | 0.4492 | Yes |
| 178 | NUP155 | NUP155 Entrez, Source | nucleoporin 155kDa | 10377 | 0.038 | 0.4510 | Yes |
| 179 | MSH6 | MSH6 Entrez, Source | mutS homolog 6 (E. coli) | 10395 | 0.038 | 0.4523 | Yes |
| 180 | ASCL1 | ASCL1 Entrez, Source | achaete-scute complex-like 1 (Drosophila) | 10402 | 0.038 | 0.4541 | Yes |
| 181 | TMEM48 | TMEM48 Entrez, Source | transmembrane protein 48 | 10414 | 0.038 | 0.4557 | Yes |
| 182 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 10437 | 0.037 | 0.4569 | Yes |
| 183 | NSUN2 | NSUN2 Entrez, Source | NOL1/NOP2/Sun domain family, member 2 | 10441 | 0.037 | 0.4588 | Yes |
| 184 | NUP43 | NUP43 Entrez, Source | nucleoporin 43kDa | 10445 | 0.037 | 0.4607 | Yes |
| 185 | RFC5 | RFC5 Entrez, Source | replication factor C (activator 1) 5, 36.5kDa | 10501 | 0.037 | 0.4605 | Yes |
| 186 | SETDB1 | SETDB1 Entrez, Source | SET domain, bifurcated 1 | 10516 | 0.036 | 0.4619 | Yes |
| 187 | SRM | SRM Entrez, Source | spermidine synthase | 10574 | 0.036 | 0.4616 | Yes |
| 188 | SKIV2L2 | SKIV2L2 Entrez, Source | superkiller viralicidic activity 2-like 2 (S. cerevisiae) | 10618 | 0.035 | 0.4618 | Yes |
| 189 | GLUL | GLUL Entrez, Source | glutamate-ammonia ligase (glutamine synthetase) | 10645 | 0.035 | 0.4627 | Yes |
| 190 | HNRNPR | 10658 | 0.035 | 0.4641 | Yes | ||
| 191 | TIPARP | TIPARP Entrez, Source | TCDD-inducible poly(ADP-ribose) polymerase | 10666 | 0.034 | 0.4656 | Yes |
| 192 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 10680 | 0.034 | 0.4670 | Yes |
| 193 | IL4I1 | IL4I1 Entrez, Source | interleukin 4 induced 1 | 10713 | 0.034 | 0.4676 | Yes |
| 194 | DNA2 | 10747 | 0.033 | 0.4681 | Yes | ||
| 195 | NCAPD2 | NCAPD2 Entrez, Source | non-SMC condensin I complex, subunit D2 | 10892 | 0.032 | 0.4642 | Yes |
| 196 | INTS7 | INTS7 Entrez, Source | integrator complex subunit 7 | 10895 | 0.032 | 0.4658 | Yes |
| 197 | HUS1 | HUS1 Entrez, Source | HUS1 checkpoint homolog (S. pombe) | 10921 | 0.031 | 0.4665 | Yes |
| 198 | DNAJC21 | 10940 | 0.031 | 0.4674 | Yes | ||
| 199 | CDV3 | CDV3 Entrez, Source | CDV3 homolog (mouse) | 10993 | 0.030 | 0.4671 | Yes |
| 200 | FKBP5 | FKBP5 Entrez, Source | FK506 binding protein 5 | 11027 | 0.030 | 0.4674 | Yes |
| 201 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 11033 | 0.030 | 0.4688 | Yes |
| 202 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 11053 | 0.030 | 0.4697 | Yes |
| 203 | DHX36 | DHX36 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 36 | 11146 | 0.029 | 0.4676 | No |
| 204 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 11167 | 0.029 | 0.4684 | No |
| 205 | RRM2 | RRM2 Entrez, Source | ribonucleotide reductase M2 polypeptide | 11196 | 0.028 | 0.4688 | No |
| 206 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 11214 | 0.028 | 0.4697 | No |
| 207 | HJURP | 11262 | 0.027 | 0.4693 | No | ||
| 208 | WDR77 | WDR77 Entrez, Source | WD repeat domain 77 | 11355 | 0.026 | 0.4671 | No |
| 209 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 11375 | 0.026 | 0.4678 | No |
| 210 | DIS3 | 11406 | 0.026 | 0.4680 | No | ||
| 211 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 11514 | 0.024 | 0.4652 | No |
| 212 | WDR33 | WDR33 Entrez, Source | WD repeat domain 33 | 11600 | 0.024 | 0.4631 | No |
| 213 | GART | GART Entrez, Source | phosphoribosylglycinamide formyltransferase, phosphoribosylglycinamide synthetase, phosphoribosylaminoimidazole synthetase | 11889 | 0.020 | 0.4529 | No |
| 214 | TK1 | TK1 Entrez, Source | thymidine kinase 1, soluble | 11893 | 0.020 | 0.4539 | No |
| 215 | EIF4E | EIF4E Entrez, Source | eukaryotic translation initiation factor 4E | 12026 | 0.018 | 0.4497 | No |
| 216 | HSP90B1 | HSP90B1 Entrez, Source | heat shock protein 90kDa beta (Grp94), member 1 | 12126 | 0.017 | 0.4468 | No |
| 217 | CENPM | CENPM Entrez, Source | centromere protein M | 12220 | 0.016 | 0.4440 | No |
| 218 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 12263 | 0.016 | 0.4432 | No |
| 219 | HYOU1 | HYOU1 Entrez, Source | hypoxia up-regulated 1 | 12328 | 0.015 | 0.4415 | No |
| 220 | ERO1L | ERO1L Entrez, Source | ERO1-like (S. cerevisiae) | 12368 | 0.014 | 0.4408 | No |
| 221 | SHMT1 | SHMT1 Entrez, Source | serine hydroxymethyltransferase 1 (soluble) | 12642 | 0.011 | 0.4307 | No |
| 222 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 12646 | 0.011 | 0.4312 | No |
| 223 | HK2 | HK2 Entrez, Source | hexokinase 2 | 12757 | 0.010 | 0.4274 | No |
| 224 | DHODH | DHODH Entrez, Source | dihydroorotate dehydrogenase | 13199 | 0.004 | 0.4104 | No |
| 225 | PDZD11 | PDZD11 Entrez, Source | PDZ domain containing 11 | 13237 | 0.004 | 0.4092 | No |
| 226 | UMPS | UMPS Entrez, Source | uridine monophosphate synthetase (orotate phosphoribosyl transferase and orotidine-5'-decarboxylase) | 13285 | 0.003 | 0.4075 | No |
| 227 | TNRC6A | TNRC6A Entrez, Source | trinucleotide repeat containing 6A | 13377 | 0.002 | 0.4041 | No |
| 228 | CENPF | CENPF Entrez, Source | centromere protein F, 350/400ka (mitosin) | 13466 | 0.001 | 0.4007 | No |
| 229 | SPAG5 | SPAG5 Entrez, Source | sperm associated antigen 5 | 13814 | -0.003 | 0.3873 | No |
| 230 | IFRD2 | IFRD2 Entrez, Source | interferon-related developmental regulator 2 | 13959 | -0.005 | 0.3820 | No |
| 231 | ME2 | ME2 Entrez, Source | malic enzyme 2, NAD(+)-dependent, mitochondrial | 13965 | -0.005 | 0.3820 | No |
| 232 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 14130 | -0.007 | 0.3760 | No |
| 233 | CDT1 | CDT1 Entrez, Source | chromatin licensing and DNA replication factor 1 | 14143 | -0.007 | 0.3759 | No |
| 234 | CDCA7L | CDCA7L Entrez, Source | cell division cycle associated 7-like | 14347 | -0.010 | 0.3685 | No |
| 235 | RBM14 | RBM14 Entrez, Source | RNA binding motif protein 14 | 14547 | -0.013 | 0.3614 | No |
| 236 | SON | SON Entrez, Source | SON DNA binding protein | 14648 | -0.014 | 0.3583 | No |
| 237 | RAD54L | RAD54L Entrez, Source | RAD54-like (S. cerevisiae) | 14784 | -0.016 | 0.3539 | No |
| 238 | TBL2 | TBL2 Entrez, Source | transducin (beta)-like 2 | 14831 | -0.017 | 0.3530 | No |
| 239 | AATF | AATF Entrez, Source | apoptosis antagonizing transcription factor | 14954 | -0.019 | 0.3493 | No |
| 240 | THRAP3 | THRAP3 Entrez, Source | thyroid hormone receptor associated protein 3 | 15111 | -0.021 | 0.3443 | No |
| 241 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 15118 | -0.021 | 0.3452 | No |
| 242 | MKI67 | MKI67 Entrez, Source | antigen identified by monoclonal antibody Ki-67 | 15149 | -0.022 | 0.3452 | No |
| 243 | GRWD1 | GRWD1 Entrez, Source | glutamate-rich WD repeat containing 1 | 15161 | -0.022 | 0.3460 | No |
| 244 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 15266 | -0.024 | 0.3432 | No |
| 245 | FANCA | FANCA Entrez, Source | Fanconi anemia, complementation group A | 15269 | -0.024 | 0.3444 | No |
| 246 | PEG3 | PEG3 Entrez, Source | paternally expressed 3 | 15338 | -0.024 | 0.3430 | No |
| 247 | RRP12 | 15341 | -0.024 | 0.3442 | No | ||
| 248 | DIMT1 | 15449 | -0.025 | 0.3414 | No | ||
| 249 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 15707 | -0.029 | 0.3330 | No |
| 250 | TIMM9 | TIMM9 Entrez, Source | translocase of inner mitochondrial membrane 9 homolog (yeast) | 15733 | -0.030 | 0.3336 | No |
| 251 | MYO10 | MYO10 Entrez, Source | myosin X | 16053 | -0.035 | 0.3230 | No |
| 252 | FRMD4B | FRMD4B Entrez, Source | FERM domain containing 4B | 16439 | -0.038 | 0.3101 | No |
| 253 | NTN1 | NTN1 Entrez, Source | netrin 1 | 16491 | -0.039 | 0.3102 | No |
| 254 | TCF19 | TCF19 Entrez, Source | transcription factor 19 (SC1) | 16559 | -0.040 | 0.3097 | No |
| 255 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 17335 | -0.055 | 0.2824 | No |
| 256 | PPP1R18 | 17574 | -0.060 | 0.2764 | No | ||
| 257 | ATM | ATM Entrez, Source | ataxia telangiectasia mutated (includes complementation groups A, C and D) | 17577 | -0.060 | 0.2796 | No |
| 258 | SOAT1 | SOAT1 Entrez, Source | sterol O-acyltransferase (acyl-Coenzyme A: cholesterol acyltransferase) 1 | 17645 | -0.062 | 0.2803 | No |
| 259 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 17779 | -0.065 | 0.2786 | No |
| 260 | NRARP | 17858 | -0.067 | 0.2791 | No | ||
| 261 | SLC4A4 | SLC4A4 Entrez, Source | solute carrier family 4, sodium bicarbonate cotransporter, member 4 | 18269 | -0.078 | 0.2673 | No |
| 262 | CCNE1 | CCNE1 Entrez, Source | cyclin E1 | 18646 | -0.089 | 0.2574 | No |
| 263 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 18703 | -0.091 | 0.2601 | No |
| 264 | PDZRN3 | PDZRN3 Entrez, Source | PDZ domain containing RING finger 3 | 20701 | -0.172 | 0.1914 | No |
| 265 | TTPA | TTPA Entrez, Source | tocopherol (alpha) transfer protein (ataxia (Friedreich-like) with vitamin E deficiency) | 21296 | -0.209 | 0.1795 | No |