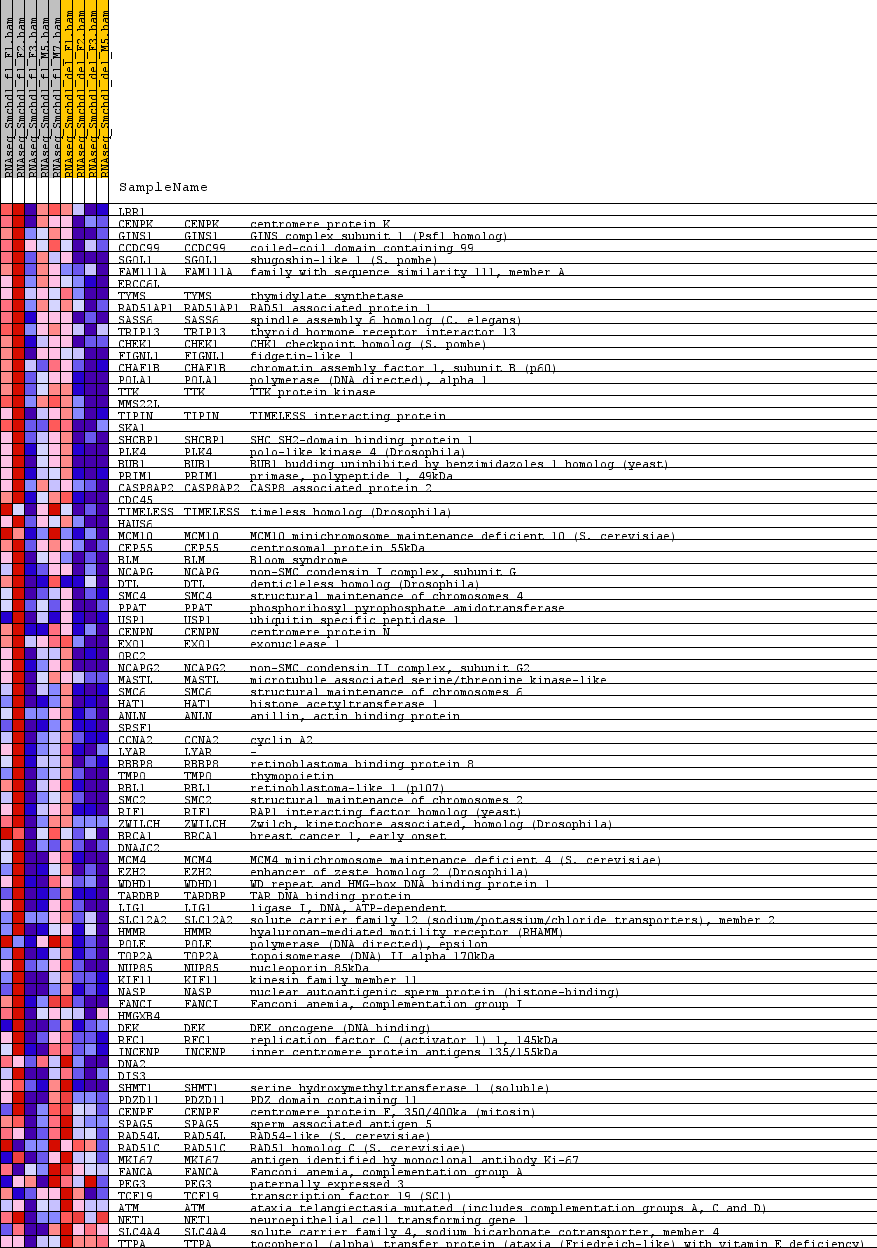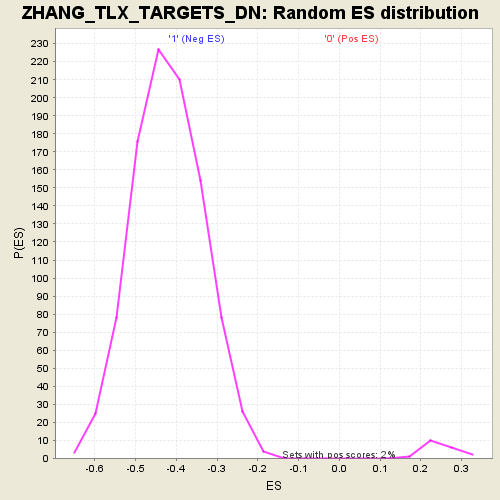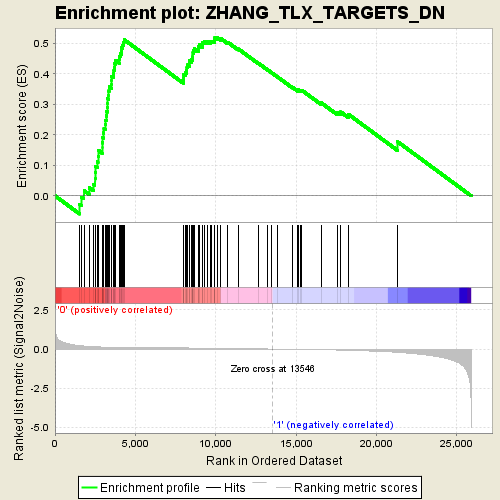
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | SMCHD1_KO_collapsed_to_symbols.SMCHD1_KO.cls #del_versus_fl.SMCHD1_KO.cls #del_versus_fl_repos |
| Phenotype | SMCHD1_KO.cls#del_versus_fl_repos |
| Upregulated in class | 0 |
| GeneSet | ZHANG_TLX_TARGETS_DN |
| Enrichment Score (ES) | 0.51898986 |
| Normalized Enrichment Score (NES) | 2.072519 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.22078115 |
| FWER p-Value | 0.161 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LRR1 | 1534 | 0.225 | -0.0284 | Yes | ||
| 2 | CENPK | CENPK Entrez, Source | centromere protein K | 1652 | 0.210 | -0.0041 | Yes |
| 3 | GINS1 | GINS1 Entrez, Source | GINS complex subunit 1 (Psf1 homolog) | 1800 | 0.195 | 0.0170 | Yes |
| 4 | CCDC99 | CCDC99 Entrez, Source | coiled-coil domain containing 99 | 2125 | 0.165 | 0.0271 | Yes |
| 5 | SGOL1 | SGOL1 Entrez, Source | shugoshin-like 1 (S. pombe) | 2403 | 0.160 | 0.0384 | Yes |
| 6 | FAM111A | FAM111A Entrez, Source | family with sequence similarity 111, member A | 2484 | 0.152 | 0.0562 | Yes |
| 7 | ERCC6L | 2491 | 0.151 | 0.0768 | Yes | ||
| 8 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 2505 | 0.150 | 0.0970 | Yes |
| 9 | RAD51AP1 | RAD51AP1 Entrez, Source | RAD51 associated protein 1 | 2626 | 0.146 | 0.1125 | Yes |
| 10 | SASS6 | SASS6 Entrez, Source | spindle assembly 6 homolog (C. elegans) | 2674 | 0.143 | 0.1303 | Yes |
| 11 | TRIP13 | TRIP13 Entrez, Source | thyroid hormone receptor interactor 13 | 2702 | 0.141 | 0.1487 | Yes |
| 12 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 2928 | 0.126 | 0.1574 | Yes |
| 13 | FIGNL1 | FIGNL1 Entrez, Source | fidgetin-like 1 | 2930 | 0.126 | 0.1747 | Yes |
| 14 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 2971 | 0.124 | 0.1902 | Yes |
| 15 | POLA1 | POLA1 Entrez, Source | polymerase (DNA directed), alpha 1 | 2993 | 0.123 | 0.2062 | Yes |
| 16 | TTK | TTK Entrez, Source | TTK protein kinase | 3042 | 0.119 | 0.2207 | Yes |
| 17 | MMS22L | 3118 | 0.115 | 0.2336 | Yes | ||
| 18 | TIPIN | TIPIN Entrez, Source | TIMELESS interacting protein | 3139 | 0.114 | 0.2485 | Yes |
| 19 | SKA1 | 3180 | 0.111 | 0.2622 | Yes | ||
| 20 | SHCBP1 | SHCBP1 Entrez, Source | SHC SH2-domain binding protein 1 | 3202 | 0.110 | 0.2765 | Yes |
| 21 | PLK4 | PLK4 Entrez, Source | polo-like kinase 4 (Drosophila) | 3251 | 0.108 | 0.2895 | Yes |
| 22 | BUB1 | BUB1 Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) | 3261 | 0.107 | 0.3039 | Yes |
| 23 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 3268 | 0.107 | 0.3184 | Yes |
| 24 | CASP8AP2 | CASP8AP2 Entrez, Source | CASP8 associated protein 2 | 3342 | 0.103 | 0.3298 | Yes |
| 25 | CDC45 | 3350 | 0.103 | 0.3437 | Yes | ||
| 26 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 3361 | 0.102 | 0.3574 | Yes |
| 27 | HAUS6 | 3501 | 0.097 | 0.3654 | Yes | ||
| 28 | MCM10 | MCM10 Entrez, Source | MCM10 minichromosome maintenance deficient 10 (S. cerevisiae) | 3508 | 0.097 | 0.3785 | Yes |
| 29 | CEP55 | CEP55 Entrez, Source | centrosomal protein 55kDa | 3514 | 0.097 | 0.3916 | Yes |
| 30 | BLM | BLM Entrez, Source | Bloom syndrome | 3626 | 0.092 | 0.3999 | Yes |
| 31 | NCAPG | NCAPG Entrez, Source | non-SMC condensin I complex, subunit G | 3663 | 0.090 | 0.4110 | Yes |
| 32 | DTL | DTL Entrez, Source | denticleless homolog (Drosophila) | 3696 | 0.089 | 0.4220 | Yes |
| 33 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 3717 | 0.088 | 0.4334 | Yes |
| 34 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 3759 | 0.087 | 0.4438 | Yes |
| 35 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 3988 | 0.086 | 0.4467 | Yes |
| 36 | CENPN | CENPN Entrez, Source | centromere protein N | 4004 | 0.085 | 0.4579 | Yes |
| 37 | EXO1 | EXO1 Entrez, Source | exonuclease 1 | 4079 | 0.083 | 0.4664 | Yes |
| 38 | ORC2 | 4143 | 0.080 | 0.4750 | Yes | ||
| 39 | NCAPG2 | NCAPG2 Entrez, Source | non-SMC condensin II complex, subunit G2 | 4155 | 0.080 | 0.4856 | Yes |
| 40 | MASTL | MASTL Entrez, Source | microtubule associated serine/threonine kinase-like | 4210 | 0.078 | 0.4942 | Yes |
| 41 | SMC6 | SMC6 Entrez, Source | structural maintenance of chromosomes 6 | 4260 | 0.077 | 0.5029 | Yes |
| 42 | HAT1 | HAT1 Entrez, Source | histone acetyltransferase 1 | 4293 | 0.075 | 0.5120 | Yes |
| 43 | ANLN | ANLN Entrez, Source | anillin, actin binding protein | 7973 | 0.074 | 0.3797 | Yes |
| 44 | SRSF1 | 7992 | 0.073 | 0.3890 | Yes | ||
| 45 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 7999 | 0.073 | 0.3989 | Yes |
| 46 | LYAR | LYAR Entrez, Source | - | 8134 | 0.072 | 0.4035 | Yes |
| 47 | RBBP8 | RBBP8 Entrez, Source | retinoblastoma binding protein 8 | 8178 | 0.071 | 0.4116 | Yes |
| 48 | TMPO | TMPO Entrez, Source | thymopoietin | 8207 | 0.070 | 0.4201 | Yes |
| 49 | RBL1 | RBL1 Entrez, Source | retinoblastoma-like 1 (p107) | 8225 | 0.070 | 0.4290 | Yes |
| 50 | SMC2 | SMC2 Entrez, Source | structural maintenance of chromosomes 2 | 8352 | 0.067 | 0.4334 | Yes |
| 51 | RIF1 | RIF1 Entrez, Source | RAP1 interacting factor homolog (yeast) | 8355 | 0.067 | 0.4425 | Yes |
| 52 | ZWILCH | ZWILCH Entrez, Source | Zwilch, kinetochore associated, homolog (Drosophila) | 8495 | 0.064 | 0.4459 | Yes |
| 53 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 8541 | 0.063 | 0.4529 | Yes |
| 54 | DNAJC2 | 8580 | 0.062 | 0.4600 | Yes | ||
| 55 | MCM4 | MCM4 Entrez, Source | MCM4 minichromosome maintenance deficient 4 (S. cerevisiae) | 8582 | 0.062 | 0.4685 | Yes |
| 56 | EZH2 | EZH2 Entrez, Source | enhancer of zeste homolog 2 (Drosophila) | 8626 | 0.061 | 0.4753 | Yes |
| 57 | WDHD1 | WDHD1 Entrez, Source | WD repeat and HMG-box DNA binding protein 1 | 8662 | 0.061 | 0.4823 | Yes |
| 58 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 8912 | 0.060 | 0.4809 | Yes |
| 59 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 8928 | 0.059 | 0.4884 | Yes |
| 60 | SLC12A2 | SLC12A2 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 2 | 8967 | 0.059 | 0.4950 | Yes |
| 61 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 9173 | 0.055 | 0.4947 | Yes |
| 62 | POLE | POLE Entrez, Source | polymerase (DNA directed), epsilon | 9191 | 0.055 | 0.5015 | Yes |
| 63 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 9270 | 0.054 | 0.5060 | Yes |
| 64 | NUP85 | NUP85 Entrez, Source | nucleoporin 85kDa | 9467 | 0.051 | 0.5054 | Yes |
| 65 | KIF11 | KIF11 Entrez, Source | kinesin family member 11 | 9652 | 0.048 | 0.5049 | Yes |
| 66 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 9757 | 0.047 | 0.5073 | Yes |
| 67 | FANCI | FANCI Entrez, Source | Fanconi anemia, complementation group I | 9924 | 0.045 | 0.5070 | Yes |
| 68 | HMGXB4 | 9941 | 0.044 | 0.5125 | Yes | ||
| 69 | DEK | DEK Entrez, Source | DEK oncogene (DNA binding) | 9948 | 0.044 | 0.5184 | Yes |
| 70 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 10084 | 0.042 | 0.5190 | Yes |
| 71 | INCENP | INCENP Entrez, Source | inner centromere protein antigens 135/155kDa | 10322 | 0.039 | 0.5152 | No |
| 72 | DNA2 | 10747 | 0.033 | 0.5034 | No | ||
| 73 | DIS3 | 11406 | 0.026 | 0.4814 | No | ||
| 74 | SHMT1 | SHMT1 Entrez, Source | serine hydroxymethyltransferase 1 (soluble) | 12642 | 0.011 | 0.4351 | No |
| 75 | PDZD11 | PDZD11 Entrez, Source | PDZ domain containing 11 | 13237 | 0.004 | 0.4126 | No |
| 76 | CENPF | CENPF Entrez, Source | centromere protein F, 350/400ka (mitosin) | 13466 | 0.001 | 0.4039 | No |
| 77 | SPAG5 | SPAG5 Entrez, Source | sperm associated antigen 5 | 13814 | -0.003 | 0.3909 | No |
| 78 | RAD54L | RAD54L Entrez, Source | RAD54-like (S. cerevisiae) | 14784 | -0.016 | 0.3556 | No |
| 79 | RAD51C | RAD51C Entrez, Source | RAD51 homolog C (S. cerevisiae) | 15118 | -0.021 | 0.3456 | No |
| 80 | MKI67 | MKI67 Entrez, Source | antigen identified by monoclonal antibody Ki-67 | 15149 | -0.022 | 0.3474 | No |
| 81 | FANCA | FANCA Entrez, Source | Fanconi anemia, complementation group A | 15269 | -0.024 | 0.3461 | No |
| 82 | PEG3 | PEG3 Entrez, Source | paternally expressed 3 | 15338 | -0.024 | 0.3467 | No |
| 83 | TCF19 | TCF19 Entrez, Source | transcription factor 19 (SC1) | 16559 | -0.040 | 0.3049 | No |
| 84 | ATM | ATM Entrez, Source | ataxia telangiectasia mutated (includes complementation groups A, C and D) | 17577 | -0.060 | 0.2738 | No |
| 85 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 17779 | -0.065 | 0.2750 | No |
| 86 | SLC4A4 | SLC4A4 Entrez, Source | solute carrier family 4, sodium bicarbonate cotransporter, member 4 | 18269 | -0.078 | 0.2667 | No |
| 87 | TTPA | TTPA Entrez, Source | tocopherol (alpha) transfer protein (ataxia (Friedreich-like) with vitamin E deficiency) | 21296 | -0.209 | 0.1783 | No |