| p-value: | 1e-15 |
| log p-value: | -3.525e+01 |
| Information Content per bp: | 1.700 |
| Number of Target Sequences with motif | 314.0 |
| Percentage of Target Sequences with motif | 21.30% |
| Number of Background Sequences with motif | 6618.7 |
| Percentage of Background Sequences with motif | 13.62% |
| Average Position of motif in Targets | 3145.9 +/- 1735.1bp |
| Average Position of motif in Background | 3025.9 +/- 1763.7bp |
| Strand Bias (log2 ratio + to - strand density) | 0.4 |
| Multiplicity (# of sites on avg that occur together) | 1.13 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
PB0203.1_Zfp691_2/Jaspar
| Match Rank: | 1 |
| Score: | 0.67
| | Offset: | -6
| | Orientation: | reverse strand |
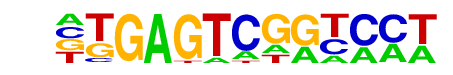
| Alignment: | ------TGAGTCGGTCCT
NTNNNAGGAGTCTCNTN- |
|


|
|
MA0099.2_AP1/Jaspar
| Match Rank: | 2 |
| Score: | 0.67
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TGAGTCGGTCCT
TGAGTCA----- |
|


|
|
AP-1(bZIP)/ThioMac-PU.1-ChIP-Seq(GSE21512)/Homer
| Match Rank: | 3 |
| Score: | 0.64
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --TGAGTCGGTCCT
GATGAGTCAT---- |
|
|
|
Bach2(bZIP)/OCILy7-Bach2-ChIP-Seq(GSE44420)/Homer
| Match Rank: | 4 |
| Score: | 0.64
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TGAGTCGGTCCT
TGACTCAGCA-- |
|


|
|
MA0099.1_Fos/Jaspar
| Match Rank: | 5 |
| Score: | 0.63
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -TGAGTCGGTCCT
GTGAGTCA----- |
|

|
|
Jun-AP1(bZIP)/K562-cJun-ChIP-Seq/Homer
| Match Rank: | 6 |
| Score: | 0.63
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --TGAGTCGGTCCT
NATGACTCATNN-- |
|

|
|
MafK(bZIP)/C2C12-MafK-ChIP-Seq(GSE36030)/Homer
| Match Rank: | 7 |
| Score: | 0.60
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---TGAGTCGGTCCT
TGCTGASTCAGC--- |
|


|
|
Nkx2.1(Homeobox)/H3122-NKX2.1-ChIP-Seq(GSE39998)/Homer
| Match Rank: | 8 |
| Score: | 0.60
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TGAGTCGGTCCT
TTGAGTGG----- |
|

|
|
MafA(bZIP)/Islet-MafA-ChIP-Seq(GSE30298)/Homer
| Match Rank: | 9 |
| Score: | 0.59
| | Offset: | 0
| | Orientation: | reverse strand |
| Alignment: | TGAGTCGGTCCT
TGAGTCAGCA-- |
|


|
|
Bach1(bZIP)/K562-Bach1-ChIP-Seq(GSE31477)/Homer
| Match Rank: | 10 |
| Score: | 0.59
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TGAGTCGGTCCT--
ATGACTCAGCANWWT |
|


|
|





















