| p-value: | 1e-11 |
| log p-value: | -2.719e+01 |
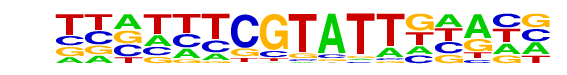
| Information Content per bp: | 1.498 |
| Number of Target Sequences with motif | 690.0 |
| Percentage of Target Sequences with motif | 46.81% |
| Number of Background Sequences with motif | 18403.3 |
| Percentage of Background Sequences with motif | 37.86% |
| Average Position of motif in Targets | 3142.8 +/- 1707.1bp |
| Average Position of motif in Background | 3027.0 +/- 1751.9bp |
| Strand Bias (log2 ratio + to - strand density) | 0.1 |
| Multiplicity (# of sites on avg that occur together) | 1.64 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
PB0187.1_Tcf7_2/Jaspar
| Match Rank: | 1 |
| Score: | 0.74
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --GTATTYTAAA---
CCGTATTATAAACAA |
|


|
|
PB0174.1_Sox30_2/Jaspar
| Match Rank: | 2 |
| Score: | 0.65
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --GTATTYTAAA----
NCGTATTATAATCNTA |
|


|
|
PB0106.1_Arid5a_2/Jaspar
| Match Rank: | 3 |
| Score: | 0.64
| | Offset: | -7
| | Orientation: | reverse strand |
| Alignment: | -------GTATTYTAAA
TNNTTTCGTATTNNANN |
|

|
|
PB0129.1_Glis2_2/Jaspar
| Match Rank: | 4 |
| Score: | 0.63
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -GTATTYTAAA---
AATATTAATAAAGA |
|


|
|
PB0152.1_Nkx3-1_2/Jaspar
| Match Rank: | 5 |
| Score: | 0.61
| | Offset: | -7
| | Orientation: | forward strand |
| Alignment: | -------GTATTYTAAA
ACTCCAAGTACTTGGAA |
|


|
|
MA0032.1_FOXC1/Jaspar
| Match Rank: | 6 |
| Score: | 0.59
| | Offset: | -5
| | Orientation: | forward strand |
| Alignment: | -----GTATTYTAAA
GGTAAGTA------- |
|


|
|
PB0079.1_Sry_1/Jaspar
| Match Rank: | 7 |
| Score: | 0.58
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --GTATTYTAAA----
NANTATTATAATTNNN |
|


|
|
PH0116.1_Nkx2-9/Jaspar
| Match Rank: | 8 |
| Score: | 0.58
| | Offset: | -7
| | Orientation: | reverse strand |
| Alignment: | -------GTATTYTAAA
NATTTAAGTACTTNAAA |
|


|
|
PH0117.1_Nkx3-1/Jaspar
| Match Rank: | 9 |
| Score: | 0.57
| | Offset: | -6
| | Orientation: | forward strand |
| Alignment: | ------GTATTYTAAA-
TACTAAGTACTTAAATG |
|


|
|
CHR/Cell-Cycle-Exp/Homer
| Match Rank: | 10 |
| Score: | 0.56
| | Offset: | -1
| | Orientation: | forward strand |
| Alignment: | -GTATTYTAAA
CGGTTTCAAA- |
|


|
|





















