| p-value: | 1e-12 |
| log p-value: | -2.916e+01 |
| Information Content per bp: | 1.773 |
| Number of Target Sequences with motif | 98.0 |
| Percentage of Target Sequences with motif | 8.17% |
| Number of Background Sequences with motif | 1750.4 |
| Percentage of Background Sequences with motif | 3.62% |
| Average Position of motif in Targets | 2801.4 +/- 1823.4bp |
| Average Position of motif in Background | 2982.6 +/- 1817.9bp |
| Strand Bias (log2 ratio + to - strand density) | -0.4 |
| Multiplicity (# of sites on avg that occur together) | 1.18 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
PB0045.1_Myb_1/Jaspar
| Match Rank: | 1 |
| Score: | 0.72
| | Offset: | -1
| | Orientation: | reverse strand |
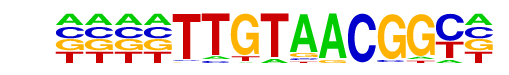
| Alignment: | -TTGTAACGGC------
NNNNTAACGGTTNNNAN |
|


|
|
PB0046.1_Mybl1_1/Jaspar
| Match Rank: | 2 |
| Score: | 0.71
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -TTGTAACGGC------
NNANTAACGGTTNNNAN |
|


|
|
BMYB(HTH)/Hela-BMYB-ChIPSeq(GSE27030)/Homer
| Match Rank: | 3 |
| Score: | 0.68
| | Offset: | 2
| | Orientation: | forward strand |
| Alignment: | TTGTAACGGC--
--NHAACBGYYV |
|


|
|
MYB(HTH)/ERMYB-Myb-ChIPSeq(GSE22095)/Homer
| Match Rank: | 4 |
| Score: | 0.66
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | TTGTAACGGC-
---YAACBGCC |
|


|
|
PB0056.1_Rfxdc2_1/Jaspar
| Match Rank: | 5 |
| Score: | 0.60
| | Offset: | -5
| | Orientation: | forward strand |
| Alignment: | -----TTGTAACGGC
CCGCATAGCAACGGA |
|


|
|
PB0194.1_Zbtb12_2/Jaspar
| Match Rank: | 6 |
| Score: | 0.59
| | Offset: | -4
| | Orientation: | reverse strand |
| Alignment: | ----TTGTAACGGC-
AGNGTTCTAATGANN |
|

|
|
PAX3:FKHR-fusion(Paired/Homeobox)/Rh4-PAX3:FKHR-ChIP-Seq/Homer
| Match Rank: | 7 |
| Score: | 0.58
| | Offset: | -5
| | Orientation: | reverse strand |
| Alignment: | -----TTGTAACGGC
NNAATTAGTCACGGT |
|


|
|
MA0100.1_Myb/Jaspar
| Match Rank: | 8 |
| Score: | 0.58
| | Offset: | 3
| | Orientation: | reverse strand |
| Alignment: | TTGTAACGGC-
---CAACNGCC |
|


|
|
Sox6(HMG)/Myotubes-Sox6-ChIP-Seq(GSE32627)/Homer
| Match Rank: | 9 |
| Score: | 0.57
| | Offset: | -3
| | Orientation: | forward strand |
| Alignment: | ---TTGTAACGGC
CCATTGTTNY--- |
|


|
|
PB0109.1_Bbx_2/Jaspar
| Match Rank: | 10 |
| Score: | 0.57
| | Offset: | -5
| | Orientation: | reverse strand |
| Alignment: | -----TTGTAACGGC--
NNNNCTGTTAACNNTNN |
|


|
|





















