| p-value: | 1e-3 |
| log p-value: | -8.794e+00 |
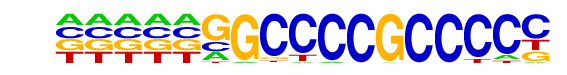
| Information Content per bp: | 1.530 |
| Number of Target Sequences with motif | 521.0 |
| Percentage of Target Sequences with motif | 43.45% |
| Number of Background Sequences with motif | 18518.2 |
| Percentage of Background Sequences with motif | 38.30% |
| Average Position of motif in Targets | 2985.8 +/- 1636.9bp |
| Average Position of motif in Background | 3049.5 +/- 1799.7bp |
| Strand Bias (log2 ratio + to - strand density) | 0.3 |
| Multiplicity (# of sites on avg that occur together) | 1.53 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
PB0052.1_Plagl1_1/Jaspar
| Match Rank: | 1 |
| Score: | 0.67
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --ATGGGAGCCCCG--
TTGGGGGCGCCCCTAG |
|


|
|
POL013.1_MED-1/Jaspar
| Match Rank: | 2 |
| Score: | 0.64
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | ATGGGAGCCCCG
--CGGAGC---- |
|


|
|
Znf263(Zf)/K562-Znf263-ChIP-Seq/Homer
| Match Rank: | 3 |
| Score: | 0.63
| | Offset: | 2
| | Orientation: | reverse strand |
| Alignment: | ATGGGAGCCCCG
--GGGAGGACNG |
|


|
|
PB0099.1_Zfp691_1/Jaspar
| Match Rank: | 4 |
| Score: | 0.55
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -ATGGGAGCCCCG----
NNNNTGAGCACTGTNNG |
|


|
|
MA0105.1_NFKB1/Jaspar
| Match Rank: | 5 |
| Score: | 0.54
| | Offset: | 1
| | Orientation: | forward strand |
| Alignment: | ATGGGAGCCCCG
-GGGGATTCCCC |
|


|
|
PB0156.1_Plagl1_2/Jaspar
| Match Rank: | 6 |
| Score: | 0.54
| | Offset: | -2
| | Orientation: | reverse strand |
| Alignment: | --ATGGGAGCCCCG---
NNNNGGTACCCCCCANN |
|


|
|
PB0203.1_Zfp691_2/Jaspar
| Match Rank: | 7 |
| Score: | 0.54
| | Offset: | -3
| | Orientation: | reverse strand |
| Alignment: | ---ATGGGAGCCCCG--
NTNNNAGGAGTCTCNTN |
|


|
|
POL010.1_DCE_S_III/Jaspar
| Match Rank: | 8 |
| Score: | 0.53
| | Offset: | 4
| | Orientation: | forward strand |
| Alignment: | ATGGGAGCCCCG
----CAGCC--- |
|


|
|
MA0095.1_YY1/Jaspar
| Match Rank: | 9 |
| Score: | 0.53
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -ATGGGAGCCCCG
GATGGC------- |
|


|
|
Sp1(Zf)/Promoter/Homer
| Match Rank: | 10 |
| Score: | 0.52
| | Offset: | 5
| | Orientation: | forward strand |
| Alignment: | ATGGGAGCCCCG-----
-----GGCCCCGCCCCC |
|

|
|





















