| p-value: | 1e-2 |
| log p-value: | -6.076e+00 |
| Information Content per bp: | 1.942 |
| Number of Target Sequences with motif | 55.0 |
| Percentage of Target Sequences with motif | 4.59% |
| Number of Background Sequences with motif | 1475.2 |
| Percentage of Background Sequences with motif | 3.05% |
| Average Position of motif in Targets | 2739.0 +/- 1143.8bp |
| Average Position of motif in Background | 3094.8 +/- 1806.4bp |
| Strand Bias (log2 ratio + to - strand density) | -1.0 |
| Multiplicity (# of sites on avg that occur together) | 2.02 |
| Motif File: | file (matrix)
reverse opposite |
| PDF Format Logos: | forward logo
reverse opposite |
MA0003.1_TFAP2A/Jaspar
| Match Rank: | 1 |
| Score: | 0.77
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | AGCCCGGGGC
-CCCNNNGGC |
|


|
|
PB0088.1_Tcfap2e_1/Jaspar
| Match Rank: | 2 |
| Score: | 0.74
| | Offset: | -2
| | Orientation: | forward strand |
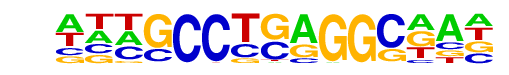
| Alignment: | --AGCCCGGGGC---
ATTGCCTGAGGCAAT |
|

|
|
PB0190.1_Tcfap2b_2/Jaspar
| Match Rank: | 3 |
| Score: | 0.70
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --AGCCCGGGGC---
ATTGCCTCAGGCAAT |
|


|
|
PB0087.1_Tcfap2c_1/Jaspar
| Match Rank: | 4 |
| Score: | 0.69
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --AGCCCGGGGC---
ATTGCCTGAGGCGAA |
|


|
|
MA0146.1_Zfx/Jaspar
| Match Rank: | 5 |
| Score: | 0.68
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -AGCCCGGGGC---
CAGGCCNNGGCCNN |
|


|
|
PB0085.1_Tcfap2a_1/Jaspar
| Match Rank: | 6 |
| Score: | 0.67
| | Offset: | -2
| | Orientation: | forward strand |
| Alignment: | --AGCCCGGGGC---
ATTCCCTGAGGGGAA |
|


|
|
AP-2alpha(AP2)/Hela-AP2alpha-ChIP-Seq/Homer
| Match Rank: | 7 |
| Score: | 0.60
| | Offset: | 1
| | Orientation: | reverse strand |
| Alignment: | AGCCCGGGGC---
-GCCTCAGGGCAT |
|


|
|
AP2gamma(AP2)/MCF7-TFAP2c-ChIP-Seq/Homer
| Match Rank: | 8 |
| Score: | 0.60
| | Offset: | -1
| | Orientation: | reverse strand |
| Alignment: | -AGCCCGGGGC----
TGCCCTGAGGGCANN |
|


|
|
MA0104.2_Mycn/Jaspar
| Match Rank: | 9 |
| Score: | 0.59
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | AGCCCGGGGC
CGCACGTGGC |
|


|
|
MA0147.1_Myc/Jaspar
| Match Rank: | 10 |
| Score: | 0.59
| | Offset: | 0
| | Orientation: | forward strand |
| Alignment: | AGCCCGGGGC
CGCACGTGGC |
|


|
|





















