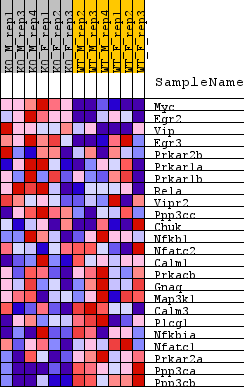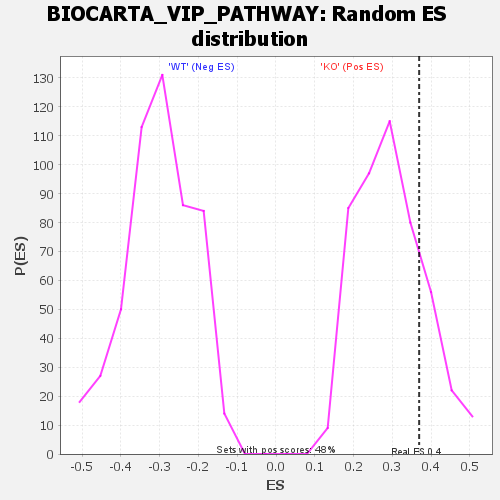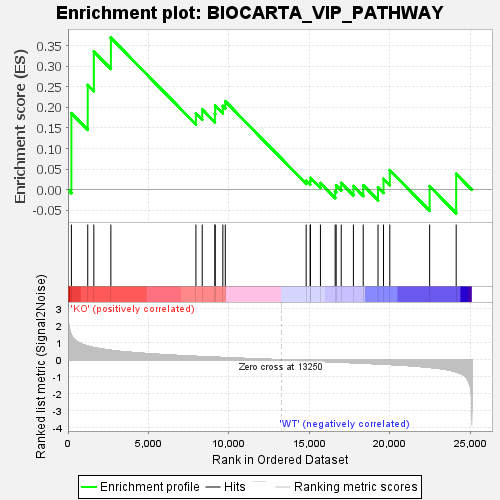
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | KO |
| GeneSet | BIOCARTA_VIP_PATHWAY |
| Enrichment Score (ES) | 0.3691626 |
| Normalized Enrichment Score (NES) | 1.2476418 |
| Nominal p-value | 0.2033543 |
| FDR q-value | 0.6630654 |
| FWER p-Value | 0.902 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Myc | Myc | 211 | 1.433 | 0.1860 | Yes |
| 2 | Egr2 | Egr2 | 1227 | 0.802 | 0.2544 | Yes |
| 3 | Vip | Vip | 1607 | 0.711 | 0.3357 | Yes |
| 4 | Egr3 | Egr3 | 2665 | 0.557 | 0.3692 | Yes |
| 5 | Prkar2b | Prkar2b | 7954 | 0.201 | 0.1856 | No |
| 6 | Prkar1a | Prkar1a | 8345 | 0.185 | 0.1951 | No |
| 7 | Prkar1b | Prkar1b | 9132 | 0.153 | 0.1845 | No |
| 8 | Rela | Rela | 9151 | 0.152 | 0.2045 | No |
| 9 | Vipr2 | Vipr2 | 9628 | 0.133 | 0.2035 | No |
| 10 | Ppp3cc | Ppp3cc | 9779 | 0.127 | 0.2147 | No |
| 11 | Chuk | Chuk | 14808 | -0.056 | 0.0219 | No |
| 12 | Nfkb1 | Nfkb1 | 15056 | -0.065 | 0.0208 | No |
| 13 | Nfatc2 | Nfatc2 | 15074 | -0.066 | 0.0291 | No |
| 14 | Calm1 | Calm1 | 15695 | -0.088 | 0.0163 | No |
| 15 | Prkacb | Prkacb | 16610 | -0.121 | -0.0037 | No |
| 16 | Gnaq | Gnaq | 16673 | -0.124 | 0.0106 | No |
| 17 | Map3k1 | Map3k1 | 16987 | -0.136 | 0.0166 | No |
| 18 | Calm3 | Calm3 | 17745 | -0.166 | 0.0090 | No |
| 19 | Plcg1 | Plcg1 | 18355 | -0.193 | 0.0110 | No |
| 20 | Nfkbia | Nfkbia | 19275 | -0.233 | 0.0060 | No |
| 21 | Nfatc1 | Nfatc1 | 19617 | -0.251 | 0.0264 | No |
| 22 | Prkar2a | Prkar2a | 20008 | -0.271 | 0.0476 | No |
| 23 | Ppp3ca | Ppp3ca | 22485 | -0.441 | 0.0087 | No |
| 24 | Ppp3cb | Ppp3cb | 24135 | -0.706 | 0.0387 | No |