Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | WT |
| GeneSet | REACTOME_ATP_DEPENDENT_CHROMATIN_REMODELERS |
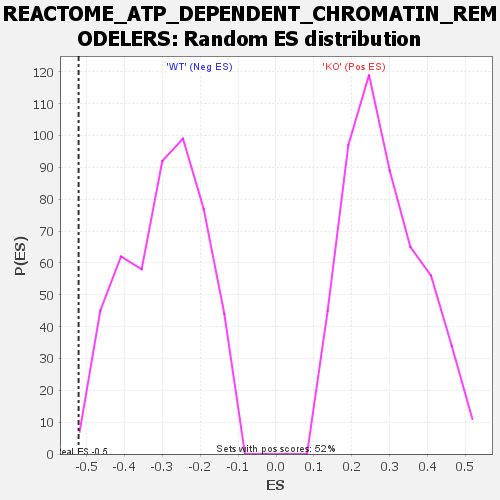
| Enrichment Score (ES) | -0.52121055 |
| Normalized Enrichment Score (NES) | -1.7600522 |
| Nominal p-value | 0.006198347 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.697 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Smarcd3 | Smarcd3 | 3822 | 0.442 | -0.1024 | No |
| 2 | Bcl7c | Bcl7c | 4282 | 0.408 | -0.0747 | No |
| 3 | Smarcc1 | Smarcc1 | 7292 | 0.231 | -0.1685 | No |
| 4 | Bcl11a | Bcl11a | 7389 | 0.227 | -0.1467 | No |
| 5 | Bcl7b | Bcl7b | 8402 | 0.183 | -0.1664 | No |
| 6 | Smarce1 | Smarce1 | 10746 | 0.089 | -0.2497 | No |
| 7 | Pbrm1 | Pbrm1 | 11153 | 0.074 | -0.2576 | No |
| 8 | Bicral | Bicral | 14294 | -0.037 | -0.3786 | No |
| 9 | Ss18l1 | Ss18l1 | 15256 | -0.072 | -0.4087 | No |
| 10 | Smarcd2 | Smarcd2 | 15286 | -0.073 | -0.4015 | No |
| 11 | Smarcb1 | Smarcb1 | 15500 | -0.081 | -0.4009 | No |
| 12 | Arid1a | Arid1a | 15784 | -0.091 | -0.4020 | No |
| 13 | Bicra | Bicra | 16970 | -0.136 | -0.4339 | No |
| 14 | Smarcc2 | Smarcc2 | 17494 | -0.157 | -0.4370 | No |
| 15 | Ss18 | Ss18 | 17733 | -0.166 | -0.4278 | No |
| 16 | Actb | Actb | 18770 | -0.211 | -0.4452 | No |
| 17 | Brd9 | Brd9 | 20080 | -0.275 | -0.4663 | No |
| 18 | Actl6a | Actl6a | 21153 | -0.336 | -0.4710 | No |
| 19 | Brd7 | Brd7 | 22413 | -0.433 | -0.4723 | Yes |
| 20 | Bcl7a | Bcl7a | 22450 | -0.437 | -0.4244 | Yes |
| 21 | Bcl11b | Bcl11b | 22486 | -0.441 | -0.3760 | Yes |
| 22 | Smarcd1 | Smarcd1 | 23281 | -0.532 | -0.3476 | Yes |
| 23 | Dpf3 | Dpf3 | 24668 | -0.934 | -0.2973 | Yes |
| 24 | Smarca4 | Smarca4 | 25102 | -2.785 | 0.0002 | Yes |