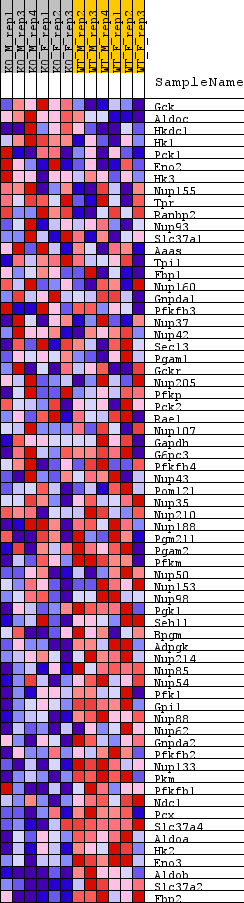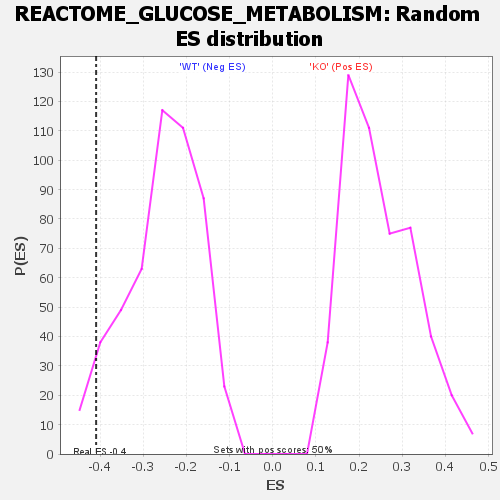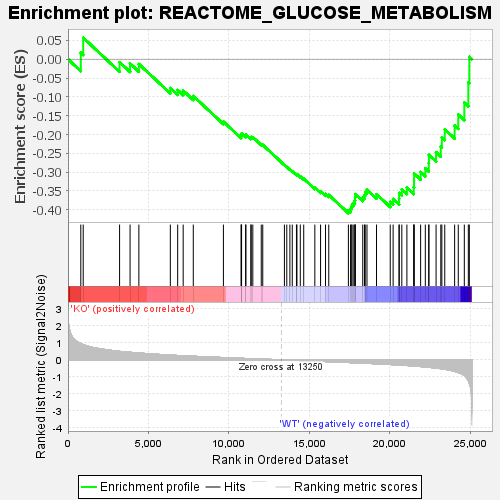
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | WT |
| GeneSet | REACTOME_GLUCOSE_METABOLISM |
| Enrichment Score (ES) | -0.40902764 |
| Normalized Enrichment Score (NES) | -1.6054646 |
| Nominal p-value | 0.035785288 |
| FDR q-value | 0.56555116 |
| FWER p-Value | 0.961 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Gck | Gck | 796 | 0.953 | 0.0170 | No |
| 2 | Aldoc | Aldoc | 946 | 0.892 | 0.0568 | No |
| 3 | Hkdc1 | Hkdc1 | 3207 | 0.497 | -0.0080 | No |
| 4 | Hk1 | Hk1 | 3859 | 0.440 | -0.0114 | No |
| 5 | Pck1 | Pck1 | 4406 | 0.398 | -0.0128 | No |
| 6 | Eno2 | Eno2 | 6362 | 0.278 | -0.0767 | No |
| 7 | Hk3 | Hk3 | 6819 | 0.253 | -0.0819 | No |
| 8 | Nup155 | Nup155 | 7161 | 0.237 | -0.0834 | No |
| 9 | Tpr | Tpr | 7789 | 0.208 | -0.0977 | No |
| 10 | Ranbp2 | Ranbp2 | 9662 | 0.132 | -0.1658 | No |
| 11 | Nup93 | Nup93 | 10761 | 0.088 | -0.2051 | No |
| 12 | Slc37a1 | Slc37a1 | 10783 | 0.087 | -0.2014 | No |
| 13 | Aaas | Aaas | 10792 | 0.087 | -0.1973 | No |
| 14 | Tpi1 | Tpi1 | 11048 | 0.078 | -0.2035 | No |
| 15 | Fbp1 | Fbp1 | 11058 | 0.077 | -0.1999 | No |
| 16 | Nup160 | Nup160 | 11368 | 0.066 | -0.2089 | No |
| 17 | Gnpda1 | Gnpda1 | 11390 | 0.065 | -0.2064 | No |
| 18 | Pfkfb3 | Pfkfb3 | 11491 | 0.062 | -0.2072 | No |
| 19 | Nup37 | Nup37 | 12016 | 0.044 | -0.2259 | No |
| 20 | Nup42 | Nup42 | 12109 | 0.040 | -0.2275 | No |
| 21 | Sec13 | Sec13 | 13455 | -0.007 | -0.2808 | No |
| 22 | Pgam1 | Pgam1 | 13603 | -0.013 | -0.2861 | No |
| 23 | Gckr | Gckr | 13796 | -0.019 | -0.2927 | No |
| 24 | Nup205 | Nup205 | 13944 | -0.025 | -0.2974 | No |
| 25 | Pfkp | Pfkp | 14204 | -0.034 | -0.3059 | No |
| 26 | Pck2 | Pck2 | 14234 | -0.035 | -0.3053 | No |
| 27 | Rae1 | Rae1 | 14442 | -0.042 | -0.3114 | No |
| 28 | Nup107 | Nup107 | 14656 | -0.051 | -0.3173 | No |
| 29 | Gapdh | Gapdh | 15347 | -0.075 | -0.3410 | No |
| 30 | G6pc3 | G6pc3 | 15704 | -0.088 | -0.3507 | No |
| 31 | Pfkfb4 | Pfkfb4 | 16011 | -0.100 | -0.3578 | No |
| 32 | Nup43 | Nup43 | 16213 | -0.107 | -0.3603 | No |
| 33 | Pom121 | Pom121 | 17433 | -0.154 | -0.4012 | Yes |
| 34 | Nup35 | Nup35 | 17569 | -0.159 | -0.3984 | Yes |
| 35 | Nup210 | Nup210 | 17610 | -0.160 | -0.3918 | Yes |
| 36 | Nup188 | Nup188 | 17664 | -0.163 | -0.3855 | Yes |
| 37 | Pgm2l1 | Pgm2l1 | 17772 | -0.168 | -0.3812 | Yes |
| 38 | Pgam2 | Pgam2 | 17835 | -0.171 | -0.3749 | Yes |
| 39 | Pfkm | Pfkm | 17851 | -0.171 | -0.3668 | Yes |
| 40 | Nup50 | Nup50 | 17861 | -0.172 | -0.3583 | Yes |
| 41 | Nup153 | Nup153 | 18326 | -0.192 | -0.3670 | Yes |
| 42 | Nup98 | Nup98 | 18434 | -0.197 | -0.3612 | Yes |
| 43 | Pgk1 | Pgk1 | 18479 | -0.199 | -0.3528 | Yes |
| 44 | Seh1l | Seh1l | 18589 | -0.203 | -0.3467 | Yes |
| 45 | Bpgm | Bpgm | 19182 | -0.229 | -0.3586 | Yes |
| 46 | Adpgk | Adpgk | 20035 | -0.273 | -0.3786 | Yes |
| 47 | Nup214 | Nup214 | 20215 | -0.282 | -0.3713 | Yes |
| 48 | Nup85 | Nup85 | 20585 | -0.303 | -0.3705 | Yes |
| 49 | Nup54 | Nup54 | 20591 | -0.303 | -0.3552 | Yes |
| 50 | Pfkl | Pfkl | 20755 | -0.313 | -0.3457 | Yes |
| 51 | Gpi1 | Gpi1 | 21071 | -0.332 | -0.3412 | Yes |
| 52 | Nup88 | Nup88 | 21498 | -0.360 | -0.3398 | Yes |
| 53 | Nup62 | Nup62 | 21516 | -0.361 | -0.3219 | Yes |
| 54 | Gnpda2 | Gnpda2 | 21518 | -0.362 | -0.3034 | Yes |
| 55 | Pfkfb2 | Pfkfb2 | 21925 | -0.393 | -0.2995 | Yes |
| 56 | Nup133 | Nup133 | 22216 | -0.416 | -0.2898 | Yes |
| 57 | Pkm | Pkm | 22432 | -0.435 | -0.2761 | Yes |
| 58 | Pfkfb1 | Pfkfb1 | 22439 | -0.436 | -0.2540 | Yes |
| 59 | Ndc1 | Ndc1 | 22885 | -0.485 | -0.2469 | Yes |
| 60 | Pcx | Pcx | 23179 | -0.518 | -0.2321 | Yes |
| 61 | Slc37a4 | Slc37a4 | 23244 | -0.528 | -0.2076 | Yes |
| 62 | Aldoa | Aldoa | 23425 | -0.553 | -0.1864 | Yes |
| 63 | Hk2 | Hk2 | 24039 | -0.678 | -0.1762 | Yes |
| 64 | Eno3 | Eno3 | 24265 | -0.744 | -0.1470 | Yes |
| 65 | Aldob | Aldob | 24641 | -0.911 | -0.1153 | Yes |
| 66 | Slc37a2 | Slc37a2 | 24890 | -1.231 | -0.0621 | Yes |
| 67 | Fbp2 | Fbp2 | 24952 | -1.379 | 0.0062 | Yes |