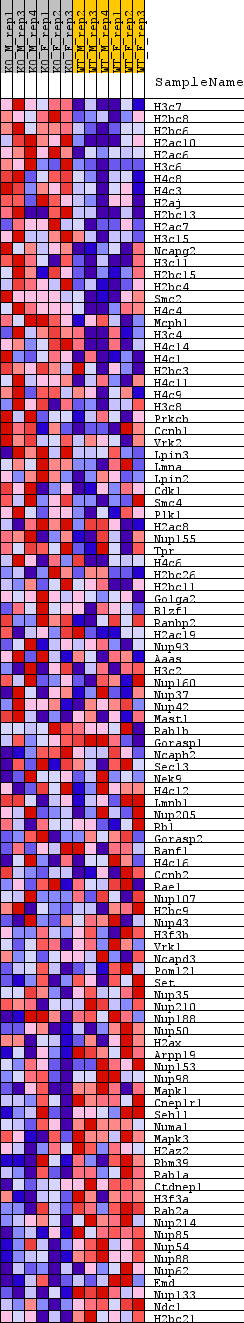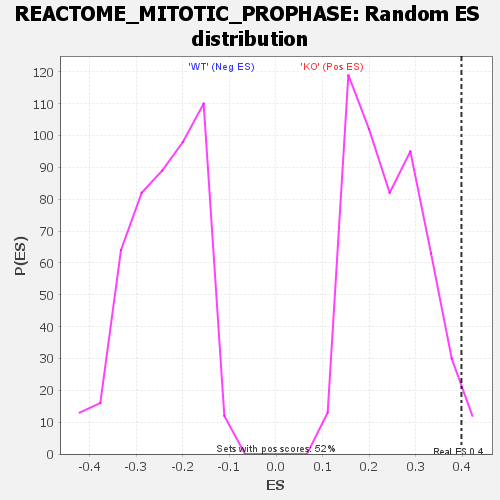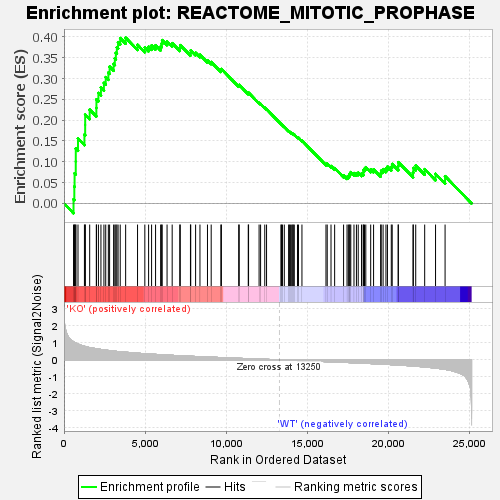
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | KO |
| GeneSet | REACTOME_MITOTIC_PROPHASE |
| Enrichment Score (ES) | 0.3978779 |
| Normalized Enrichment Score (NES) | 1.6429235 |
| Nominal p-value | 0.023255814 |
| FDR q-value | 0.7666419 |
| FWER p-Value | 0.913 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | H3c7 | H3c7 | 587 | 1.051 | 0.0100 | Yes |
| 2 | H2bc8 | H2bc8 | 636 | 1.025 | 0.0407 | Yes |
| 3 | H2bc6 | H2bc6 | 652 | 1.014 | 0.0724 | Yes |
| 4 | H2ac10 | H2ac10 | 724 | 0.979 | 0.1007 | Yes |
| 5 | H2ac6 | H2ac6 | 726 | 0.979 | 0.1318 | Yes |
| 6 | H3c6 | H3c6 | 862 | 0.927 | 0.1559 | Yes |
| 7 | H4c8 | H4c8 | 1258 | 0.795 | 0.1655 | Yes |
| 8 | H4c3 | H4c3 | 1304 | 0.783 | 0.1886 | Yes |
| 9 | H2aj | H2aj | 1305 | 0.782 | 0.2135 | Yes |
| 10 | H2bc13 | H2bc13 | 1582 | 0.716 | 0.2252 | Yes |
| 11 | H2ac7 | H2ac7 | 1993 | 0.649 | 0.2295 | Yes |
| 12 | H3c15 | H3c15 | 1998 | 0.647 | 0.2500 | Yes |
| 13 | Ncapg2 | Ncapg2 | 2119 | 0.630 | 0.2652 | Yes |
| 14 | H3c11 | H3c11 | 2275 | 0.607 | 0.2783 | Yes |
| 15 | H2bc15 | H2bc15 | 2459 | 0.582 | 0.2895 | Yes |
| 16 | H2bc4 | H2bc4 | 2570 | 0.570 | 0.3033 | Yes |
| 17 | Smc2 | Smc2 | 2732 | 0.549 | 0.3143 | Yes |
| 18 | H4c4 | H4c4 | 2809 | 0.540 | 0.3285 | Yes |
| 19 | Mcph1 | Mcph1 | 3055 | 0.513 | 0.3350 | Yes |
| 20 | H3c4 | H3c4 | 3131 | 0.505 | 0.3481 | Yes |
| 21 | H4c14 | H4c14 | 3193 | 0.498 | 0.3615 | Yes |
| 22 | H4c1 | H4c1 | 3255 | 0.492 | 0.3747 | Yes |
| 23 | H2bc3 | H2bc3 | 3330 | 0.486 | 0.3872 | Yes |
| 24 | H4c11 | H4c11 | 3463 | 0.475 | 0.3971 | Yes |
| 25 | H4c9 | H4c9 | 3797 | 0.444 | 0.3979 | Yes |
| 26 | H3c8 | H3c8 | 4530 | 0.389 | 0.3810 | No |
| 27 | Prkcb | Prkcb | 4979 | 0.359 | 0.3745 | No |
| 28 | Ccnb1 | Ccnb1 | 5211 | 0.345 | 0.3762 | No |
| 29 | Vrk2 | Vrk2 | 5395 | 0.334 | 0.3795 | No |
| 30 | Lpin3 | Lpin3 | 5640 | 0.318 | 0.3799 | No |
| 31 | Lmna | Lmna | 5952 | 0.301 | 0.3770 | No |
| 32 | Lpin2 | Lpin2 | 6012 | 0.297 | 0.3841 | No |
| 33 | Cdk1 | Cdk1 | 6047 | 0.296 | 0.3922 | No |
| 34 | Smc4 | Smc4 | 6348 | 0.278 | 0.3890 | No |
| 35 | Plk1 | Plk1 | 6663 | 0.261 | 0.3848 | No |
| 36 | H2ac8 | H2ac8 | 7131 | 0.238 | 0.3737 | No |
| 37 | Nup155 | Nup155 | 7161 | 0.237 | 0.3801 | No |
| 38 | Tpr | Tpr | 7789 | 0.208 | 0.3616 | No |
| 39 | H4c6 | H4c6 | 7806 | 0.208 | 0.3676 | No |
| 40 | H2bc26 | H2bc26 | 8106 | 0.194 | 0.3618 | No |
| 41 | H2bc11 | H2bc11 | 8374 | 0.184 | 0.3570 | No |
| 42 | Golga2 | Golga2 | 8837 | 0.165 | 0.3438 | No |
| 43 | Blzf1 | Blzf1 | 9066 | 0.156 | 0.3396 | No |
| 44 | Ranbp2 | Ranbp2 | 9662 | 0.132 | 0.3200 | No |
| 45 | H2ac19 | H2ac19 | 9696 | 0.130 | 0.3228 | No |
| 46 | Nup93 | Nup93 | 10761 | 0.088 | 0.2831 | No |
| 47 | Aaas | Aaas | 10792 | 0.087 | 0.2846 | No |
| 48 | H3c2 | H3c2 | 11349 | 0.067 | 0.2645 | No |
| 49 | Nup160 | Nup160 | 11368 | 0.066 | 0.2659 | No |
| 50 | Nup37 | Nup37 | 12016 | 0.044 | 0.2414 | No |
| 51 | Nup42 | Nup42 | 12109 | 0.040 | 0.2390 | No |
| 52 | Mastl | Mastl | 12366 | 0.031 | 0.2298 | No |
| 53 | Rab1b | Rab1b | 12476 | 0.027 | 0.2263 | No |
| 54 | Gorasp1 | Gorasp1 | 13362 | -0.004 | 0.1910 | No |
| 55 | Ncaph2 | Ncaph2 | 13394 | -0.005 | 0.1900 | No |
| 56 | Sec13 | Sec13 | 13455 | -0.007 | 0.1878 | No |
| 57 | Nek9 | Nek9 | 13569 | -0.011 | 0.1836 | No |
| 58 | H4c12 | H4c12 | 13836 | -0.021 | 0.1737 | No |
| 59 | Lmnb1 | Lmnb1 | 13883 | -0.022 | 0.1725 | No |
| 60 | Nup205 | Nup205 | 13944 | -0.025 | 0.1709 | No |
| 61 | Rb1 | Rb1 | 14025 | -0.028 | 0.1686 | No |
| 62 | Gorasp2 | Gorasp2 | 14088 | -0.030 | 0.1671 | No |
| 63 | Banf1 | Banf1 | 14123 | -0.031 | 0.1667 | No |
| 64 | H4c16 | H4c16 | 14184 | -0.034 | 0.1654 | No |
| 65 | Ccnb2 | Ccnb2 | 14381 | -0.040 | 0.1588 | No |
| 66 | Rae1 | Rae1 | 14442 | -0.042 | 0.1578 | No |
| 67 | Nup107 | Nup107 | 14656 | -0.051 | 0.1509 | No |
| 68 | H2bc9 | H2bc9 | 16136 | -0.104 | 0.0950 | No |
| 69 | Nup43 | Nup43 | 16213 | -0.107 | 0.0954 | No |
| 70 | H3f3b | H3f3b | 16446 | -0.115 | 0.0898 | No |
| 71 | Vrk1 | Vrk1 | 16670 | -0.124 | 0.0848 | No |
| 72 | Ncapd3 | Ncapd3 | 17220 | -0.145 | 0.0675 | No |
| 73 | Pom121 | Pom121 | 17433 | -0.154 | 0.0639 | No |
| 74 | Set | Set | 17513 | -0.157 | 0.0657 | No |
| 75 | Nup35 | Nup35 | 17569 | -0.159 | 0.0686 | No |
| 76 | Nup210 | Nup210 | 17610 | -0.160 | 0.0721 | No |
| 77 | Nup188 | Nup188 | 17664 | -0.163 | 0.0752 | No |
| 78 | Nup50 | Nup50 | 17861 | -0.172 | 0.0728 | No |
| 79 | H2ax | H2ax | 18012 | -0.178 | 0.0725 | No |
| 80 | Arpp19 | Arpp19 | 18122 | -0.183 | 0.0740 | No |
| 81 | Nup153 | Nup153 | 18326 | -0.192 | 0.0720 | No |
| 82 | Nup98 | Nup98 | 18434 | -0.197 | 0.0739 | No |
| 83 | Mapk1 | Mapk1 | 18437 | -0.197 | 0.0801 | No |
| 84 | Cnep1r1 | Cnep1r1 | 18509 | -0.200 | 0.0837 | No |
| 85 | Seh1l | Seh1l | 18589 | -0.203 | 0.0870 | No |
| 86 | Numa1 | Numa1 | 18890 | -0.217 | 0.0819 | No |
| 87 | Mapk3 | Mapk3 | 19068 | -0.224 | 0.0819 | No |
| 88 | H2az2 | H2az2 | 19504 | -0.244 | 0.0723 | No |
| 89 | Rbm39 | Rbm39 | 19538 | -0.246 | 0.0788 | No |
| 90 | Rab1a | Rab1a | 19654 | -0.253 | 0.0823 | No |
| 91 | Ctdnep1 | Ctdnep1 | 19830 | -0.262 | 0.0836 | No |
| 92 | H3f3a | H3f3a | 19922 | -0.267 | 0.0885 | No |
| 93 | Rab2a | Rab2a | 20159 | -0.279 | 0.0879 | No |
| 94 | Nup214 | Nup214 | 20215 | -0.282 | 0.0947 | No |
| 95 | Nup85 | Nup85 | 20585 | -0.303 | 0.0896 | No |
| 96 | Nup54 | Nup54 | 20591 | -0.303 | 0.0990 | No |
| 97 | Nup88 | Nup88 | 21498 | -0.360 | 0.0743 | No |
| 98 | Nup62 | Nup62 | 21516 | -0.361 | 0.0851 | No |
| 99 | Emd | Emd | 21664 | -0.373 | 0.0911 | No |
| 100 | Nup133 | Nup133 | 22216 | -0.416 | 0.0823 | No |
| 101 | Ndc1 | Ndc1 | 22885 | -0.485 | 0.0710 | No |
| 102 | H2bc21 | H2bc21 | 23472 | -0.560 | 0.0654 | No |