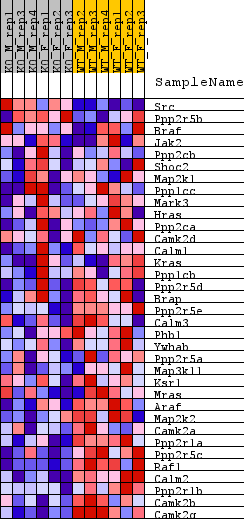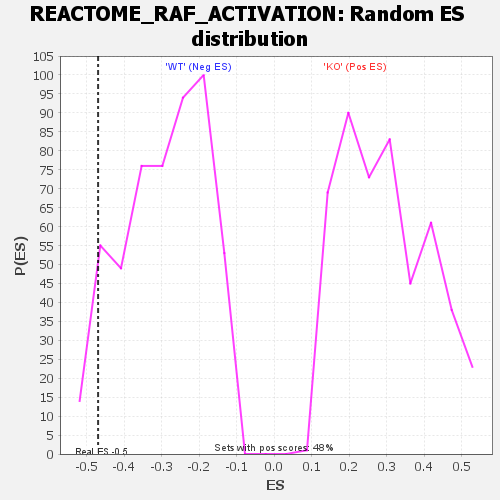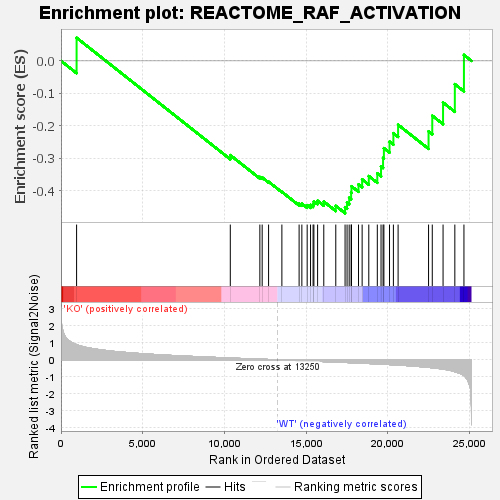
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | WT |
| GeneSet | REACTOME_RAF_ACTIVATION |
| Enrichment Score (ES) | -0.47003826 |
| Normalized Enrichment Score (NES) | -1.6109496 |
| Nominal p-value | 0.06963249 |
| FDR q-value | 0.6145699 |
| FWER p-Value | 0.954 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Src | Src | 954 | 0.889 | 0.0713 | No |
| 2 | Ppp2r5b | Ppp2r5b | 10355 | 0.104 | -0.2908 | No |
| 3 | Braf | Braf | 12163 | 0.038 | -0.3582 | No |
| 4 | Jak2 | Jak2 | 12305 | 0.033 | -0.3597 | No |
| 5 | Ppp2cb | Ppp2cb | 12697 | 0.020 | -0.3729 | No |
| 6 | Shoc2 | Shoc2 | 13510 | -0.009 | -0.4041 | No |
| 7 | Map2k1 | Map2k1 | 14564 | -0.047 | -0.4403 | No |
| 8 | Ppp1cc | Ppp1cc | 14732 | -0.054 | -0.4404 | No |
| 9 | Mark3 | Mark3 | 15057 | -0.065 | -0.4453 | No |
| 10 | Hras | Hras | 15263 | -0.073 | -0.4445 | No |
| 11 | Ppp2ca | Ppp2ca | 15419 | -0.078 | -0.4412 | No |
| 12 | Camk2d | Camk2d | 15468 | -0.079 | -0.4333 | No |
| 13 | Calm1 | Calm1 | 15695 | -0.088 | -0.4315 | No |
| 14 | Kras | Kras | 16073 | -0.102 | -0.4340 | No |
| 15 | Ppp1cb | Ppp1cb | 16807 | -0.129 | -0.4474 | No |
| 16 | Ppp2r5d | Ppp2r5d | 17375 | -0.151 | -0.4514 | Yes |
| 17 | Brap | Brap | 17497 | -0.157 | -0.4370 | Yes |
| 18 | Ppp2r5e | Ppp2r5e | 17626 | -0.161 | -0.4223 | Yes |
| 19 | Calm3 | Calm3 | 17745 | -0.166 | -0.4065 | Yes |
| 20 | Phb1 | Phb1 | 17769 | -0.168 | -0.3868 | Yes |
| 21 | Ywhab | Ywhab | 18198 | -0.186 | -0.3810 | Yes |
| 22 | Ppp2r5a | Ppp2r5a | 18419 | -0.196 | -0.3656 | Yes |
| 23 | Map3k11 | Map3k11 | 18826 | -0.214 | -0.3555 | Yes |
| 24 | Ksr1 | Ksr1 | 19347 | -0.236 | -0.3472 | Yes |
| 25 | Mras | Mras | 19575 | -0.249 | -0.3256 | Yes |
| 26 | Araf | Araf | 19699 | -0.256 | -0.2991 | Yes |
| 27 | Map2k2 | Map2k2 | 19750 | -0.258 | -0.2694 | Yes |
| 28 | Camk2a | Camk2a | 20094 | -0.276 | -0.2492 | Yes |
| 29 | Ppp2r1a | Ppp2r1a | 20331 | -0.288 | -0.2232 | Yes |
| 30 | Ppp2r5c | Ppp2r5c | 20619 | -0.305 | -0.1971 | Yes |
| 31 | Raf1 | Raf1 | 22482 | -0.440 | -0.2173 | Yes |
| 32 | Calm2 | Calm2 | 22709 | -0.465 | -0.1691 | Yes |
| 33 | Ppp2r1b | Ppp2r1b | 23369 | -0.544 | -0.1285 | Yes |
| 34 | Camk2b | Camk2b | 24089 | -0.694 | -0.0719 | Yes |
| 35 | Camk2g | Camk2g | 24646 | -0.915 | 0.0184 | Yes |