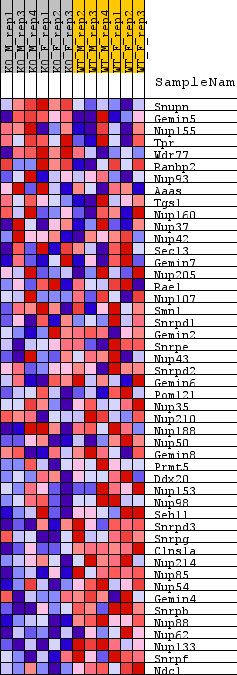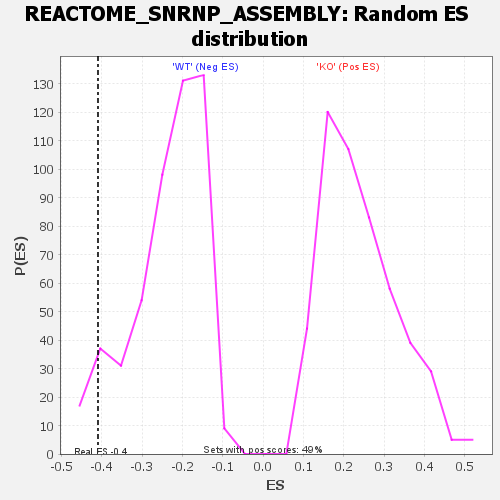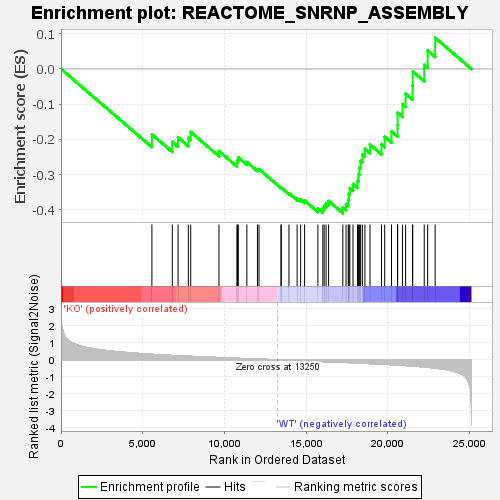
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | WT |
| GeneSet | REACTOME_SNRNP_ASSEMBLY |
| Enrichment Score (ES) | -0.4096739 |
| Normalized Enrichment Score (NES) | -1.7318839 |
| Nominal p-value | 0.0627451 |
| FDR q-value | 0.9841203 |
| FWER p-Value | 0.75 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Snupn | Snupn | 5560 | 0.323 | -0.1856 | No |
| 2 | Gemin5 | Gemin5 | 6809 | 0.254 | -0.2069 | No |
| 3 | Nup155 | Nup155 | 7161 | 0.237 | -0.1943 | No |
| 4 | Tpr | Tpr | 7789 | 0.208 | -0.1959 | No |
| 5 | Wdr77 | Wdr77 | 7934 | 0.202 | -0.1790 | No |
| 6 | Ranbp2 | Ranbp2 | 9662 | 0.132 | -0.2331 | No |
| 7 | Nup93 | Nup93 | 10761 | 0.088 | -0.2670 | No |
| 8 | Aaas | Aaas | 10792 | 0.087 | -0.2584 | No |
| 9 | Tgs1 | Tgs1 | 10846 | 0.085 | -0.2509 | No |
| 10 | Nup160 | Nup160 | 11368 | 0.066 | -0.2643 | No |
| 11 | Nup37 | Nup37 | 12016 | 0.044 | -0.2852 | No |
| 12 | Nup42 | Nup42 | 12109 | 0.040 | -0.2844 | No |
| 13 | Sec13 | Sec13 | 13455 | -0.007 | -0.3372 | No |
| 14 | Gemin7 | Gemin7 | 13471 | -0.008 | -0.3370 | No |
| 15 | Nup205 | Nup205 | 13944 | -0.025 | -0.3530 | No |
| 16 | Rae1 | Rae1 | 14442 | -0.042 | -0.3681 | No |
| 17 | Nup107 | Nup107 | 14656 | -0.051 | -0.3709 | No |
| 18 | Smn1 | Smn1 | 14901 | -0.059 | -0.3739 | No |
| 19 | Snrpd1 | Snrpd1 | 15713 | -0.089 | -0.3964 | No |
| 20 | Gemin2 | Gemin2 | 16016 | -0.100 | -0.3972 | Yes |
| 21 | Snrpe | Snrpe | 16094 | -0.103 | -0.3887 | Yes |
| 22 | Nup43 | Nup43 | 16213 | -0.107 | -0.3814 | Yes |
| 23 | Snrpd2 | Snrpd2 | 16367 | -0.112 | -0.3749 | Yes |
| 24 | Gemin6 | Gemin6 | 17239 | -0.146 | -0.3933 | Yes |
| 25 | Pom121 | Pom121 | 17433 | -0.154 | -0.3837 | Yes |
| 26 | Nup35 | Nup35 | 17569 | -0.159 | -0.3712 | Yes |
| 27 | Nup210 | Nup210 | 17610 | -0.160 | -0.3548 | Yes |
| 28 | Nup188 | Nup188 | 17664 | -0.163 | -0.3386 | Yes |
| 29 | Nup50 | Nup50 | 17861 | -0.172 | -0.3271 | Yes |
| 30 | Gemin8 | Gemin8 | 18132 | -0.184 | -0.3172 | Yes |
| 31 | Prmt5 | Prmt5 | 18200 | -0.186 | -0.2989 | Yes |
| 32 | Ddx20 | Ddx20 | 18253 | -0.189 | -0.2798 | Yes |
| 33 | Nup153 | Nup153 | 18326 | -0.192 | -0.2611 | Yes |
| 34 | Nup98 | Nup98 | 18434 | -0.197 | -0.2432 | Yes |
| 35 | Seh1l | Seh1l | 18589 | -0.203 | -0.2265 | Yes |
| 36 | Snrpd3 | Snrpd3 | 18901 | -0.217 | -0.2145 | Yes |
| 37 | Snrpg | Snrpg | 19603 | -0.250 | -0.2143 | Yes |
| 38 | Clns1a | Clns1a | 19799 | -0.261 | -0.1928 | Yes |
| 39 | Nup214 | Nup214 | 20215 | -0.282 | -0.1777 | Yes |
| 40 | Nup85 | Nup85 | 20585 | -0.303 | -0.1584 | Yes |
| 41 | Nup54 | Nup54 | 20591 | -0.303 | -0.1245 | Yes |
| 42 | Gemin4 | Gemin4 | 20885 | -0.320 | -0.1003 | Yes |
| 43 | Snrpb | Snrpb | 21078 | -0.332 | -0.0706 | Yes |
| 44 | Nup88 | Nup88 | 21498 | -0.360 | -0.0468 | Yes |
| 45 | Nup62 | Nup62 | 21516 | -0.361 | -0.0069 | Yes |
| 46 | Nup133 | Nup133 | 22216 | -0.416 | 0.0120 | Yes |
| 47 | Snrpf | Snrpf | 22427 | -0.435 | 0.0524 | Yes |
| 48 | Ndc1 | Ndc1 | 22885 | -0.485 | 0.0887 | Yes |