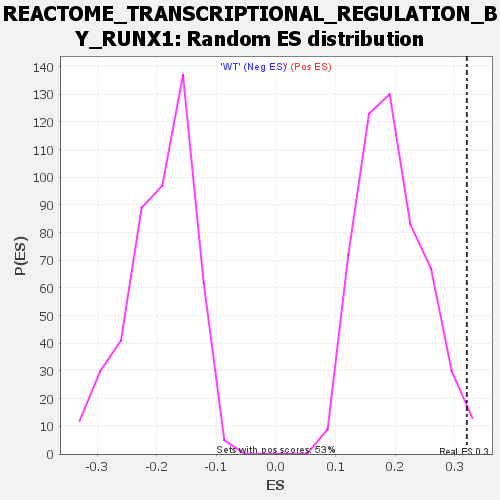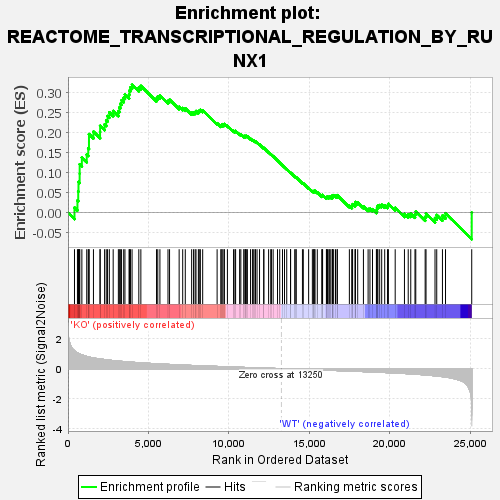
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | KO |
| GeneSet | REACTOME_TRANSCRIPTIONAL_REGULATION_BY_RUNX1 |
| Enrichment Score (ES) | 0.3205336 |
| Normalized Enrichment Score (NES) | 1.6378598 |
| Nominal p-value | 0.0056926 |
| FDR q-value | 0.6614499 |
| FWER p-Value | 0.922 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Ccnd2 | Ccnd2 | 403 | 1.195 | 0.0126 | Yes |
| 2 | H3c7 | H3c7 | 587 | 1.051 | 0.0305 | Yes |
| 3 | H2bc8 | H2bc8 | 636 | 1.025 | 0.0532 | Yes |
| 4 | H2bc6 | H2bc6 | 652 | 1.014 | 0.0770 | Yes |
| 5 | H2ac10 | H2ac10 | 724 | 0.979 | 0.0977 | Yes |
| 6 | H2ac6 | H2ac6 | 726 | 0.979 | 0.1211 | Yes |
| 7 | H3c6 | H3c6 | 862 | 0.927 | 0.1380 | Yes |
| 8 | Lmo2 | Lmo2 | 1165 | 0.819 | 0.1456 | Yes |
| 9 | H4c8 | H4c8 | 1258 | 0.795 | 0.1610 | Yes |
| 10 | H4c3 | H4c3 | 1304 | 0.783 | 0.1780 | Yes |
| 11 | H2aj | H2aj | 1305 | 0.782 | 0.1968 | Yes |
| 12 | H2bc13 | H2bc13 | 1582 | 0.716 | 0.2030 | Yes |
| 13 | H2ac7 | H2ac7 | 1993 | 0.649 | 0.2022 | Yes |
| 14 | H3c15 | H3c15 | 1998 | 0.647 | 0.2176 | Yes |
| 15 | H3c11 | H3c11 | 2275 | 0.607 | 0.2211 | Yes |
| 16 | Ctsk | Ctsk | 2377 | 0.592 | 0.2313 | Yes |
| 17 | H2bc15 | H2bc15 | 2459 | 0.582 | 0.2420 | Yes |
| 18 | H2bc4 | H2bc4 | 2570 | 0.570 | 0.2513 | Yes |
| 19 | H4c4 | H4c4 | 2809 | 0.540 | 0.2548 | Yes |
| 20 | H3c4 | H3c4 | 3131 | 0.505 | 0.2540 | Yes |
| 21 | H4c14 | H4c14 | 3193 | 0.498 | 0.2636 | Yes |
| 22 | H4c1 | H4c1 | 3255 | 0.492 | 0.2729 | Yes |
| 23 | H2bc3 | H2bc3 | 3330 | 0.486 | 0.2816 | Yes |
| 24 | H4c11 | H4c11 | 3463 | 0.475 | 0.2878 | Yes |
| 25 | Ccnd3 | Ccnd3 | 3540 | 0.467 | 0.2960 | Yes |
| 26 | H4c9 | H4c9 | 3797 | 0.444 | 0.2964 | Yes |
| 27 | Smarcd3 | Smarcd3 | 3822 | 0.442 | 0.3060 | Yes |
| 28 | Cdk6 | Cdk6 | 3884 | 0.438 | 0.3141 | Yes |
| 29 | Phc3 | Phc3 | 3983 | 0.431 | 0.3205 | Yes |
| 30 | Lmo1 | Lmo1 | 4403 | 0.398 | 0.3133 | No |
| 31 | H3c8 | H3c8 | 4530 | 0.389 | 0.3176 | No |
| 32 | Psma5 | Psma5 | 5504 | 0.326 | 0.2865 | No |
| 33 | Ctsl | Ctsl | 5581 | 0.321 | 0.2912 | No |
| 34 | Psmd8 | Psmd8 | 5715 | 0.314 | 0.2934 | No |
| 35 | Esr1 | Esr1 | 6217 | 0.286 | 0.2802 | No |
| 36 | Foxp3 | Foxp3 | 6314 | 0.280 | 0.2831 | No |
| 37 | Cdk7 | Cdk7 | 6910 | 0.249 | 0.2652 | No |
| 38 | H2ac8 | H2ac8 | 7131 | 0.238 | 0.2621 | No |
| 39 | Smarcc1 | Smarcc1 | 7292 | 0.231 | 0.2613 | No |
| 40 | Ring1 | Ring1 | 7688 | 0.213 | 0.2506 | No |
| 41 | H4c6 | H4c6 | 7806 | 0.208 | 0.2509 | No |
| 42 | Psma7 | Psma7 | 7903 | 0.203 | 0.2519 | No |
| 43 | Rps27a | Rps27a | 7958 | 0.201 | 0.2546 | No |
| 44 | H2bc26 | H2bc26 | 8106 | 0.194 | 0.2533 | No |
| 45 | Psmd14 | Psmd14 | 8161 | 0.192 | 0.2558 | No |
| 46 | Kmt2a | Kmt2a | 8229 | 0.190 | 0.2577 | No |
| 47 | H2bc11 | H2bc11 | 8374 | 0.184 | 0.2563 | No |
| 48 | Itch | Itch | 9271 | 0.148 | 0.2240 | No |
| 49 | Mnat1 | Mnat1 | 9499 | 0.138 | 0.2182 | No |
| 50 | Sin3b | Sin3b | 9569 | 0.136 | 0.2187 | No |
| 51 | Cbx6 | Cbx6 | 9580 | 0.135 | 0.2215 | No |
| 52 | H2ac19 | H2ac19 | 9696 | 0.130 | 0.2201 | No |
| 53 | Psmc1 | Psmc1 | 9723 | 0.129 | 0.2221 | No |
| 54 | Pcgf5 | Pcgf5 | 9911 | 0.122 | 0.2175 | No |
| 55 | Kat2b | Kat2b | 10292 | 0.107 | 0.2049 | No |
| 56 | Psmd13 | Psmd13 | 10378 | 0.103 | 0.2040 | No |
| 57 | Ubb | Ubb | 10421 | 0.102 | 0.2047 | No |
| 58 | Gata3 | Gata3 | 10671 | 0.092 | 0.1970 | No |
| 59 | Smarce1 | Smarce1 | 10746 | 0.089 | 0.1961 | No |
| 60 | Prmt1 | Prmt1 | 10936 | 0.082 | 0.1905 | No |
| 61 | Psmd2 | Psmd2 | 10945 | 0.081 | 0.1922 | No |
| 62 | Cbx8 | Cbx8 | 11042 | 0.078 | 0.1902 | No |
| 63 | Hdac1 | Hdac1 | 11044 | 0.078 | 0.1920 | No |
| 64 | Psmc3 | Psmc3 | 11085 | 0.076 | 0.1923 | No |
| 65 | Pbrm1 | Pbrm1 | 11153 | 0.074 | 0.1913 | No |
| 66 | H3c2 | H3c2 | 11349 | 0.067 | 0.1851 | No |
| 67 | Psma6 | Psma6 | 11467 | 0.063 | 0.1819 | No |
| 68 | Pax5 | Pax5 | 11520 | 0.061 | 0.1813 | No |
| 69 | Psmc5 | Psmc5 | 11622 | 0.057 | 0.1786 | No |
| 70 | Rybp | Rybp | 11659 | 0.055 | 0.1785 | No |
| 71 | Pml | Pml | 11754 | 0.052 | 0.1760 | No |
| 72 | Psmd6 | Psmd6 | 11909 | 0.047 | 0.1710 | No |
| 73 | Psmb1 | Psmb1 | 12172 | 0.038 | 0.1614 | No |
| 74 | Rnf2 | Rnf2 | 12175 | 0.038 | 0.1622 | No |
| 75 | Cbx4 | Cbx4 | 12473 | 0.027 | 0.1510 | No |
| 76 | Ptpn11 | Ptpn11 | 12607 | 0.023 | 0.1462 | No |
| 77 | Psmd1 | Psmd1 | 12648 | 0.022 | 0.1452 | No |
| 78 | Psmd7 | Psmd7 | 12755 | 0.018 | 0.1413 | No |
| 79 | Psmc6 | Psmc6 | 13026 | 0.008 | 0.1307 | No |
| 80 | Phc1 | Phc1 | 13164 | 0.003 | 0.1253 | No |
| 81 | Ash2l | Ash2l | 13336 | -0.003 | 0.1185 | No |
| 82 | Tal1 | Tal1 | 13461 | -0.008 | 0.1137 | No |
| 83 | Tcf12 | Tcf12 | 13600 | -0.013 | 0.1085 | No |
| 84 | H4c12 | H4c12 | 13836 | -0.021 | 0.0996 | No |
| 85 | Phc2 | Phc2 | 13840 | -0.021 | 0.1000 | No |
| 86 | Zfpm1 | Zfpm1 | 14096 | -0.031 | 0.0905 | No |
| 87 | Uba52rt | Uba52rt | 14172 | -0.033 | 0.0883 | No |
| 88 | H4c16 | H4c16 | 14184 | -0.034 | 0.0887 | No |
| 89 | Psmd12 | Psmd12 | 14576 | -0.047 | 0.0741 | No |
| 90 | Cbx2 | Cbx2 | 14629 | -0.049 | 0.0732 | No |
| 91 | Ep300 | Ep300 | 14962 | -0.062 | 0.0614 | No |
| 92 | Psmc4 | Psmc4 | 15193 | -0.070 | 0.0539 | No |
| 93 | Ccnd1 | Ccnd1 | 15281 | -0.073 | 0.0522 | No |
| 94 | Smarcd2 | Smarcd2 | 15286 | -0.073 | 0.0538 | No |
| 95 | Uba52 | Uba52 | 15308 | -0.074 | 0.0547 | No |
| 96 | Tcf3 | Tcf3 | 15360 | -0.076 | 0.0545 | No |
| 97 | Smarcb1 | Smarcb1 | 15500 | -0.081 | 0.0509 | No |
| 98 | Arid1a | Arid1a | 15784 | -0.091 | 0.0417 | No |
| 99 | Ldb1 | Ldb1 | 15805 | -0.091 | 0.0431 | No |
| 100 | Setd1b | Setd1b | 15829 | -0.092 | 0.0444 | No |
| 101 | Psma3 | Psma3 | 16066 | -0.102 | 0.0374 | No |
| 102 | Csnk2a2 | Csnk2a2 | 16082 | -0.102 | 0.0392 | No |
| 103 | H2bc9 | H2bc9 | 16136 | -0.104 | 0.0396 | No |
| 104 | Psmb7 | Psmb7 | 16224 | -0.107 | 0.0387 | No |
| 105 | Adrm1 | Adrm1 | 16245 | -0.108 | 0.0405 | No |
| 106 | Psmb5 | Psmb5 | 16338 | -0.111 | 0.0395 | No |
| 107 | Psma4 | Psma4 | 16432 | -0.115 | 0.0385 | No |
| 108 | Psmd11 | Psmd11 | 16444 | -0.115 | 0.0408 | No |
| 109 | H3f3b | H3f3b | 16446 | -0.115 | 0.0435 | No |
| 110 | Psma2 | Psma2 | 16516 | -0.118 | 0.0436 | No |
| 111 | Psmd3 | Psmd3 | 16632 | -0.122 | 0.0419 | No |
| 112 | Gata2 | Gata2 | 16737 | -0.126 | 0.0408 | No |
| 113 | Ubc | Ubc | 16741 | -0.126 | 0.0437 | No |
| 114 | Smarcc2 | Smarcc2 | 17494 | -0.157 | 0.0174 | No |
| 115 | Psma1 | Psma1 | 17651 | -0.162 | 0.0150 | No |
| 116 | Psmb3 | Psmb3 | 17663 | -0.163 | 0.0185 | No |
| 117 | Prmt6 | Prmt6 | 17695 | -0.165 | 0.0212 | No |
| 118 | Psmc2 | Psmc2 | 17846 | -0.171 | 0.0193 | No |
| 119 | Psmb4 | Psmb4 | 17870 | -0.172 | 0.0225 | No |
| 120 | Yap1 | Yap1 | 17872 | -0.172 | 0.0266 | No |
| 121 | H2ax | H2ax | 18012 | -0.178 | 0.0253 | No |
| 122 | Hipk2 | Hipk2 | 18368 | -0.194 | 0.0158 | No |
| 123 | Elf1 | Elf1 | 18659 | -0.207 | 0.0091 | No |
| 124 | Cbfb | Cbfb | 18774 | -0.212 | 0.0096 | No |
| 125 | Yaf2 | Yaf2 | 18932 | -0.218 | 0.0086 | No |
| 126 | Elf2 | Elf2 | 19174 | -0.229 | 0.0045 | No |
| 127 | Sin3a | Sin3a | 19207 | -0.231 | 0.0087 | No |
| 128 | Bmi1 | Bmi1 | 19222 | -0.231 | 0.0137 | No |
| 129 | Psmb6 | Psmb6 | 19269 | -0.233 | 0.0175 | No |
| 130 | Rbbp5 | Rbbp5 | 19379 | -0.238 | 0.0188 | No |
| 131 | H2az2 | H2az2 | 19504 | -0.244 | 0.0197 | No |
| 132 | Setd1a | Setd1a | 19691 | -0.255 | 0.0184 | No |
| 133 | Kmt2b | Kmt2b | 19864 | -0.264 | 0.0178 | No |
| 134 | H3f3a | H3f3a | 19922 | -0.267 | 0.0220 | No |
| 135 | Csnk2a1 | Csnk2a1 | 20343 | -0.289 | 0.0121 | No |
| 136 | Auts2 | Auts2 | 20919 | -0.321 | -0.0032 | No |
| 137 | Actl6a | Actl6a | 21153 | -0.336 | -0.0045 | No |
| 138 | Psmb2 | Psmb2 | 21311 | -0.347 | -0.0024 | No |
| 139 | Abl1 | Abl1 | 21574 | -0.366 | -0.0041 | No |
| 140 | Csnk2b | Csnk2b | 21623 | -0.370 | 0.0028 | No |
| 141 | Wdr5 | Wdr5 | 22206 | -0.415 | -0.0105 | No |
| 142 | Ccnh | Ccnh | 22260 | -0.419 | -0.0026 | No |
| 143 | Scmh1 | Scmh1 | 22819 | -0.477 | -0.0135 | No |
| 144 | Kmt2d | Kmt2d | 22926 | -0.490 | -0.0059 | No |
| 145 | Smarcd1 | Smarcd1 | 23281 | -0.532 | -0.0073 | No |
| 146 | H2bc21 | H2bc21 | 23472 | -0.560 | -0.0015 | No |
| 147 | Smarca4 | Smarca4 | 25102 | -2.785 | 0.0002 | No |