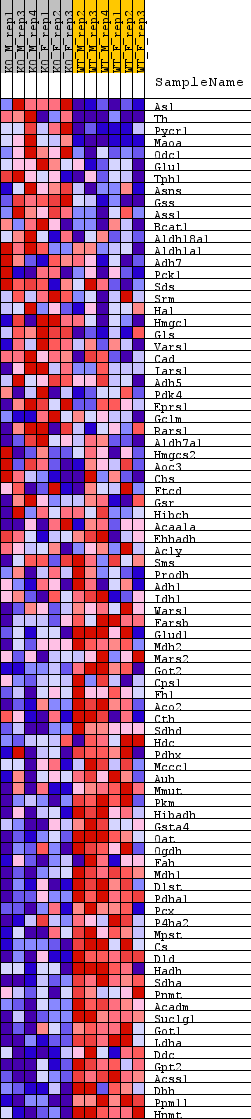
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | WT |
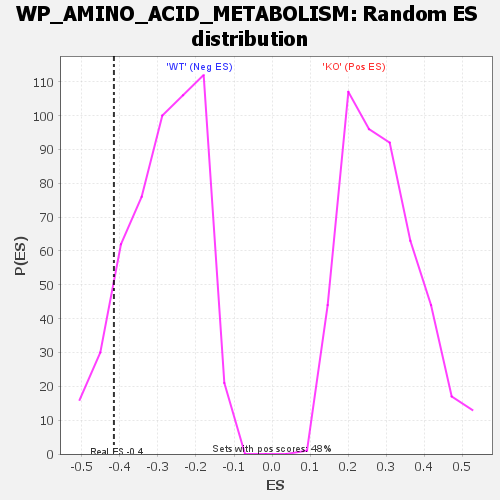
| GeneSet | WP_AMINO_ACID_METABOLISM |
| Enrichment Score (ES) | -0.41552952 |
| Normalized Enrichment Score (NES) | -1.4536703 |
| Nominal p-value | 0.107074566 |
| FDR q-value | 0.38560593 |
| FWER p-Value | 0.824 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Asl | Asl | 224 | 1.418 | 0.0285 | No |
| 2 | Th | Th | 621 | 1.031 | 0.0399 | No |
| 3 | Pycr1 | Pycr1 | 952 | 0.890 | 0.0502 | No |
| 4 | Maoa | Maoa | 1033 | 0.857 | 0.0696 | No |
| 5 | Odc1 | Odc1 | 1857 | 0.672 | 0.0544 | No |
| 6 | Glul | Glul | 1892 | 0.666 | 0.0706 | No |
| 7 | Tph1 | Tph1 | 2101 | 0.633 | 0.0790 | No |
| 8 | Asns | Asns | 2210 | 0.617 | 0.0910 | No |
| 9 | Gss | Gss | 2400 | 0.589 | 0.0990 | No |
| 10 | Ass1 | Ass1 | 2629 | 0.562 | 0.1047 | No |
| 11 | Bcat1 | Bcat1 | 3613 | 0.460 | 0.0776 | No |
| 12 | Aldh18a1 | Aldh18a1 | 3774 | 0.446 | 0.0830 | No |
| 13 | Aldh1a1 | Aldh1a1 | 3992 | 0.430 | 0.0856 | No |
| 14 | Adh7 | Adh7 | 4179 | 0.415 | 0.0892 | No |
| 15 | Pck1 | Pck1 | 4406 | 0.398 | 0.0906 | No |
| 16 | Sds | Sds | 4474 | 0.393 | 0.0983 | No |
| 17 | Srm | Srm | 5859 | 0.306 | 0.0511 | No |
| 18 | Hal | Hal | 6546 | 0.267 | 0.0307 | No |
| 19 | Hmgcl | Hmgcl | 6724 | 0.258 | 0.0305 | No |
| 20 | Gls | Gls | 6848 | 0.252 | 0.0322 | No |
| 21 | Vars1 | Vars1 | 6896 | 0.250 | 0.0369 | No |
| 22 | Cad | Cad | 7140 | 0.238 | 0.0335 | No |
| 23 | Iars1 | Iars1 | 9020 | 0.158 | -0.0374 | No |
| 24 | Adh5 | Adh5 | 9089 | 0.155 | -0.0361 | No |
| 25 | Pdk4 | Pdk4 | 10327 | 0.106 | -0.0827 | No |
| 26 | Eprs1 | Eprs1 | 10438 | 0.101 | -0.0845 | No |
| 27 | Gclm | Gclm | 10596 | 0.095 | -0.0882 | No |
| 28 | Rars1 | Rars1 | 10677 | 0.092 | -0.0890 | No |
| 29 | Aldh7a1 | Aldh7a1 | 10979 | 0.080 | -0.0989 | No |
| 30 | Hmgcs2 | Hmgcs2 | 11118 | 0.075 | -0.1024 | No |
| 31 | Aoc3 | Aoc3 | 11298 | 0.068 | -0.1078 | No |
| 32 | Cbs | Cbs | 11638 | 0.056 | -0.1199 | No |
| 33 | Ftcd | Ftcd | 12443 | 0.028 | -0.1512 | No |
| 34 | Gsr | Gsr | 12668 | 0.021 | -0.1597 | No |
| 35 | Hibch | Hibch | 13172 | 0.003 | -0.1797 | No |
| 36 | Acaa1a | Acaa1a | 13587 | -0.012 | -0.1959 | No |
| 37 | Ehhadh | Ehhadh | 15210 | -0.071 | -0.2589 | No |
| 38 | Acly | Acly | 16156 | -0.105 | -0.2939 | No |
| 39 | Sms | Sms | 16697 | -0.124 | -0.3122 | No |
| 40 | Prodh | Prodh | 17817 | -0.170 | -0.3524 | No |
| 41 | Adh1 | Adh1 | 18787 | -0.212 | -0.3855 | No |
| 42 | Idh1 | Idh1 | 19028 | -0.222 | -0.3892 | No |
| 43 | Wars1 | Wars1 | 19416 | -0.240 | -0.3984 | No |
| 44 | Farsb | Farsb | 19846 | -0.263 | -0.4086 | Yes |
| 45 | Glud1 | Glud1 | 19892 | -0.265 | -0.4034 | Yes |
| 46 | Mdh2 | Mdh2 | 20172 | -0.280 | -0.4072 | Yes |
| 47 | Mars2 | Mars2 | 20208 | -0.282 | -0.4011 | Yes |
| 48 | Got2 | Got2 | 20277 | -0.285 | -0.3963 | Yes |
| 49 | Cps1 | Cps1 | 20695 | -0.310 | -0.4048 | Yes |
| 50 | Fh1 | Fh1 | 20746 | -0.312 | -0.3986 | Yes |
| 51 | Aco2 | Aco2 | 20932 | -0.322 | -0.3975 | Yes |
| 52 | Cth | Cth | 21036 | -0.329 | -0.3929 | Yes |
| 53 | Sdhd | Sdhd | 21102 | -0.334 | -0.3867 | Yes |
| 54 | Hdc | Hdc | 21630 | -0.370 | -0.3980 | Yes |
| 55 | Pdhx | Pdhx | 21695 | -0.375 | -0.3906 | Yes |
| 56 | Mccc1 | Mccc1 | 21817 | -0.384 | -0.3853 | Yes |
| 57 | Auh | Auh | 22183 | -0.413 | -0.3890 | Yes |
| 58 | Mmut | Mmut | 22425 | -0.435 | -0.3872 | Yes |
| 59 | Pkm | Pkm | 22432 | -0.435 | -0.3760 | Yes |
| 60 | Hibadh | Hibadh | 22557 | -0.449 | -0.3691 | Yes |
| 61 | Gsta4 | Gsta4 | 22593 | -0.452 | -0.3585 | Yes |
| 62 | Oat | Oat | 22707 | -0.465 | -0.3508 | Yes |
| 63 | Ogdh | Ogdh | 22746 | -0.468 | -0.3399 | Yes |
| 64 | Fah | Fah | 22868 | -0.482 | -0.3320 | Yes |
| 65 | Mdh1 | Mdh1 | 22922 | -0.490 | -0.3212 | Yes |
| 66 | Dlst | Dlst | 22974 | -0.495 | -0.3102 | Yes |
| 67 | Pdha1 | Pdha1 | 23160 | -0.515 | -0.3040 | Yes |
| 68 | Pcx | Pcx | 23179 | -0.518 | -0.2910 | Yes |
| 69 | P4ha2 | P4ha2 | 23196 | -0.520 | -0.2780 | Yes |
| 70 | Mpst | Mpst | 23351 | -0.542 | -0.2698 | Yes |
| 71 | Cs | Cs | 23678 | -0.593 | -0.2672 | Yes |
| 72 | Dld | Dld | 23723 | -0.603 | -0.2530 | Yes |
| 73 | Hadh | Hadh | 23900 | -0.638 | -0.2432 | Yes |
| 74 | Sdha | Sdha | 24002 | -0.667 | -0.2296 | Yes |
| 75 | Pnmt | Pnmt | 24210 | -0.726 | -0.2187 | Yes |
| 76 | Acadm | Acadm | 24213 | -0.728 | -0.1996 | Yes |
| 77 | Suclg1 | Suclg1 | 24285 | -0.753 | -0.1826 | Yes |
| 78 | Got1 | Got1 | 24390 | -0.788 | -0.1659 | Yes |
| 79 | Ldha | Ldha | 24653 | -0.921 | -0.1521 | Yes |
| 80 | Ddc | Ddc | 24677 | -0.942 | -0.1282 | Yes |
| 81 | Gpt2 | Gpt2 | 24678 | -0.943 | -0.1033 | Yes |
| 82 | Acss1 | Acss1 | 24734 | -0.999 | -0.0791 | Yes |
| 83 | Dbh | Dbh | 24754 | -1.023 | -0.0529 | Yes |
| 84 | Ppm1l | Ppm1l | 24870 | -1.188 | -0.0261 | Yes |
| 85 | Hnmt | Hnmt | 24934 | -1.347 | 0.0069 | Yes |