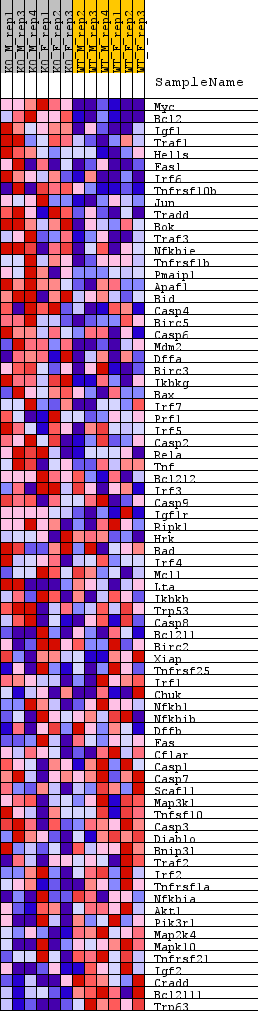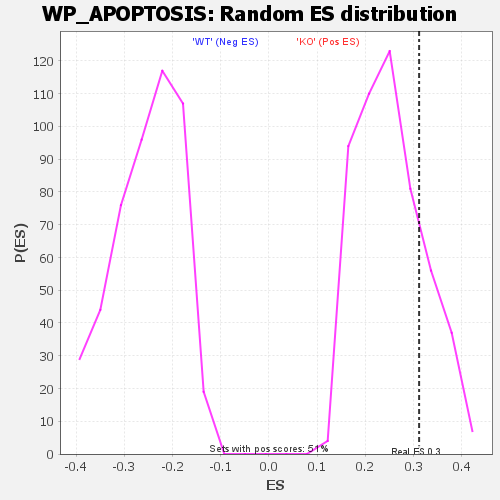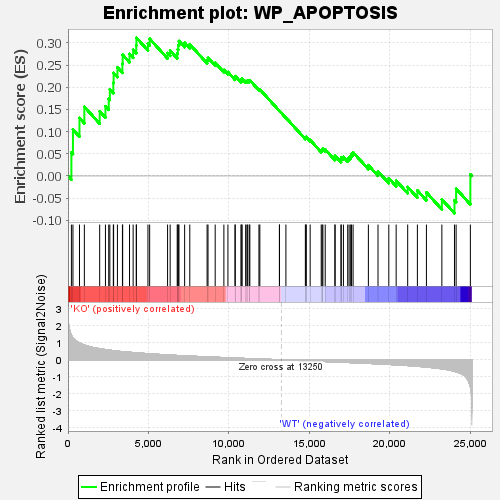
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | KO |
| GeneSet | WP_APOPTOSIS |
| Enrichment Score (ES) | 0.31141114 |
| Normalized Enrichment Score (NES) | 1.2354933 |
| Nominal p-value | 0.1953125 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.978 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Myc | Myc | 211 | 1.433 | 0.0531 | Yes |
| 2 | Bcl2 | Bcl2 | 307 | 1.295 | 0.1049 | Yes |
| 3 | Igf1 | Igf1 | 714 | 0.982 | 0.1309 | Yes |
| 4 | Traf1 | Traf1 | 1017 | 0.864 | 0.1559 | Yes |
| 5 | Hells | Hells | 1970 | 0.651 | 0.1458 | Yes |
| 6 | Fasl | Fasl | 2328 | 0.599 | 0.1573 | Yes |
| 7 | Irf6 | Irf6 | 2533 | 0.573 | 0.1737 | Yes |
| 8 | Tnfrsf10b | Tnfrsf10b | 2601 | 0.566 | 0.1953 | Yes |
| 9 | Jun | Jun | 2818 | 0.539 | 0.2098 | Yes |
| 10 | Tradd | Tradd | 2836 | 0.537 | 0.2322 | Yes |
| 11 | Bok | Bok | 3073 | 0.511 | 0.2447 | Yes |
| 12 | Traf3 | Traf3 | 3384 | 0.481 | 0.2530 | Yes |
| 13 | Nfkbie | Nfkbie | 3399 | 0.480 | 0.2730 | Yes |
| 14 | Tnfrsf1b | Tnfrsf1b | 3825 | 0.442 | 0.2750 | Yes |
| 15 | Pmaip1 | Pmaip1 | 4050 | 0.425 | 0.2843 | Yes |
| 16 | Apaf1 | Apaf1 | 4244 | 0.410 | 0.2942 | Yes |
| 17 | Bid | Bid | 4255 | 0.409 | 0.3114 | Yes |
| 18 | Casp4 | Casp4 | 4971 | 0.359 | 0.2983 | No |
| 19 | Birc5 | Birc5 | 5077 | 0.351 | 0.3091 | No |
| 20 | Casp6 | Casp6 | 6191 | 0.287 | 0.2770 | No |
| 21 | Mdm2 | Mdm2 | 6351 | 0.278 | 0.2826 | No |
| 22 | Dffa | Dffa | 6790 | 0.255 | 0.2760 | No |
| 23 | Birc3 | Birc3 | 6827 | 0.253 | 0.2855 | No |
| 24 | Ikbkg | Ikbkg | 6856 | 0.252 | 0.2952 | No |
| 25 | Bax | Bax | 6900 | 0.250 | 0.3042 | No |
| 26 | Irf7 | Irf7 | 7252 | 0.233 | 0.3001 | No |
| 27 | Prf1 | Prf1 | 7572 | 0.218 | 0.2967 | No |
| 28 | Irf5 | Irf5 | 8648 | 0.172 | 0.2612 | No |
| 29 | Casp2 | Casp2 | 8705 | 0.170 | 0.2662 | No |
| 30 | Rela | Rela | 9151 | 0.152 | 0.2550 | No |
| 31 | Tnf | Tnf | 9692 | 0.130 | 0.2390 | No |
| 32 | Bcl2l2 | Bcl2l2 | 9943 | 0.121 | 0.2342 | No |
| 33 | Irf3 | Irf3 | 10385 | 0.103 | 0.2210 | No |
| 34 | Casp9 | Casp9 | 10396 | 0.102 | 0.2250 | No |
| 35 | Igf1r | Igf1r | 10752 | 0.089 | 0.2146 | No |
| 36 | Ripk1 | Ripk1 | 10811 | 0.086 | 0.2160 | No |
| 37 | Hrk | Hrk | 10814 | 0.086 | 0.2196 | No |
| 38 | Bad | Bad | 11045 | 0.078 | 0.2138 | No |
| 39 | Irf4 | Irf4 | 11124 | 0.075 | 0.2139 | No |
| 40 | Mcl1 | Mcl1 | 11169 | 0.073 | 0.2153 | No |
| 41 | Lta | Lta | 11267 | 0.069 | 0.2144 | No |
| 42 | Ikbkb | Ikbkb | 11305 | 0.068 | 0.2158 | No |
| 43 | Trp53 | Trp53 | 11872 | 0.048 | 0.1953 | No |
| 44 | Casp8 | Casp8 | 11927 | 0.046 | 0.1951 | No |
| 45 | Bcl2l1 | Bcl2l1 | 13146 | 0.003 | 0.1466 | No |
| 46 | Birc2 | Birc2 | 13147 | 0.003 | 0.1467 | No |
| 47 | Xiap | Xiap | 13548 | -0.011 | 0.1312 | No |
| 48 | Tnfrsf25 | Tnfrsf25 | 14758 | -0.054 | 0.0853 | No |
| 49 | Irf1 | Irf1 | 14797 | -0.056 | 0.0861 | No |
| 50 | Chuk | Chuk | 14808 | -0.056 | 0.0881 | No |
| 51 | Nfkb1 | Nfkb1 | 15056 | -0.065 | 0.0811 | No |
| 52 | Nfkbib | Nfkbib | 15744 | -0.089 | 0.0574 | No |
| 53 | Dffb | Dffb | 15801 | -0.091 | 0.0591 | No |
| 54 | Fas | Fas | 15847 | -0.093 | 0.0613 | No |
| 55 | Cflar | Cflar | 15992 | -0.099 | 0.0598 | No |
| 56 | Casp1 | Casp1 | 16598 | -0.121 | 0.0408 | No |
| 57 | Casp7 | Casp7 | 16604 | -0.121 | 0.0458 | No |
| 58 | Scaf11 | Scaf11 | 16973 | -0.136 | 0.0369 | No |
| 59 | Map3k1 | Map3k1 | 16987 | -0.136 | 0.0423 | No |
| 60 | Tnfsf10 | Tnfsf10 | 17124 | -0.142 | 0.0429 | No |
| 61 | Casp3 | Casp3 | 17391 | -0.152 | 0.0388 | No |
| 62 | Diablo | Diablo | 17478 | -0.156 | 0.0421 | No |
| 63 | Bnip3l | Bnip3l | 17577 | -0.159 | 0.0450 | No |
| 64 | Traf2 | Traf2 | 17629 | -0.161 | 0.0499 | No |
| 65 | Irf2 | Irf2 | 17726 | -0.166 | 0.0532 | No |
| 66 | Tnfrsf1a | Tnfrsf1a | 18676 | -0.207 | 0.0242 | No |
| 67 | Nfkbia | Nfkbia | 19275 | -0.233 | 0.0103 | No |
| 68 | Akt1 | Akt1 | 19946 | -0.268 | -0.0050 | No |
| 69 | Pik3r1 | Pik3r1 | 20403 | -0.293 | -0.0106 | No |
| 70 | Map2k4 | Map2k4 | 21120 | -0.334 | -0.0249 | No |
| 71 | Mapk10 | Mapk10 | 21723 | -0.377 | -0.0327 | No |
| 72 | Tnfrsf21 | Tnfrsf21 | 22285 | -0.421 | -0.0371 | No |
| 73 | Igf2 | Igf2 | 23245 | -0.528 | -0.0527 | No |
| 74 | Cradd | Cradd | 24029 | -0.674 | -0.0550 | No |
| 75 | Bcl2l11 | Bcl2l11 | 24131 | -0.706 | -0.0288 | No |
| 76 | Trp63 | Trp63 | 25017 | -1.577 | 0.0036 | No |