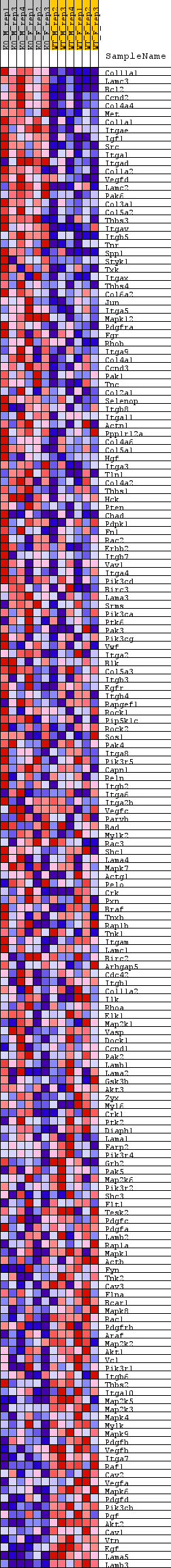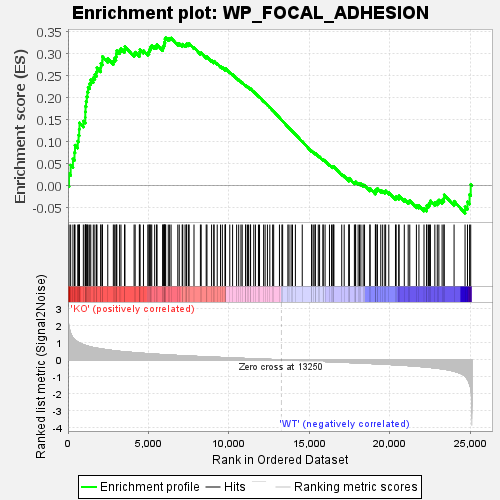
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | KO |
| GeneSet | WP_FOCAL_ADHESION |
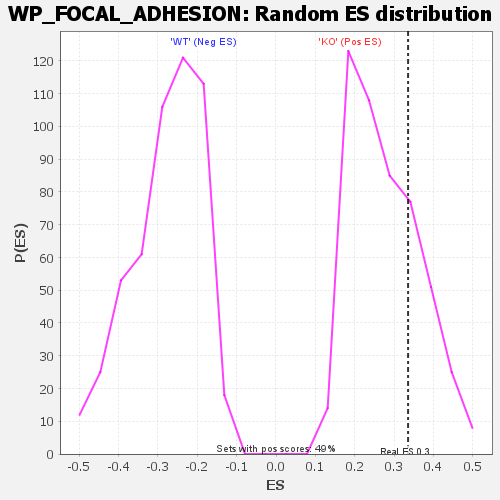
| Enrichment Score (ES) | 0.33575198 |
| Normalized Enrichment Score (NES) | 1.2061418 |
| Nominal p-value | 0.27698573 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.981 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Col11a1 | Col11a1 | 64 | 1.921 | 0.0269 | Yes |
| 2 | Lamc3 | Lamc3 | 165 | 1.554 | 0.0466 | Yes |
| 3 | Bcl2 | Bcl2 | 307 | 1.295 | 0.0608 | Yes |
| 4 | Ccnd2 | Ccnd2 | 403 | 1.195 | 0.0753 | Yes |
| 5 | Col4a4 | Col4a4 | 435 | 1.170 | 0.0920 | Yes |
| 6 | Met | Met | 606 | 1.040 | 0.1011 | Yes |
| 7 | Col1a1 | Col1a1 | 659 | 1.011 | 0.1145 | Yes |
| 8 | Itgae | Itgae | 695 | 0.995 | 0.1283 | Yes |
| 9 | Igf1 | Igf1 | 714 | 0.982 | 0.1426 | Yes |
| 10 | Src | Src | 954 | 0.889 | 0.1467 | Yes |
| 11 | Itgal | Itgal | 1072 | 0.845 | 0.1549 | Yes |
| 12 | Itgad | Itgad | 1075 | 0.844 | 0.1678 | Yes |
| 13 | Col1a2 | Col1a2 | 1089 | 0.840 | 0.1801 | Yes |
| 14 | Vegfd | Vegfd | 1124 | 0.830 | 0.1915 | Yes |
| 15 | Lamc2 | Lamc2 | 1167 | 0.817 | 0.2023 | Yes |
| 16 | Pak6 | Pak6 | 1206 | 0.808 | 0.2132 | Yes |
| 17 | Col3a1 | Col3a1 | 1250 | 0.798 | 0.2237 | Yes |
| 18 | Col5a2 | Col5a2 | 1348 | 0.767 | 0.2315 | Yes |
| 19 | Thbs3 | Thbs3 | 1416 | 0.751 | 0.2403 | Yes |
| 20 | Itgav | Itgav | 1573 | 0.719 | 0.2451 | Yes |
| 21 | Itgb5 | Itgb5 | 1671 | 0.699 | 0.2519 | Yes |
| 22 | Tnr | Tnr | 1774 | 0.682 | 0.2582 | Yes |
| 23 | Spp1 | Spp1 | 1800 | 0.678 | 0.2676 | Yes |
| 24 | Styk1 | Styk1 | 2036 | 0.642 | 0.2680 | Yes |
| 25 | Txk | Txk | 2043 | 0.642 | 0.2776 | Yes |
| 26 | Itgax | Itgax | 2136 | 0.627 | 0.2835 | Yes |
| 27 | Thbs4 | Thbs4 | 2137 | 0.627 | 0.2932 | Yes |
| 28 | Col6a2 | Col6a2 | 2472 | 0.580 | 0.2886 | Yes |
| 29 | Jun | Jun | 2818 | 0.539 | 0.2831 | Yes |
| 30 | Itga5 | Itga5 | 2879 | 0.531 | 0.2888 | Yes |
| 31 | Mapk12 | Mapk12 | 2982 | 0.520 | 0.2926 | Yes |
| 32 | Pdgfra | Pdgfra | 3001 | 0.518 | 0.2999 | Yes |
| 33 | Fgr | Fgr | 3036 | 0.514 | 0.3064 | Yes |
| 34 | Rhob | Rhob | 3209 | 0.497 | 0.3071 | Yes |
| 35 | Itga9 | Itga9 | 3293 | 0.489 | 0.3112 | Yes |
| 36 | Col4a1 | Col4a1 | 3515 | 0.470 | 0.3096 | Yes |
| 37 | Ccnd3 | Ccnd3 | 3540 | 0.467 | 0.3158 | Yes |
| 38 | Pak1 | Pak1 | 4109 | 0.420 | 0.2994 | Yes |
| 39 | Tnc | Tnc | 4175 | 0.415 | 0.3032 | Yes |
| 40 | Col2a1 | Col2a1 | 4435 | 0.395 | 0.2988 | Yes |
| 41 | Selenop | Selenop | 4462 | 0.394 | 0.3038 | Yes |
| 42 | Itgb8 | Itgb8 | 4476 | 0.393 | 0.3093 | Yes |
| 43 | Itga11 | Itga11 | 4692 | 0.377 | 0.3064 | Yes |
| 44 | Actn1 | Actn1 | 4963 | 0.360 | 0.3011 | Yes |
| 45 | Ppp1r12a | Ppp1r12a | 5033 | 0.354 | 0.3038 | Yes |
| 46 | Col4a6 | Col4a6 | 5053 | 0.354 | 0.3084 | Yes |
| 47 | Col5a1 | Col5a1 | 5107 | 0.350 | 0.3117 | Yes |
| 48 | Hgf | Hgf | 5151 | 0.348 | 0.3153 | Yes |
| 49 | Itga3 | Itga3 | 5208 | 0.345 | 0.3183 | Yes |
| 50 | Tln1 | Tln1 | 5389 | 0.334 | 0.3162 | Yes |
| 51 | Col4a2 | Col4a2 | 5507 | 0.326 | 0.3165 | Yes |
| 52 | Thbs1 | Thbs1 | 5535 | 0.324 | 0.3204 | Yes |
| 53 | Hck | Hck | 5886 | 0.305 | 0.3110 | Yes |
| 54 | Pten | Pten | 5905 | 0.304 | 0.3149 | Yes |
| 55 | Chad | Chad | 5949 | 0.301 | 0.3178 | Yes |
| 56 | Pdpk1 | Pdpk1 | 5983 | 0.299 | 0.3211 | Yes |
| 57 | Fn1 | Fn1 | 5989 | 0.299 | 0.3255 | Yes |
| 58 | Rac2 | Rac2 | 6030 | 0.296 | 0.3284 | Yes |
| 59 | Erbb2 | Erbb2 | 6031 | 0.296 | 0.3329 | Yes |
| 60 | Itgb7 | Itgb7 | 6074 | 0.294 | 0.3358 | Yes |
| 61 | Vav1 | Vav1 | 6235 | 0.285 | 0.3337 | No |
| 62 | Itga4 | Itga4 | 6320 | 0.280 | 0.3346 | No |
| 63 | Pik3cd | Pik3cd | 6409 | 0.275 | 0.3353 | No |
| 64 | Birc3 | Birc3 | 6827 | 0.253 | 0.3224 | No |
| 65 | Lama3 | Lama3 | 6916 | 0.249 | 0.3227 | No |
| 66 | Srms | Srms | 7101 | 0.239 | 0.3190 | No |
| 67 | Pik3ca | Pik3ca | 7134 | 0.238 | 0.3214 | No |
| 68 | Ptk6 | Ptk6 | 7279 | 0.232 | 0.3191 | No |
| 69 | Pak3 | Pak3 | 7359 | 0.228 | 0.3195 | No |
| 70 | Pik3cg | Pik3cg | 7381 | 0.227 | 0.3221 | No |
| 71 | Vwf | Vwf | 7500 | 0.221 | 0.3207 | No |
| 72 | Itga2 | Itga2 | 7528 | 0.220 | 0.3230 | No |
| 73 | Blk | Blk | 7834 | 0.207 | 0.3140 | No |
| 74 | Col5a3 | Col5a3 | 8232 | 0.189 | 0.3009 | No |
| 75 | Itgb3 | Itgb3 | 8280 | 0.188 | 0.3019 | No |
| 76 | Egfr | Egfr | 8592 | 0.174 | 0.2921 | No |
| 77 | Itgb4 | Itgb4 | 8640 | 0.172 | 0.2929 | No |
| 78 | Rapgef1 | Rapgef1 | 8927 | 0.161 | 0.2839 | No |
| 79 | Rock1 | Rock1 | 9065 | 0.156 | 0.2808 | No |
| 80 | Pip5k1c | Pip5k1c | 9080 | 0.155 | 0.2826 | No |
| 81 | Rock2 | Rock2 | 9278 | 0.148 | 0.2769 | No |
| 82 | Sos1 | Sos1 | 9483 | 0.139 | 0.2709 | No |
| 83 | Pak4 | Pak4 | 9596 | 0.134 | 0.2684 | No |
| 84 | Itga8 | Itga8 | 9763 | 0.128 | 0.2637 | No |
| 85 | Pik3r5 | Pik3r5 | 9785 | 0.127 | 0.2648 | No |
| 86 | Capn1 | Capn1 | 9795 | 0.126 | 0.2664 | No |
| 87 | Reln | Reln | 10058 | 0.116 | 0.2577 | No |
| 88 | Itgb2 | Itgb2 | 10234 | 0.110 | 0.2523 | No |
| 89 | Itga6 | Itga6 | 10484 | 0.099 | 0.2439 | No |
| 90 | Itga2b | Itga2b | 10613 | 0.095 | 0.2402 | No |
| 91 | Vegfc | Vegfc | 10753 | 0.089 | 0.2360 | No |
| 92 | Parvb | Parvb | 10830 | 0.086 | 0.2342 | No |
| 93 | Bad | Bad | 11045 | 0.078 | 0.2268 | No |
| 94 | Mylk2 | Mylk2 | 11050 | 0.077 | 0.2279 | No |
| 95 | Rac3 | Rac3 | 11165 | 0.073 | 0.2244 | No |
| 96 | Shc1 | Shc1 | 11228 | 0.071 | 0.2230 | No |
| 97 | Lama4 | Lama4 | 11339 | 0.067 | 0.2196 | No |
| 98 | Mapk7 | Mapk7 | 11352 | 0.067 | 0.2202 | No |
| 99 | Actg1 | Actg1 | 11550 | 0.059 | 0.2132 | No |
| 100 | Pelo | Pelo | 11653 | 0.055 | 0.2099 | No |
| 101 | Crk | Crk | 11842 | 0.049 | 0.2031 | No |
| 102 | Pxn | Pxn | 11908 | 0.047 | 0.2012 | No |
| 103 | Braf | Braf | 12163 | 0.038 | 0.1916 | No |
| 104 | Tnxb | Tnxb | 12255 | 0.035 | 0.1885 | No |
| 105 | Rap1b | Rap1b | 12399 | 0.030 | 0.1832 | No |
| 106 | Tnk1 | Tnk1 | 12548 | 0.025 | 0.1777 | No |
| 107 | Itgam | Itgam | 12724 | 0.019 | 0.1710 | No |
| 108 | Lamc1 | Lamc1 | 12788 | 0.016 | 0.1687 | No |
| 109 | Birc2 | Birc2 | 13147 | 0.003 | 0.1544 | No |
| 110 | Arhgap5 | Arhgap5 | 13301 | -0.002 | 0.1483 | No |
| 111 | Cdc42 | Cdc42 | 13345 | -0.004 | 0.1466 | No |
| 112 | Itgb1 | Itgb1 | 13671 | -0.015 | 0.1338 | No |
| 113 | Col11a2 | Col11a2 | 13741 | -0.018 | 0.1313 | No |
| 114 | Ilk | Ilk | 13873 | -0.022 | 0.1264 | No |
| 115 | Rhoa | Rhoa | 13958 | -0.025 | 0.1234 | No |
| 116 | Elk1 | Elk1 | 14138 | -0.032 | 0.1167 | No |
| 117 | Map2k1 | Map2k1 | 14564 | -0.047 | 0.1004 | No |
| 118 | Vasp | Vasp | 15144 | -0.068 | 0.0782 | No |
| 119 | Dock1 | Dock1 | 15167 | -0.069 | 0.0783 | No |
| 120 | Ccnd1 | Ccnd1 | 15281 | -0.073 | 0.0749 | No |
| 121 | Pak2 | Pak2 | 15364 | -0.076 | 0.0728 | No |
| 122 | Lamb1 | Lamb1 | 15377 | -0.076 | 0.0735 | No |
| 123 | Lama2 | Lama2 | 15562 | -0.083 | 0.0674 | No |
| 124 | Gsk3b | Gsk3b | 15642 | -0.086 | 0.0655 | No |
| 125 | Akt3 | Akt3 | 15857 | -0.093 | 0.0584 | No |
| 126 | Zyx | Zyx | 15861 | -0.094 | 0.0597 | No |
| 127 | Myl6 | Myl6 | 15984 | -0.098 | 0.0563 | No |
| 128 | Crkl | Crkl | 16257 | -0.109 | 0.0471 | No |
| 129 | Ptk2 | Ptk2 | 16385 | -0.113 | 0.0437 | No |
| 130 | Diaph1 | Diaph1 | 16468 | -0.116 | 0.0422 | No |
| 131 | Lama1 | Lama1 | 16499 | -0.117 | 0.0428 | No |
| 132 | Farp2 | Farp2 | 16534 | -0.119 | 0.0432 | No |
| 133 | Pik3r4 | Pik3r4 | 17019 | -0.137 | 0.0259 | No |
| 134 | Grb2 | Grb2 | 17172 | -0.143 | 0.0220 | No |
| 135 | Pak5 | Pak5 | 17470 | -0.155 | 0.0125 | No |
| 136 | Map2k6 | Map2k6 | 17471 | -0.155 | 0.0148 | No |
| 137 | Pik3r2 | Pik3r2 | 17481 | -0.156 | 0.0169 | No |
| 138 | Shc3 | Shc3 | 17806 | -0.170 | 0.0065 | No |
| 139 | Flt1 | Flt1 | 17860 | -0.172 | 0.0070 | No |
| 140 | Tesk2 | Tesk2 | 17884 | -0.173 | 0.0087 | No |
| 141 | Pdgfc | Pdgfc | 18039 | -0.180 | 0.0053 | No |
| 142 | Pdgfa | Pdgfa | 18127 | -0.184 | 0.0046 | No |
| 143 | Lamb2 | Lamb2 | 18211 | -0.187 | 0.0041 | No |
| 144 | Rap1a | Rap1a | 18352 | -0.193 | 0.0015 | No |
| 145 | Mapk1 | Mapk1 | 18437 | -0.197 | 0.0011 | No |
| 146 | Actb | Actb | 18770 | -0.211 | -0.0090 | No |
| 147 | Fyn | Fyn | 18780 | -0.212 | -0.0061 | No |
| 148 | Tnk2 | Tnk2 | 19109 | -0.226 | -0.0158 | No |
| 149 | Cav3 | Cav3 | 19123 | -0.227 | -0.0128 | No |
| 150 | Flna | Flna | 19132 | -0.227 | -0.0097 | No |
| 151 | Bcar1 | Bcar1 | 19219 | -0.231 | -0.0096 | No |
| 152 | Mapk8 | Mapk8 | 19234 | -0.232 | -0.0066 | No |
| 153 | Rac1 | Rac1 | 19447 | -0.241 | -0.0114 | No |
| 154 | Pdgfrb | Pdgfrb | 19565 | -0.248 | -0.0123 | No |
| 155 | Araf | Araf | 19699 | -0.256 | -0.0137 | No |
| 156 | Map2k2 | Map2k2 | 19750 | -0.258 | -0.0118 | No |
| 157 | Akt1 | Akt1 | 19946 | -0.268 | -0.0155 | No |
| 158 | Vcl | Vcl | 20361 | -0.290 | -0.0277 | No |
| 159 | Pik3r1 | Pik3r1 | 20403 | -0.293 | -0.0248 | No |
| 160 | Itgb6 | Itgb6 | 20566 | -0.302 | -0.0267 | No |
| 161 | Thbs2 | Thbs2 | 20582 | -0.303 | -0.0227 | No |
| 162 | Itga10 | Itga10 | 20909 | -0.321 | -0.0308 | No |
| 163 | Map2k5 | Map2k5 | 21151 | -0.336 | -0.0353 | No |
| 164 | Map2k3 | Map2k3 | 21250 | -0.343 | -0.0340 | No |
| 165 | Mapk4 | Mapk4 | 21662 | -0.373 | -0.0448 | No |
| 166 | Mylk | Mylk | 21819 | -0.384 | -0.0452 | No |
| 167 | Mapk9 | Mapk9 | 22134 | -0.409 | -0.0515 | No |
| 168 | Pdgfb | Pdgfb | 22298 | -0.423 | -0.0516 | No |
| 169 | Vegfb | Vegfb | 22307 | -0.424 | -0.0454 | No |
| 170 | Itga7 | Itga7 | 22411 | -0.433 | -0.0429 | No |
| 171 | Raf1 | Raf1 | 22482 | -0.440 | -0.0390 | No |
| 172 | Cav2 | Cav2 | 22535 | -0.446 | -0.0342 | No |
| 173 | Vegfa | Vegfa | 22808 | -0.476 | -0.0379 | No |
| 174 | Mapk6 | Mapk6 | 22959 | -0.494 | -0.0363 | No |
| 175 | Pdgfd | Pdgfd | 23059 | -0.503 | -0.0326 | No |
| 176 | Pik3cb | Pik3cb | 23260 | -0.530 | -0.0325 | No |
| 177 | Pgf | Pgf | 23372 | -0.545 | -0.0286 | No |
| 178 | Akt2 | Akt2 | 23388 | -0.547 | -0.0208 | No |
| 179 | Cav1 | Cav1 | 24004 | -0.667 | -0.0353 | No |
| 180 | Vtn | Vtn | 24687 | -0.955 | -0.0480 | No |
| 181 | Egf | Egf | 24838 | -1.130 | -0.0367 | No |
| 182 | Lama5 | Lama5 | 24969 | -1.430 | -0.0200 | No |
| 183 | Lamb3 | Lamb3 | 25039 | -1.666 | 0.0027 | No |