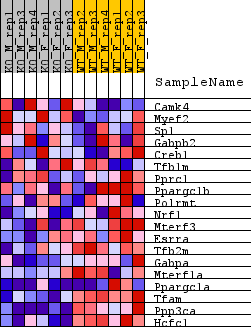
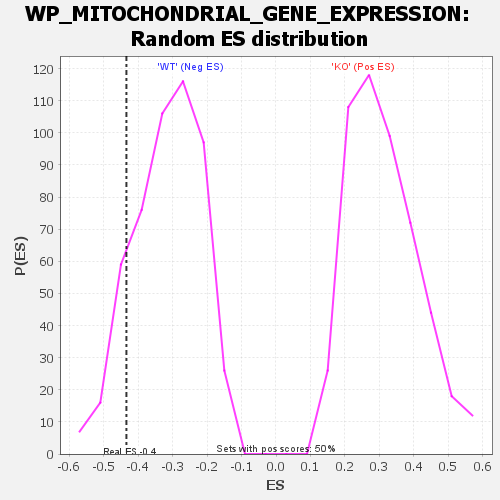
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | WT |
| GeneSet | WP_MITOCHONDRIAL_GENE_EXPRESSION |
| Enrichment Score (ES) | -0.43430293 |
| Normalized Enrichment Score (NES) | -1.3720132 |
| Nominal p-value | 0.12326044 |
| FDR q-value | 0.4934431 |
| FWER p-Value | 0.921 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Camk4 | Camk4 | 1966 | 0.652 | 0.0449 | No |
| 2 | Myef2 | Myef2 | 4387 | 0.400 | 0.0240 | No |
| 3 | Sp1 | Sp1 | 8395 | 0.183 | -0.1011 | No |
| 4 | Gabpb2 | Gabpb2 | 10968 | 0.081 | -0.1883 | No |
| 5 | Creb1 | Creb1 | 12406 | 0.029 | -0.2400 | No |
| 6 | Tfb1m | Tfb1m | 12493 | 0.027 | -0.2384 | No |
| 7 | Pprc1 | Pprc1 | 15609 | -0.085 | -0.3465 | No |
| 8 | Ppargc1b | Ppargc1b | 16963 | -0.135 | -0.3748 | No |
| 9 | Polrmt | Polrmt | 18072 | -0.181 | -0.3848 | Yes |
| 10 | Nrf1 | Nrf1 | 18363 | -0.194 | -0.3597 | Yes |
| 11 | Mterf3 | Mterf3 | 20235 | -0.283 | -0.3808 | Yes |
| 12 | Esrra | Esrra | 20636 | -0.306 | -0.3390 | Yes |
| 13 | Tfb2m | Tfb2m | 20947 | -0.323 | -0.2903 | Yes |
| 14 | Gabpa | Gabpa | 21014 | -0.328 | -0.2310 | Yes |
| 15 | Mterf1a | Mterf1a | 21570 | -0.366 | -0.1839 | Yes |
| 16 | Ppargc1a | Ppargc1a | 21739 | -0.378 | -0.1192 | Yes |
| 17 | Tfam | Tfam | 21956 | -0.395 | -0.0532 | Yes |
| 18 | Ppp3ca | Ppp3ca | 22485 | -0.441 | 0.0091 | Yes |
| 19 | Hcfc1 | Hcfc1 | 23072 | -0.505 | 0.0811 | Yes |