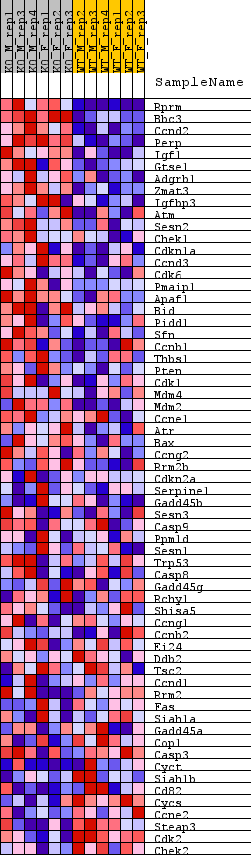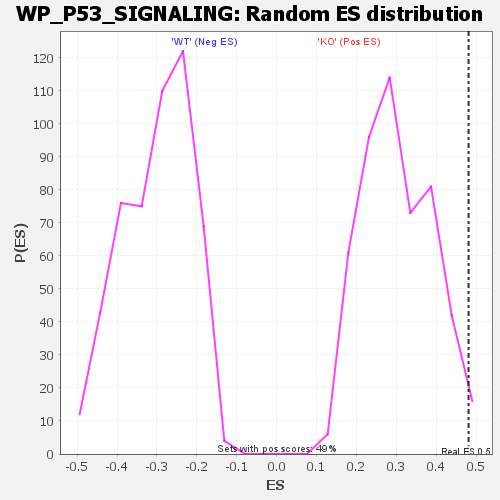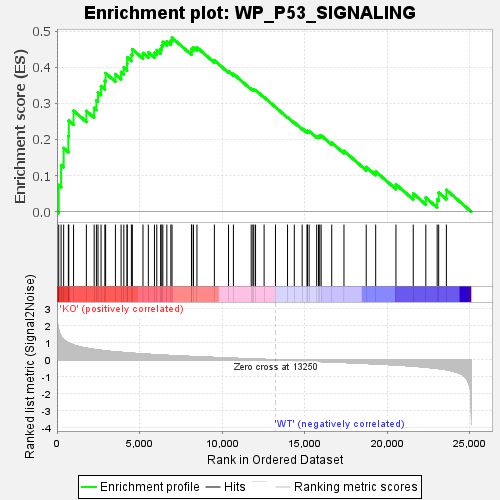
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | KO |
| GeneSet | WP_P53_SIGNALING |
| Enrichment Score (ES) | 0.48132995 |
| Normalized Enrichment Score (NES) | 1.5839511 |
| Nominal p-value | 0.018404908 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.603 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Rprm | Rprm | 105 | 1.750 | 0.0738 | Yes |
| 2 | Bbc3 | Bbc3 | 249 | 1.366 | 0.1290 | Yes |
| 3 | Ccnd2 | Ccnd2 | 403 | 1.195 | 0.1761 | Yes |
| 4 | Perp | Perp | 688 | 0.997 | 0.2092 | Yes |
| 5 | Igf1 | Igf1 | 714 | 0.982 | 0.2520 | Yes |
| 6 | Gtse1 | Gtse1 | 1003 | 0.874 | 0.2794 | Yes |
| 7 | Adgrb1 | Adgrb1 | 1779 | 0.681 | 0.2788 | Yes |
| 8 | Zmat3 | Zmat3 | 2251 | 0.611 | 0.2872 | Yes |
| 9 | Igfbp3 | Igfbp3 | 2388 | 0.591 | 0.3081 | Yes |
| 10 | Atm | Atm | 2484 | 0.579 | 0.3301 | Yes |
| 11 | Sesn2 | Sesn2 | 2669 | 0.557 | 0.3476 | Yes |
| 12 | Chek1 | Chek1 | 2902 | 0.529 | 0.3619 | Yes |
| 13 | Cdkn1a | Cdkn1a | 2946 | 0.523 | 0.3835 | Yes |
| 14 | Ccnd3 | Ccnd3 | 3540 | 0.467 | 0.3806 | Yes |
| 15 | Cdk6 | Cdk6 | 3884 | 0.438 | 0.3864 | Yes |
| 16 | Pmaip1 | Pmaip1 | 4050 | 0.425 | 0.3988 | Yes |
| 17 | Apaf1 | Apaf1 | 4244 | 0.410 | 0.4094 | Yes |
| 18 | Bid | Bid | 4255 | 0.409 | 0.4272 | Yes |
| 19 | Pidd1 | Pidd1 | 4509 | 0.390 | 0.4345 | Yes |
| 20 | Sfn | Sfn | 4566 | 0.387 | 0.4495 | Yes |
| 21 | Ccnb1 | Ccnb1 | 5211 | 0.345 | 0.4391 | Yes |
| 22 | Thbs1 | Thbs1 | 5535 | 0.324 | 0.4407 | Yes |
| 23 | Pten | Pten | 5905 | 0.304 | 0.4395 | Yes |
| 24 | Cdk1 | Cdk1 | 6047 | 0.296 | 0.4470 | Yes |
| 25 | Mdm4 | Mdm4 | 6280 | 0.282 | 0.4503 | Yes |
| 26 | Mdm2 | Mdm2 | 6351 | 0.278 | 0.4599 | Yes |
| 27 | Ccne1 | Ccne1 | 6414 | 0.275 | 0.4697 | Yes |
| 28 | Atr | Atr | 6657 | 0.262 | 0.4717 | Yes |
| 29 | Bax | Bax | 6900 | 0.250 | 0.4732 | Yes |
| 30 | Ccng2 | Ccng2 | 6971 | 0.246 | 0.4813 | Yes |
| 31 | Rrm2b | Rrm2b | 8149 | 0.193 | 0.4429 | No |
| 32 | Cdkn2a | Cdkn2a | 8169 | 0.192 | 0.4507 | No |
| 33 | Serpine1 | Serpine1 | 8273 | 0.188 | 0.4550 | No |
| 34 | Gadd45b | Gadd45b | 8481 | 0.179 | 0.4547 | No |
| 35 | Sesn3 | Sesn3 | 9537 | 0.137 | 0.4187 | No |
| 36 | Casp9 | Casp9 | 10396 | 0.102 | 0.3890 | No |
| 37 | Ppm1d | Ppm1d | 10691 | 0.091 | 0.3813 | No |
| 38 | Sesn1 | Sesn1 | 11769 | 0.051 | 0.3406 | No |
| 39 | Trp53 | Trp53 | 11872 | 0.048 | 0.3387 | No |
| 40 | Casp8 | Casp8 | 11927 | 0.046 | 0.3386 | No |
| 41 | Gadd45g | Gadd45g | 12033 | 0.043 | 0.3363 | No |
| 42 | Rchy1 | Rchy1 | 12551 | 0.025 | 0.3168 | No |
| 43 | Shisa5 | Shisa5 | 13239 | 0.000 | 0.2894 | No |
| 44 | Ccng1 | Ccng1 | 13971 | -0.026 | 0.2613 | No |
| 45 | Ccnb2 | Ccnb2 | 14381 | -0.040 | 0.2468 | No |
| 46 | Ei24 | Ei24 | 14857 | -0.058 | 0.2304 | No |
| 47 | Ddb2 | Ddb2 | 15151 | -0.068 | 0.2218 | No |
| 48 | Tsc2 | Tsc2 | 15183 | -0.070 | 0.2237 | No |
| 49 | Ccnd1 | Ccnd1 | 15281 | -0.073 | 0.2231 | No |
| 50 | Rrm2 | Rrm2 | 15737 | -0.089 | 0.2089 | No |
| 51 | Fas | Fas | 15847 | -0.093 | 0.2086 | No |
| 52 | Siah1a | Siah1a | 15922 | -0.096 | 0.2100 | No |
| 53 | Gadd45a | Gadd45a | 16008 | -0.099 | 0.2110 | No |
| 54 | Cop1 | Cop1 | 16649 | -0.123 | 0.1909 | No |
| 55 | Casp3 | Casp3 | 17391 | -0.152 | 0.1681 | No |
| 56 | Cyct | Cyct | 18739 | -0.210 | 0.1237 | No |
| 57 | Siah1b | Siah1b | 19313 | -0.235 | 0.1113 | No |
| 58 | Cd82 | Cd82 | 20541 | -0.300 | 0.0757 | No |
| 59 | Cycs | Cycs | 21589 | -0.368 | 0.0503 | No |
| 60 | Ccne2 | Ccne2 | 22352 | -0.428 | 0.0389 | No |
| 61 | Steap3 | Steap3 | 23039 | -0.501 | 0.0338 | No |
| 62 | Cdk2 | Cdk2 | 23131 | -0.512 | 0.0530 | No |
| 63 | Chek2 | Chek2 | 23598 | -0.580 | 0.0603 | No |