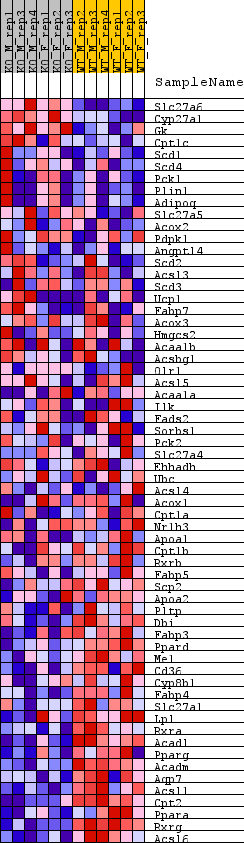
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | KO_vs_WT.KO_vs_WT.cls#KO_versus_WT.KO_vs_WT.cls#KO_versus_WT_repos |
| Phenotype | KO_vs_WT.cls#KO_versus_WT_repos |
| Upregulated in class | WT |
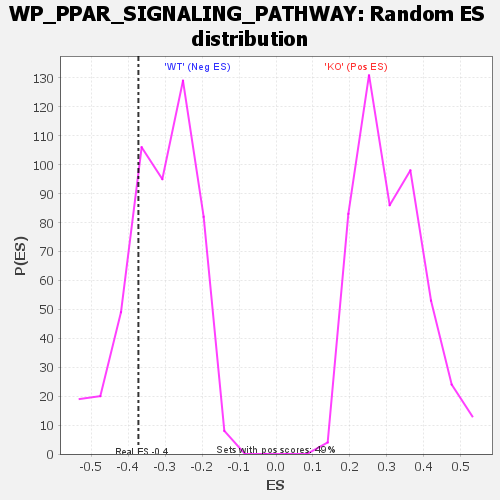
| GeneSet | WP_PPAR_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.37210858 |
| Normalized Enrichment Score (NES) | -1.2023213 |
| Nominal p-value | 0.24409449 |
| FDR q-value | 0.79387873 |
| FWER p-Value | 0.98 |

| SYMBOL | TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|
| 1 | Slc27a6 | Slc27a6 | 375 | 1.227 | 0.0409 | No |
| 2 | Cyp27a1 | Cyp27a1 | 918 | 0.904 | 0.0604 | No |
| 3 | Gk | Gk | 1922 | 0.662 | 0.0506 | No |
| 4 | Cpt1c | Cpt1c | 2775 | 0.545 | 0.0414 | No |
| 5 | Scd1 | Scd1 | 4076 | 0.423 | 0.0087 | No |
| 6 | Scd4 | Scd4 | 4279 | 0.408 | 0.0192 | No |
| 7 | Pck1 | Pck1 | 4406 | 0.398 | 0.0323 | No |
| 8 | Plin1 | Plin1 | 4515 | 0.390 | 0.0458 | No |
| 9 | Adipoq | Adipoq | 5271 | 0.342 | 0.0312 | No |
| 10 | Slc27a5 | Slc27a5 | 5441 | 0.331 | 0.0395 | No |
| 11 | Acox2 | Acox2 | 5467 | 0.329 | 0.0535 | No |
| 12 | Pdpk1 | Pdpk1 | 5983 | 0.299 | 0.0466 | No |
| 13 | Angptl4 | Angptl4 | 6512 | 0.269 | 0.0377 | No |
| 14 | Scd2 | Scd2 | 7871 | 0.205 | -0.0072 | No |
| 15 | Acsl3 | Acsl3 | 7938 | 0.201 | -0.0006 | No |
| 16 | Scd3 | Scd3 | 8129 | 0.193 | 0.0006 | No |
| 17 | Ucp1 | Ucp1 | 8851 | 0.164 | -0.0207 | No |
| 18 | Fabp7 | Fabp7 | 9413 | 0.142 | -0.0367 | No |
| 19 | Acox3 | Acox3 | 10909 | 0.083 | -0.0926 | No |
| 20 | Hmgcs2 | Hmgcs2 | 11118 | 0.075 | -0.0975 | No |
| 21 | Acaa1b | Acaa1b | 12362 | 0.031 | -0.1457 | No |
| 22 | Acsbg1 | Acsbg1 | 12521 | 0.026 | -0.1508 | No |
| 23 | Olr1 | Olr1 | 12903 | 0.012 | -0.1655 | No |
| 24 | Acsl5 | Acsl5 | 13404 | -0.005 | -0.1852 | No |
| 25 | Acaa1a | Acaa1a | 13587 | -0.012 | -0.1919 | No |
| 26 | Ilk | Ilk | 13873 | -0.022 | -0.2023 | No |
| 27 | Fads2 | Fads2 | 13952 | -0.025 | -0.2043 | No |
| 28 | Sorbs1 | Sorbs1 | 14018 | -0.027 | -0.2056 | No |
| 29 | Pck2 | Pck2 | 14234 | -0.035 | -0.2126 | No |
| 30 | Slc27a4 | Slc27a4 | 14841 | -0.058 | -0.2342 | No |
| 31 | Ehhadh | Ehhadh | 15210 | -0.071 | -0.2456 | No |
| 32 | Ubc | Ubc | 16741 | -0.126 | -0.3010 | No |
| 33 | Acsl4 | Acsl4 | 17140 | -0.142 | -0.3104 | No |
| 34 | Acox1 | Acox1 | 17147 | -0.143 | -0.3041 | No |
| 35 | Cpt1a | Cpt1a | 17462 | -0.155 | -0.3096 | No |
| 36 | Nr1h3 | Nr1h3 | 17700 | -0.165 | -0.3116 | No |
| 37 | Apoa1 | Apoa1 | 18823 | -0.214 | -0.3466 | No |
| 38 | Cpt1b | Cpt1b | 19008 | -0.222 | -0.3439 | No |
| 39 | Rxrb | Rxrb | 19588 | -0.249 | -0.3557 | No |
| 40 | Fabp5 | Fabp5 | 19906 | -0.266 | -0.3562 | No |
| 41 | Scp2 | Scp2 | 19998 | -0.271 | -0.3475 | No |
| 42 | Apoa2 | Apoa2 | 20615 | -0.305 | -0.3582 | Yes |
| 43 | Pltp | Pltp | 20809 | -0.315 | -0.3516 | Yes |
| 44 | Dbi | Dbi | 21086 | -0.333 | -0.3474 | Yes |
| 45 | Fabp3 | Fabp3 | 21341 | -0.349 | -0.3417 | Yes |
| 46 | Ppard | Ppard | 21841 | -0.386 | -0.3440 | Yes |
| 47 | Me1 | Me1 | 22164 | -0.412 | -0.3381 | Yes |
| 48 | Cd36 | Cd36 | 22440 | -0.436 | -0.3293 | Yes |
| 49 | Cyp8b1 | Cyp8b1 | 22524 | -0.445 | -0.3123 | Yes |
| 50 | Fabp4 | Fabp4 | 23014 | -0.499 | -0.3091 | Yes |
| 51 | Slc27a1 | Slc27a1 | 23240 | -0.528 | -0.2941 | Yes |
| 52 | Lpl | Lpl | 23266 | -0.531 | -0.2709 | Yes |
| 53 | Rxra | Rxra | 23278 | -0.532 | -0.2471 | Yes |
| 54 | Acadl | Acadl | 23626 | -0.584 | -0.2344 | Yes |
| 55 | Pparg | Pparg | 23780 | -0.615 | -0.2125 | Yes |
| 56 | Acadm | Acadm | 24213 | -0.728 | -0.1966 | Yes |
| 57 | Aqp7 | Aqp7 | 24240 | -0.737 | -0.1640 | Yes |
| 58 | Acsl1 | Acsl1 | 24244 | -0.739 | -0.1305 | Yes |
| 59 | Cpt2 | Cpt2 | 24299 | -0.759 | -0.0981 | Yes |
| 60 | Ppara | Ppara | 24360 | -0.778 | -0.0651 | Yes |
| 61 | Rxrg | Rxrg | 24710 | -0.973 | -0.0347 | Yes |
| 62 | Acsl6 | Acsl6 | 24817 | -1.109 | 0.0116 | Yes |