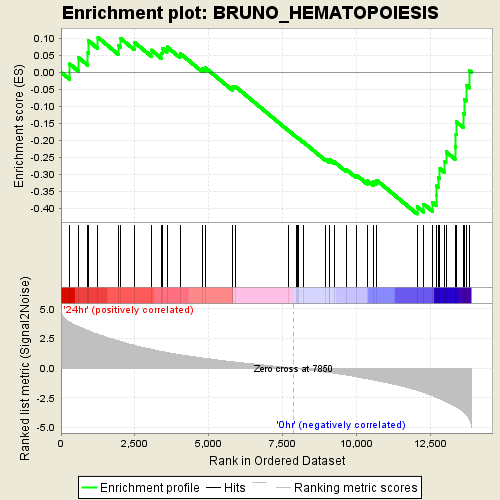
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #24hr_versus_0hr.class.cls #24hr_versus_0hr_repos |
| Phenotype | class.cls#24hr_versus_0hr_repos |
| Upregulated in class | 0hr |
| GeneSet | BRUNO_HEMATOPOIESIS |
| Enrichment Score (ES) | -0.41559297 |
| Normalized Enrichment Score (NES) | -1.5819143 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.941 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RCN1 | RCN1 Entrez, Source | reticulocalbin 1, EF-hand calcium binding domain | 288 | 3.909 | 0.0243 | No |
| 2 | CCRN4L | CCRN4L Entrez, Source | CCR4 carbon catabolite repression 4-like (S. cerevisiae) | 591 | 3.497 | 0.0428 | No |
| 3 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 908 | 3.180 | 0.0567 | No |
| 4 | SQSTM1 | SQSTM1 Entrez, Source | sequestosome 1 | 919 | 3.171 | 0.0925 | No |
| 5 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 1247 | 2.855 | 0.1019 | No |
| 6 | ENAH | ENAH Entrez, Source | enabled homolog (Drosophila) | 1931 | 2.313 | 0.0792 | No |
| 7 | CAPN5 | CAPN5 Entrez, Source | calpain 5 | 2019 | 2.251 | 0.0989 | No |
| 8 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 2481 | 1.922 | 0.0877 | No |
| 9 | LAT2 | LAT2 Entrez, Source | linker for activation of T cells family, member 2 | 3048 | 1.591 | 0.0652 | No |
| 10 | CALCA | CALCA Entrez, Source | calcitonin/calcitonin-related polypeptide, alpha | 3399 | 1.405 | 0.0561 | No |
| 11 | HMGCR | HMGCR Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A reductase | 3435 | 1.389 | 0.0696 | No |
| 12 | REPS1 | REPS1 Entrez, Source | RALBP1 associated Eps domain containing 1 | 3584 | 1.323 | 0.0742 | No |
| 13 | AK1 | AK1 Entrez, Source | adenylate kinase 1 | 4025 | 1.129 | 0.0554 | No |
| 14 | ELF1 | ELF1 Entrez, Source | E74-like factor 1 (ets domain transcription factor) | 4776 | 0.855 | 0.0111 | No |
| 15 | HK2 | HK2 Entrez, Source | hexokinase 2 | 4887 | 0.823 | 0.0126 | No |
| 16 | GLB1 | GLB1 Entrez, Source | galactosidase, beta 1 | 5799 | 0.539 | -0.0470 | No |
| 17 | SPINT1 | SPINT1 Entrez, Source | serine peptidase inhibitor, Kunitz type 1 | 5809 | 0.536 | -0.0415 | No |
| 18 | FKBP1A | FKBP1A Entrez, Source | FK506 binding protein 1A, 12kDa | 5901 | 0.507 | -0.0422 | No |
| 19 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 7686 | 0.044 | -0.1706 | No |
| 20 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 7979 | -0.034 | -0.1913 | No |
| 21 | S100A6 | S100A6 Entrez, Source | S100 calcium binding protein A6 | 7981 | -0.035 | -0.1910 | No |
| 22 | TJP1 | TJP1 Entrez, Source | tight junction protein 1 (zona occludens 1) | 8044 | -0.055 | -0.1949 | No |
| 23 | SESN1 | SESN1 Entrez, Source | sestrin 1 | 8184 | -0.094 | -0.2038 | No |
| 24 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 8955 | -0.334 | -0.2556 | No |
| 25 | CPE | CPE Entrez, Source | carboxypeptidase E | 9077 | -0.374 | -0.2600 | No |
| 26 | SNAP23 | SNAP23 Entrez, Source | synaptosomal-associated protein, 23kDa | 9091 | -0.378 | -0.2566 | No |
| 27 | NOV | NOV Entrez, Source | nephroblastoma overexpressed gene | 9236 | -0.424 | -0.2621 | No |
| 28 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 9647 | -0.574 | -0.2851 | No |
| 29 | CTSC | CTSC Entrez, Source | cathepsin C | 9990 | -0.719 | -0.3016 | No |
| 30 | FGF3 | FGF3 Entrez, Source | fibroblast growth factor 3 (murine mammary tumor virus integration site (v-int-2) oncogene homolog) | 10363 | -0.906 | -0.3180 | No |
| 31 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 10571 | -1.006 | -0.3214 | No |
| 32 | ZFP106 | ZFP106 Entrez, Source | zinc finger protein 106 homolog (mouse) | 10673 | -1.053 | -0.3165 | No |
| 33 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 12045 | -1.872 | -0.3940 | Yes |
| 34 | SERPINF1 | SERPINF1 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 | 12270 | -2.054 | -0.3865 | Yes |
| 35 | CD9 | CD9 Entrez, Source | CD9 molecule | 12574 | -2.333 | -0.3815 | Yes |
| 36 | CCL3 | CCL3 Entrez, Source | chemokine (C-C motif) ligand 3 | 12690 | -2.456 | -0.3614 | Yes |
| 37 | DAG1 | DAG1 Entrez, Source | dystroglycan 1 (dystrophin-associated glycoprotein 1) | 12693 | -2.459 | -0.3332 | Yes |
| 38 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 12758 | -2.533 | -0.3086 | Yes |
| 39 | VAMP5 | VAMP5 Entrez, Source | vesicle-associated membrane protein 5 (myobrevin) | 12816 | -2.592 | -0.2828 | Yes |
| 40 | MAP3K1 | MAP3K1 Entrez, Source | mitogen-activated protein kinase kinase kinase 1 | 12981 | -2.769 | -0.2627 | Yes |
| 41 | GALK1 | GALK1 Entrez, Source | galactokinase 1 | 13022 | -2.819 | -0.2330 | Yes |
| 42 | BCL2 | BCL2 Entrez, Source | B-cell CLL/lymphoma 2 | 13329 | -3.226 | -0.2179 | Yes |
| 43 | CCNG2 | CCNG2 Entrez, Source | cyclin G2 | 13358 | -3.262 | -0.1823 | Yes |
| 44 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 13365 | -3.270 | -0.1450 | Yes |
| 45 | SLC39A6 | SLC39A6 Entrez, Source | solute carrier family 39 (zinc transporter), member 6 | 13613 | -3.687 | -0.1203 | Yes |
| 46 | RBL2 | RBL2 Entrez, Source | retinoblastoma-like 2 (p130) | 13650 | -3.770 | -0.0794 | Yes |
| 47 | PBX1 | PBX1 Entrez, Source | pre-B-cell leukemia transcription factor 1 | 13720 | -3.957 | -0.0387 | Yes |
| 48 | ST6GAL1 | ST6GAL1 Entrez, Source | ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 | 13827 | -4.370 | 0.0040 | Yes |

