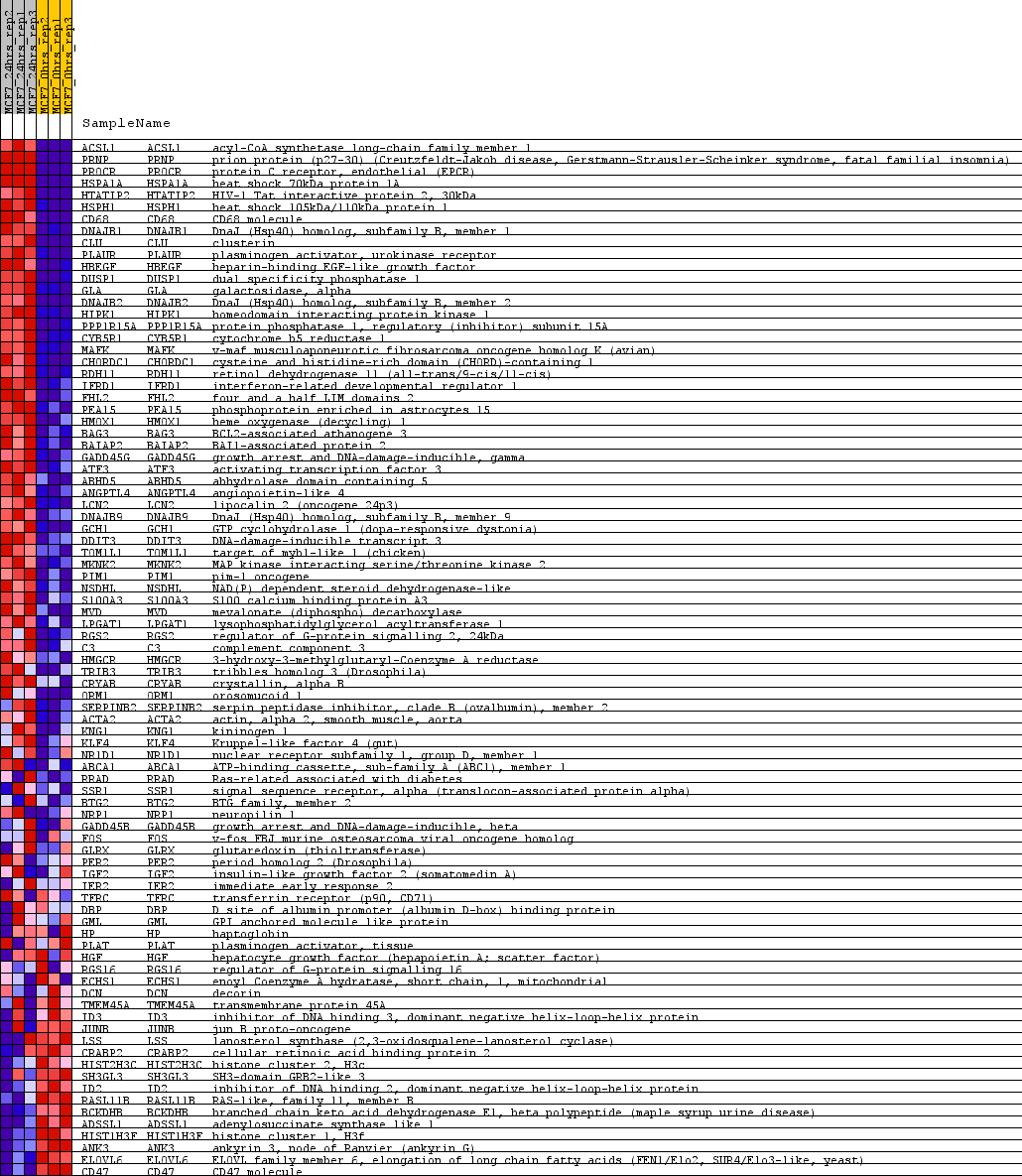
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #24hr_versus_0hr.class.cls #24hr_versus_0hr_repos |
| Phenotype | class.cls#24hr_versus_0hr_repos |
| Upregulated in class | 24hr |
| GeneSet | GERY_CEBP_TARGETS |
| Enrichment Score (ES) | 0.5395381 |
| Normalized Enrichment Score (NES) | 1.7443682 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.48896796 |
| FWER p-Value | 0.751 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 21 | 4.585 | 0.0249 | Yes |
| 2 | PRNP | PRNP Entrez, Source | prion protein (p27-30) (Creutzfeldt-Jakob disease, Gerstmann-Strausler-Scheinker syndrome, fatal familial insomnia) | 46 | 4.474 | 0.0490 | Yes |
| 3 | PROCR | PROCR Entrez, Source | protein C receptor, endothelial (EPCR) | 54 | 4.447 | 0.0741 | Yes |
| 4 | HSPA1A | HSPA1A Entrez, Source | heat shock 70kDa protein 1A | 101 | 4.293 | 0.0955 | Yes |
| 5 | HTATIP2 | HTATIP2 Entrez, Source | HIV-1 Tat interactive protein 2, 30kDa | 151 | 4.157 | 0.1159 | Yes |
| 6 | HSPH1 | HSPH1 Entrez, Source | heat shock 105kDa/110kDa protein 1 | 210 | 4.014 | 0.1349 | Yes |
| 7 | CD68 | CD68 Entrez, Source | CD68 molecule | 215 | 4.007 | 0.1577 | Yes |
| 8 | DNAJB1 | DNAJB1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 1 | 248 | 3.967 | 0.1782 | Yes |
| 9 | CLU | CLU Entrez, Source | clusterin | 299 | 3.889 | 0.1970 | Yes |
| 10 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 316 | 3.859 | 0.2181 | Yes |
| 11 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 384 | 3.748 | 0.2349 | Yes |
| 12 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 417 | 3.702 | 0.2539 | Yes |
| 13 | GLA | GLA Entrez, Source | galactosidase, alpha | 492 | 3.618 | 0.2694 | Yes |
| 14 | DNAJB2 | DNAJB2 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 2 | 508 | 3.596 | 0.2891 | Yes |
| 15 | HIPK1 | HIPK1 Entrez, Source | homeodomain interacting protein kinase 1 | 513 | 3.593 | 0.3095 | Yes |
| 16 | PPP1R15A | PPP1R15A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 15A | 515 | 3.592 | 0.3301 | Yes |
| 17 | CYB5R1 | CYB5R1 Entrez, Source | cytochrome b5 reductase 1 | 633 | 3.465 | 0.3416 | Yes |
| 18 | MAFK | MAFK Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog K (avian) | 713 | 3.394 | 0.3555 | Yes |
| 19 | CHORDC1 | CHORDC1 Entrez, Source | cysteine and histidine-rich domain (CHORD)-containing 1 | 832 | 3.259 | 0.3657 | Yes |
| 20 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 866 | 3.224 | 0.3819 | Yes |
| 21 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 1017 | 3.050 | 0.3886 | Yes |
| 22 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 1099 | 2.975 | 0.3999 | Yes |
| 23 | PEA15 | PEA15 Entrez, Source | phosphoprotein enriched in astrocytes 15 | 1238 | 2.866 | 0.4064 | Yes |
| 24 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 1474 | 2.658 | 0.4047 | Yes |
| 25 | BAG3 | BAG3 Entrez, Source | BCL2-associated athanogene 3 | 1565 | 2.576 | 0.4130 | Yes |
| 26 | BAIAP2 | BAIAP2 Entrez, Source | BAI1-associated protein 2 | 1592 | 2.562 | 0.4259 | Yes |
| 27 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 1624 | 2.537 | 0.4383 | Yes |
| 28 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1699 | 2.474 | 0.4472 | Yes |
| 29 | ABHD5 | ABHD5 Entrez, Source | abhydrolase domain containing 5 | 1726 | 2.461 | 0.4595 | Yes |
| 30 | ANGPTL4 | ANGPTL4 Entrez, Source | angiopoietin-like 4 | 1817 | 2.392 | 0.4668 | Yes |
| 31 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 1871 | 2.351 | 0.4765 | Yes |
| 32 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 1904 | 2.326 | 0.4876 | Yes |
| 33 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 1913 | 2.322 | 0.5004 | Yes |
| 34 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 1970 | 2.287 | 0.5095 | Yes |
| 35 | TOM1L1 | TOM1L1 Entrez, Source | target of myb1-like 1 (chicken) | 2122 | 2.172 | 0.5111 | Yes |
| 36 | MKNK2 | MKNK2 Entrez, Source | MAP kinase interacting serine/threonine kinase 2 | 2224 | 2.095 | 0.5158 | Yes |
| 37 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 2274 | 2.055 | 0.5241 | Yes |
| 38 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 2467 | 1.930 | 0.5214 | Yes |
| 39 | S100A3 | S100A3 Entrez, Source | S100 calcium binding protein A3 | 2533 | 1.882 | 0.5275 | Yes |
| 40 | MVD | MVD Entrez, Source | mevalonate (diphospho) decarboxylase | 2544 | 1.877 | 0.5376 | Yes |
| 41 | LPGAT1 | LPGAT1 Entrez, Source | lysophosphatidylglycerol acyltransferase 1 | 2662 | 1.808 | 0.5395 | Yes |
| 42 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 3021 | 1.610 | 0.5229 | No |
| 43 | C3 | C3 Entrez, Source | complement component 3 | 3371 | 1.416 | 0.5057 | No |
| 44 | HMGCR | HMGCR Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A reductase | 3435 | 1.389 | 0.5092 | No |
| 45 | TRIB3 | TRIB3 Entrez, Source | tribbles homolog 3 (Drosophila) | 3475 | 1.374 | 0.5143 | No |
| 46 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 3609 | 1.313 | 0.5122 | No |
| 47 | ORM1 | ORM1 Entrez, Source | orosomucoid 1 | 3699 | 1.272 | 0.5131 | No |
| 48 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 3806 | 1.226 | 0.5125 | No |
| 49 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 4004 | 1.137 | 0.5048 | No |
| 50 | KNG1 | KNG1 Entrez, Source | kininogen 1 | 4023 | 1.130 | 0.5100 | No |
| 51 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 4580 | 0.928 | 0.4750 | No |
| 52 | NR1D1 | NR1D1 Entrez, Source | nuclear receptor subfamily 1, group D, member 1 | 5000 | 0.785 | 0.4492 | No |
| 53 | ABCA1 | ABCA1 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 1 | 5146 | 0.740 | 0.4429 | No |
| 54 | RRAD | RRAD Entrez, Source | Ras-related associated with diabetes | 5558 | 0.607 | 0.4166 | No |
| 55 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 5705 | 0.566 | 0.4093 | No |
| 56 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 5873 | 0.516 | 0.4002 | No |
| 57 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 6005 | 0.484 | 0.3935 | No |
| 58 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 6322 | 0.398 | 0.3729 | No |
| 59 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 6728 | 0.302 | 0.3453 | No |
| 60 | GLRX | GLRX Entrez, Source | glutaredoxin (thioltransferase) | 6991 | 0.229 | 0.3276 | No |
| 61 | PER2 | PER2 Entrez, Source | period homolog 2 (Drosophila) | 7152 | 0.185 | 0.3171 | No |
| 62 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 7311 | 0.145 | 0.3065 | No |
| 63 | IER2 | IER2 Entrez, Source | immediate early response 2 | 7430 | 0.115 | 0.2986 | No |
| 64 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 7814 | 0.011 | 0.2709 | No |
| 65 | DBP | DBP Entrez, Source | D site of albumin promoter (albumin D-box) binding protein | 7934 | -0.025 | 0.2624 | No |
| 66 | GML | GML Entrez, Source | GPI anchored molecule like protein | 8014 | -0.045 | 0.2569 | No |
| 67 | HP | HP Entrez, Source | haptoglobin | 8248 | -0.109 | 0.2407 | No |
| 68 | PLAT | PLAT Entrez, Source | plasminogen activator, tissue | 8509 | -0.185 | 0.2229 | No |
| 69 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 8581 | -0.207 | 0.2189 | No |
| 70 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 8658 | -0.231 | 0.2148 | No |
| 71 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 9221 | -0.417 | 0.1764 | No |
| 72 | DCN | DCN Entrez, Source | decorin | 9277 | -0.436 | 0.1750 | No |
| 73 | TMEM45A | TMEM45A Entrez, Source | transmembrane protein 45A | 9483 | -0.514 | 0.1631 | No |
| 74 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 9773 | -0.632 | 0.1458 | No |
| 75 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 10192 | -0.822 | 0.1202 | No |
| 76 | LSS | LSS Entrez, Source | lanosterol synthase (2,3-oxidosqualene-lanosterol cyclase) | 10430 | -0.936 | 0.1084 | No |
| 77 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 10449 | -0.946 | 0.1126 | No |
| 78 | HIST2H3C | HIST2H3C Entrez, Source | histone cluster 2, H3c | 10536 | -0.987 | 0.1120 | No |
| 79 | SH3GL3 | SH3GL3 Entrez, Source | SH3-domain GRB2-like 3 | 10802 | -1.126 | 0.0993 | No |
| 80 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 11547 | -1.538 | 0.0543 | No |
| 81 | RASL11B | RASL11B Entrez, Source | RAS-like, family 11, member B | 12282 | -2.064 | 0.0129 | No |
| 82 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 12491 | -2.250 | 0.0108 | No |
| 83 | ADSSL1 | ADSSL1 Entrez, Source | adenylosuccinate synthase like 1 | 12807 | -2.580 | 0.0029 | No |
| 84 | HIST1H3F | HIST1H3F Entrez, Source | histone cluster 1, H3f | 12875 | -2.651 | 0.0133 | No |
| 85 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 13101 | -2.932 | 0.0139 | No |
| 86 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 13365 | -3.270 | 0.0137 | No |
| 87 | CD47 | CD47 Entrez, Source | CD47 molecule | 13767 | -4.120 | 0.0084 | No |