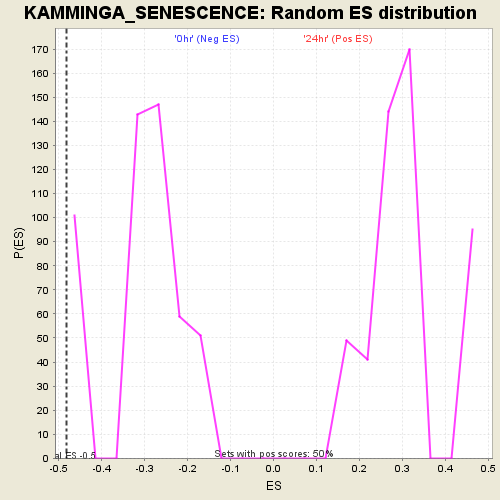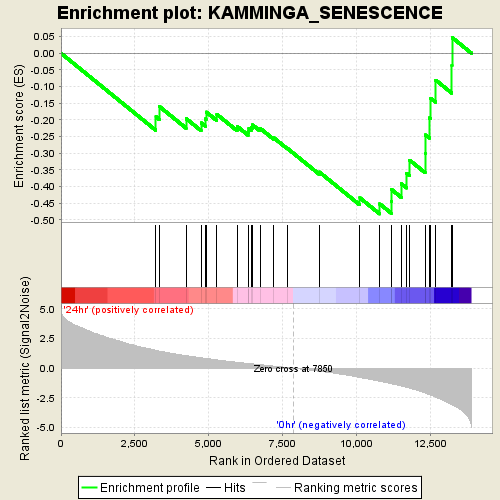
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #24hr_versus_0hr.class.cls #24hr_versus_0hr_repos |
| Phenotype | class.cls#24hr_versus_0hr_repos |
| Upregulated in class | 0hr |
| GeneSet | KAMMINGA_SENESCENCE |
| Enrichment Score (ES) | -0.48255455 |
| Normalized Enrichment Score (NES) | -1.5737627 |
| Nominal p-value | 0.105788425 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.941 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FHIT | FHIT Entrez, Source | fragile histidine triad gene | 3210 | 1.503 | -0.1906 | No |
| 2 | NOL9 | NOL9 Entrez, Source | nucleolar protein 9 | 3328 | 1.435 | -0.1599 | No |
| 3 | GNAO1 | GNAO1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O | 4230 | 1.058 | -0.1960 | No |
| 4 | CCAR1 | CCAR1 Entrez, Source | cell division cycle and apoptosis regulator 1 | 4739 | 0.869 | -0.2089 | No |
| 5 | CPN1 | CPN1 Entrez, Source | carboxypeptidase N, polypeptide 1, 50kD | 4886 | 0.823 | -0.1970 | No |
| 6 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 4909 | 0.816 | -0.1763 | No |
| 7 | QTRT1 | QTRT1 Entrez, Source | queuine tRNA-ribosyltransferase 1 (tRNA-guanine transglycosylase) | 5270 | 0.694 | -0.1833 | No |
| 8 | MLLT6 | MLLT6 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 6 | 5957 | 0.494 | -0.2194 | No |
| 9 | CLK2 | CLK2 Entrez, Source | CDC-like kinase 2 | 6327 | 0.397 | -0.2352 | No |
| 10 | ABHD8 | ABHD8 Entrez, Source | abhydrolase domain containing 8 | 6353 | 0.391 | -0.2263 | No |
| 11 | DDX54 | DDX54 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 54 | 6432 | 0.373 | -0.2217 | No |
| 12 | WHSC2 | WHSC2 Entrez, Source | Wolf-Hirschhorn syndrome candidate 2 | 6476 | 0.362 | -0.2149 | No |
| 13 | DHRSX | DHRSX Entrez, Source | dehydrogenase/reductase (SDR family) X-linked | 6731 | 0.301 | -0.2250 | No |
| 14 | GRP | GRP Entrez, Source | gastrin-releasing peptide | 7193 | 0.175 | -0.2535 | No |
| 15 | RABAC1 | RABAC1 Entrez, Source | Rab acceptor 1 (prenylated) | 7663 | 0.050 | -0.2860 | No |
| 16 | ARID1A | ARID1A Entrez, Source | AT rich interactive domain 1A (SWI- like) | 8734 | -0.254 | -0.3563 | No |
| 17 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 10109 | -0.782 | -0.4341 | No |
| 18 | PLEKHA7 | PLEKHA7 Entrez, Source | pleckstrin homology domain containing, family A member 7 | 10781 | -1.111 | -0.4522 | Yes |
| 19 | GTF2E1 | GTF2E1 Entrez, Source | general transcription factor IIE, polypeptide 1, alpha 56kDa | 11171 | -1.312 | -0.4444 | Yes |
| 20 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 11183 | -1.317 | -0.4092 | Yes |
| 21 | ELAVL1 | ELAVL1 Entrez, Source | ELAV (embryonic lethal, abnormal vision, Drosophila)-like 1 (Hu antigen R) | 11511 | -1.510 | -0.3916 | Yes |
| 22 | MKI67 | MKI67 Entrez, Source | antigen identified by monoclonal antibody Ki-67 | 11692 | -1.628 | -0.3601 | Yes |
| 23 | ALG10 | ALG10 Entrez, Source | asparagine-linked glycosylation 10 homolog (yeast, alpha-1,2-glucosyltransferase) | 11791 | -1.693 | -0.3209 | Yes |
| 24 | TCEA3 | TCEA3 Entrez, Source | transcription elongation factor A (SII), 3 | 12317 | -2.089 | -0.3017 | Yes |
| 25 | SUOX | SUOX Entrez, Source | sulfite oxidase | 12326 | -2.098 | -0.2450 | Yes |
| 26 | THBS4 | THBS4 Entrez, Source | thrombospondin 4 | 12464 | -2.230 | -0.1940 | Yes |
| 27 | EZH2 | EZH2 Entrez, Source | enhancer of zeste homolog 2 (Drosophila) | 12505 | -2.264 | -0.1350 | Yes |
| 28 | ARHGEF10L | ARHGEF10L Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 10-like | 12672 | -2.439 | -0.0804 | Yes |
| 29 | HMGN1 | HMGN1 Entrez, Source | high-mobility group nucleosome binding domain 1 | 13221 | -3.067 | -0.0361 | Yes |
| 30 | PLEKHG2 | PLEKHG2 Entrez, Source | pleckstrin homology domain containing, family G (with RhoGef domain) member 2 | 13226 | -3.069 | 0.0474 | Yes |