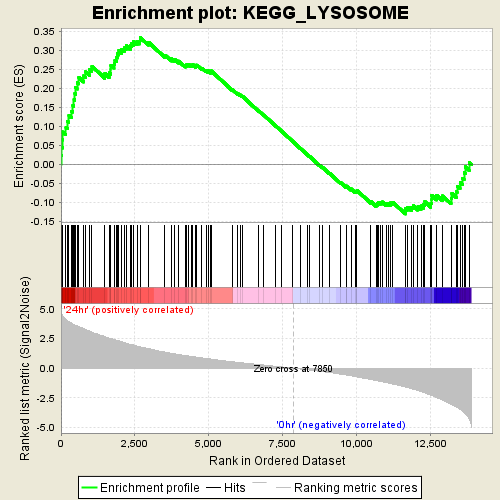
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #24hr_versus_0hr.class.cls #24hr_versus_0hr_repos |
| Phenotype | class.cls#24hr_versus_0hr_repos |
| Upregulated in class | 24hr |
| GeneSet | KEGG_LYSOSOME |
| Enrichment Score (ES) | 0.33479238 |
| Normalized Enrichment Score (NES) | 1.6755824 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.68584675 |
| FWER p-Value | 0.733 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NPC1 | NPC1 Entrez, Source | Niemann-Pick disease, type C1 | 1 | 4.884 | 0.0236 | Yes |
| 2 | AP4B1 | AP4B1 Entrez, Source | adaptor-related protein complex 4, beta 1 subunit | 15 | 4.674 | 0.0453 | Yes |
| 3 | AP1S2 | AP1S2 Entrez, Source | adaptor-related protein complex 1, sigma 2 subunit | 43 | 4.484 | 0.0651 | Yes |
| 4 | AP4E1 | AP4E1 Entrez, Source | adaptor-related protein complex 4, epsilon 1 subunit | 53 | 4.452 | 0.0860 | Yes |
| 5 | CLTB | CLTB Entrez, Source | clathrin, light chain (Lcb) | 164 | 4.096 | 0.0979 | Yes |
| 6 | CD68 | CD68 Entrez, Source | CD68 molecule | 215 | 4.007 | 0.1137 | Yes |
| 7 | CTSE | CTSE Entrez, Source | cathepsin E | 265 | 3.944 | 0.1292 | Yes |
| 8 | LGMN | LGMN Entrez, Source | legumain | 357 | 3.790 | 0.1410 | Yes |
| 9 | AP1G1 | AP1G1 Entrez, Source | adaptor-related protein complex 1, gamma 1 subunit | 398 | 3.728 | 0.1562 | Yes |
| 10 | AP3B2 | AP3B2 Entrez, Source | adaptor-related protein complex 3, beta 2 subunit | 436 | 3.673 | 0.1713 | Yes |
| 11 | NAGLU | NAGLU Entrez, Source | N-acetylglucosaminidase, alpha- (Sanfilippo disease IIIB) | 467 | 3.643 | 0.1868 | Yes |
| 12 | GLA | GLA Entrez, Source | galactosidase, alpha | 492 | 3.618 | 0.2026 | Yes |
| 13 | ENTPD4 | ENTPD4 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 4 | 543 | 3.560 | 0.2162 | Yes |
| 14 | MAN2B1 | MAN2B1 Entrez, Source | mannosidase, alpha, class 2B, member 1 | 600 | 3.487 | 0.2290 | Yes |
| 15 | GBA | GBA Entrez, Source | glucosidase, beta; acid (includes glucosylceramidase) | 762 | 3.343 | 0.2335 | Yes |
| 16 | CD63 | CD63 Entrez, Source | CD63 molecule | 833 | 3.259 | 0.2443 | Yes |
| 17 | ACP2 | ACP2 Entrez, Source | acid phosphatase 2, lysosomal | 965 | 3.113 | 0.2498 | Yes |
| 18 | AP3S2 | AP3S2 Entrez, Source | adaptor-related protein complex 3, sigma 2 subunit | 1043 | 3.024 | 0.2589 | Yes |
| 19 | M6PR | M6PR Entrez, Source | mannose-6-phosphate receptor (cation dependent) | 1481 | 2.649 | 0.2400 | Yes |
| 20 | PSAP | PSAP Entrez, Source | prosaposin (variant Gaucher disease and variant metachromatic leukodystrophy) | 1628 | 2.533 | 0.2417 | Yes |
| 21 | ATP6V0A2 | ATP6V0A2 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a2 | 1683 | 2.486 | 0.2498 | Yes |
| 22 | AP4S1 | AP4S1 Entrez, Source | adaptor-related protein complex 4, sigma 1 subunit | 1685 | 2.485 | 0.2618 | Yes |
| 23 | SGSH | SGSH Entrez, Source | N-sulfoglucosamine sulfohydrolase (sulfamidase) | 1791 | 2.406 | 0.2659 | Yes |
| 24 | SCARB2 | SCARB2 Entrez, Source | scavenger receptor class B, member 2 | 1821 | 2.390 | 0.2753 | Yes |
| 25 | ATP6V0B | ATP6V0B Entrez, Source | ATPase, H+ transporting, lysosomal 21kDa, V0 subunit b | 1880 | 2.341 | 0.2825 | Yes |
| 26 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 1906 | 2.324 | 0.2919 | Yes |
| 27 | CTSW | CTSW Entrez, Source | cathepsin W (lymphopain) | 1954 | 2.294 | 0.2996 | Yes |
| 28 | IGF2R | IGF2R Entrez, Source | insulin-like growth factor 2 receptor | 2051 | 2.223 | 0.3034 | Yes |
| 29 | GGA3 | GGA3 Entrez, Source | golgi associated, gamma adaptin ear containing, ARF binding protein 3 | 2138 | 2.160 | 0.3077 | Yes |
| 30 | CLTC | CLTC Entrez, Source | clathrin, heavy chain (Hc) | 2202 | 2.108 | 0.3133 | Yes |
| 31 | CTSD | CTSD Entrez, Source | cathepsin D (lysosomal aspartyl peptidase) | 2344 | 2.015 | 0.3128 | Yes |
| 32 | ATP6V0A1 | ATP6V0A1 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a1 | 2388 | 1.987 | 0.3193 | Yes |
| 33 | HEXB | HEXB Entrez, Source | hexosaminidase B (beta polypeptide) | 2457 | 1.942 | 0.3238 | Yes |
| 34 | AP3M1 | AP3M1 Entrez, Source | adaptor-related protein complex 3, mu 1 subunit | 2573 | 1.861 | 0.3245 | Yes |
| 35 | IDS | IDS Entrez, Source | iduronate 2-sulfatase (Hunter syndrome) | 2670 | 1.803 | 0.3263 | Yes |
| 36 | AP3D1 | AP3D1 Entrez, Source | adaptor-related protein complex 3, delta 1 subunit | 2674 | 1.803 | 0.3348 | Yes |
| 37 | AP1S3 | AP1S3 Entrez, Source | adaptor-related protein complex 1, sigma 3 subunit | 2953 | 1.646 | 0.3226 | No |
| 38 | LAMP1 | LAMP1 Entrez, Source | lysosomal-associated membrane protein 1 | 3515 | 1.358 | 0.2885 | No |
| 39 | CTNS | CTNS Entrez, Source | cystinosis, nephropathic | 3733 | 1.259 | 0.2788 | No |
| 40 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 3842 | 1.211 | 0.2768 | No |
| 41 | AP1M1 | AP1M1 Entrez, Source | adaptor-related protein complex 1, mu 1 subunit | 3975 | 1.152 | 0.2728 | No |
| 42 | ATP6V1H | ATP6V1H Entrez, Source | ATPase, H+ transporting, lysosomal 50/57kDa, V1 subunit H | 4197 | 1.064 | 0.2620 | No |
| 43 | ATP6V0D1 | ATP6V0D1 Entrez, Source | ATPase, H+ transporting, lysosomal 38kDa, V0 subunit d1 | 4253 | 1.052 | 0.2631 | No |
| 44 | NEU1 | NEU1 Entrez, Source | sialidase 1 (lysosomal sialidase) | 4303 | 1.033 | 0.2645 | No |
| 45 | HEXA | HEXA Entrez, Source | hexosaminidase A (alpha polypeptide) | 4402 | 0.994 | 0.2622 | No |
| 46 | CTSZ | CTSZ Entrez, Source | cathepsin Z | 4456 | 0.976 | 0.2631 | No |
| 47 | LAMP3 | LAMP3 Entrez, Source | lysosomal-associated membrane protein 3 | 4563 | 0.934 | 0.2599 | No |
| 48 | SMPD1 | SMPD1 Entrez, Source | sphingomyelin phosphodiesterase 1, acid lysosomal (acid sphingomyelinase) | 4593 | 0.921 | 0.2623 | No |
| 49 | TPP1 | TPP1 Entrez, Source | tripeptidyl peptidase I | 4762 | 0.861 | 0.2543 | No |
| 50 | LAMP2 | LAMP2 Entrez, Source | lysosomal-associated membrane protein 2 | 4906 | 0.817 | 0.2478 | No |
| 51 | CLTA | CLTA Entrez, Source | clathrin, light chain (Lca) | 4972 | 0.796 | 0.2470 | No |
| 52 | NAGPA | NAGPA Entrez, Source | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase | 5061 | 0.768 | 0.2443 | No |
| 53 | SLC17A5 | SLC17A5 Entrez, Source | solute carrier family 17 (anion/sugar transporter), member 5 | 5078 | 0.761 | 0.2468 | No |
| 54 | GLB1 | GLB1 Entrez, Source | galactosidase, beta 1 | 5799 | 0.539 | 0.1972 | No |
| 55 | SLC11A1 | SLC11A1 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 1 | 5967 | 0.492 | 0.1875 | No |
| 56 | HYAL1 | HYAL1 Entrez, Source | hyaluronoglucosaminidase 1 | 6056 | 0.469 | 0.1833 | No |
| 57 | GALNS | GALNS Entrez, Source | galactosamine (N-acetyl)-6-sulfate sulfatase (Morquio syndrome, mucopolysaccharidosis type IVA) | 6130 | 0.452 | 0.1802 | No |
| 58 | CTSG | CTSG Entrez, Source | cathepsin G | 6693 | 0.310 | 0.1409 | No |
| 59 | PPT2 | PPT2 Entrez, Source | palmitoyl-protein thioesterase 2 | 6860 | 0.267 | 0.1302 | No |
| 60 | MCOLN1 | MCOLN1 Entrez, Source | mucolipin 1 | 7268 | 0.158 | 0.1014 | No |
| 61 | GGA2 | GGA2 Entrez, Source | golgi associated, gamma adaptin ear containing, ARF binding protein 2 | 7459 | 0.105 | 0.0881 | No |
| 62 | ARSB | ARSB Entrez, Source | arylsulfatase B | 7823 | 0.008 | 0.0618 | No |
| 63 | AP3S1 | AP3S1 Entrez, Source | adaptor-related protein complex 3, sigma 1 subunit | 8103 | -0.073 | 0.0419 | No |
| 64 | AGA | AGA Entrez, Source | aspartylglucosaminidase | 8321 | -0.127 | 0.0268 | No |
| 65 | CTSB | CTSB Entrez, Source | cathepsin B | 8414 | -0.157 | 0.0209 | No |
| 66 | ACP5 | ACP5 Entrez, Source | acid phosphatase 5, tartrate resistant | 8748 | -0.261 | -0.0020 | No |
| 67 | AP3B1 | AP3B1 Entrez, Source | adaptor-related protein complex 3, beta 1 subunit | 8847 | -0.293 | -0.0077 | No |
| 68 | MANBA | MANBA Entrez, Source | mannosidase, beta A, lysosomal | 9084 | -0.375 | -0.0231 | No |
| 69 | CLTCL1 | CLTCL1 Entrez, Source | clathrin, heavy chain-like 1 | 9451 | -0.502 | -0.0472 | No |
| 70 | TCIRG1 | TCIRG1 Entrez, Source | T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 subunit A3 | 9651 | -0.575 | -0.0588 | No |
| 71 | CD164 | CD164 Entrez, Source | CD164 molecule, sialomucin | 9658 | -0.577 | -0.0565 | No |
| 72 | CLN3 | CLN3 Entrez, Source | ceroid-lipofuscinosis, neuronal 3, juvenile (Batten, Spielmeyer-Vogt disease) | 9809 | -0.644 | -0.0643 | No |
| 73 | CTSL2 | CTSL2 Entrez, Source | cathepsin L2 | 9945 | -0.698 | -0.0707 | No |
| 74 | CTSC | CTSC Entrez, Source | cathepsin C | 9990 | -0.719 | -0.0704 | No |
| 75 | AP3M2 | AP3M2 Entrez, Source | adaptor-related protein complex 3, mu 2 subunit | 10010 | -0.734 | -0.0682 | No |
| 76 | AP1M2 | AP1M2 Entrez, Source | adaptor-related protein complex 1, mu 2 subunit | 10480 | -0.961 | -0.0976 | No |
| 77 | CLN5 | CLN5 Entrez, Source | ceroid-lipofuscinosis, neuronal 5 | 10657 | -1.044 | -0.1053 | No |
| 78 | AP1S1 | AP1S1 Entrez, Source | adaptor-related protein complex 1, sigma 1 subunit | 10696 | -1.066 | -0.1029 | No |
| 79 | LAPTM4A | LAPTM4A Entrez, Source | lysosomal-associated protein transmembrane 4 alpha | 10737 | -1.090 | -0.1005 | No |
| 80 | NAGA | NAGA Entrez, Source | N-acetylgalactosaminidase, alpha- | 10794 | -1.120 | -0.0992 | No |
| 81 | NPC2 | NPC2 Entrez, Source | Niemann-Pick disease, type C2 | 10863 | -1.157 | -0.0985 | No |
| 82 | ARSA | ARSA Entrez, Source | arylsulfatase A | 11002 | -1.224 | -0.1026 | No |
| 83 | ABCB9 | ABCB9 Entrez, Source | ATP-binding cassette, sub-family B (MDR/TAP), member 9 | 11073 | -1.257 | -0.1015 | No |
| 84 | CTSK | CTSK Entrez, Source | cathepsin K (pycnodysostosis) | 11154 | -1.303 | -0.1010 | No |
| 85 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 11222 | -1.338 | -0.0994 | No |
| 86 | LIPA | LIPA Entrez, Source | lipase A, lysosomal acid, cholesterol esterase (Wolman disease) | 11650 | -1.604 | -0.1226 | No |
| 87 | SORT1 | SORT1 Entrez, Source | sortilin 1 | 11654 | -1.607 | -0.1151 | No |
| 88 | AP4M1 | AP4M1 Entrez, Source | adaptor-related protein complex 4, mu 1 subunit | 11725 | -1.649 | -0.1122 | No |
| 89 | DNASE2 | DNASE2 Entrez, Source | deoxyribonuclease II, lysosomal | 11857 | -1.745 | -0.1132 | No |
| 90 | GALC | GALC Entrez, Source | galactosylceramidase | 11912 | -1.782 | -0.1085 | No |
| 91 | ATP6AP1 | ATP6AP1 Entrez, Source | ATPase, H+ transporting, lysosomal accessory protein 1 | 12073 | -1.894 | -0.1109 | No |
| 92 | SLC11A2 | SLC11A2 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 | 12178 | -1.981 | -0.1089 | No |
| 93 | PPT1 | PPT1 Entrez, Source | palmitoyl-protein thioesterase 1 (ceroid-lipofuscinosis, neuronal 1, infantile) | 12265 | -2.052 | -0.1052 | No |
| 94 | GNPTG | GNPTG Entrez, Source | N-acetylglucosamine-1-phosphate transferase, gamma subunit | 12306 | -2.081 | -0.0980 | No |
| 95 | GNPTAB | GNPTAB Entrez, Source | N-acetylglucosamine-1-phosphate transferase, alpha and beta subunits | 12509 | -2.268 | -0.1016 | No |
| 96 | LAPTM4B | LAPTM4B Entrez, Source | lysosomal associated protein transmembrane 4 beta | 12531 | -2.291 | -0.0921 | No |
| 97 | FUCA1 | FUCA1 Entrez, Source | fucosidase, alpha-L- 1, tissue | 12537 | -2.299 | -0.0813 | No |
| 98 | IDUA | IDUA Entrez, Source | iduronidase, alpha-L- | 12706 | -2.478 | -0.0815 | No |
| 99 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 12901 | -2.676 | -0.0826 | No |
| 100 | ABCA2 | ABCA2 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 2 | 13194 | -3.039 | -0.0890 | No |
| 101 | ATP6V0A4 | ATP6V0A4 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a4 | 13219 | -3.065 | -0.0759 | No |
| 102 | GGA1 | GGA1 Entrez, Source | golgi associated, gamma adaptin ear containing, ARF binding protein 1 | 13369 | -3.276 | -0.0709 | No |
| 103 | SUMF1 | SUMF1 Entrez, Source | sulfatase modifying factor 1 | 13399 | -3.310 | -0.0569 | No |
| 104 | CTSH | CTSH Entrez, Source | cathepsin H | 13498 | -3.474 | -0.0472 | No |
| 105 | GAA | GAA Entrez, Source | glucosidase, alpha; acid (Pompe disease, glycogen storage disease type II) | 13584 | -3.618 | -0.0358 | No |
| 106 | GUSB | GUSB Entrez, Source | glucuronidase, beta | 13636 | -3.736 | -0.0214 | No |
| 107 | CTSF | CTSF Entrez, Source | cathepsin F | 13675 | -3.827 | -0.0056 | No |
| 108 | NAPSA | NAPSA Entrez, Source | napsin A aspartic peptidase | 13807 | -4.263 | 0.0055 | No |

