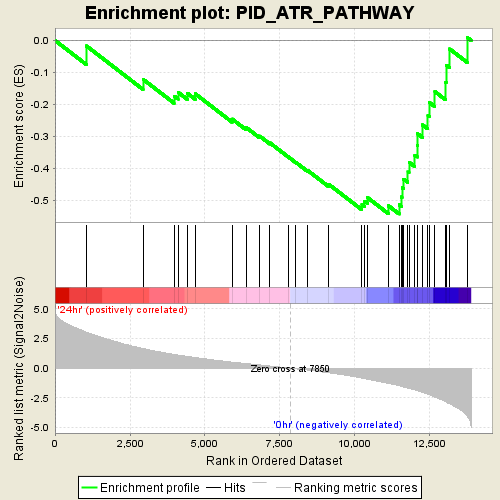
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #24hr_versus_0hr.class.cls #24hr_versus_0hr_repos |
| Phenotype | class.cls#24hr_versus_0hr_repos |
| Upregulated in class | 0hr |
| GeneSet | PID_ATR_PATHWAY |
| Enrichment Score (ES) | -0.5426616 |
| Normalized Enrichment Score (NES) | -1.4191134 |
| Nominal p-value | 0.18055555 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 1046 | 3.022 | -0.0170 | No |
| 2 | YWHAZ | YWHAZ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide | 2936 | 1.653 | -0.1214 | No |
| 3 | HUS1 | HUS1 Entrez, Source | HUS1 checkpoint homolog (S. pombe) | 3971 | 1.155 | -0.1737 | No |
| 4 | FBXW11 | FBXW11 Entrez, Source | F-box and WD-40 domain protein 11 | 4105 | 1.098 | -0.1620 | No |
| 5 | RAD1 | RAD1 Entrez, Source | RAD1 homolog (S. pombe) | 4411 | 0.991 | -0.1648 | No |
| 6 | PPP2CA | PPP2CA Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, alpha isoform | 4676 | 0.892 | -0.1666 | No |
| 7 | CEP164 | CEP164 Entrez, Source | centrosomal protein 164kDa | 5905 | 0.506 | -0.2455 | No |
| 8 | RAD9A | RAD9A Entrez, Source | RAD9 homolog A (S. pombe) | 6382 | 0.384 | -0.2724 | No |
| 9 | SMARCAL1 | SMARCAL1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a-like 1 | 6821 | 0.278 | -0.2987 | No |
| 10 | RPA2 | RPA2 Entrez, Source | replication protein A2, 32kDa | 7170 | 0.179 | -0.3204 | No |
| 11 | YWHAB | YWHAB Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide | 7774 | 0.021 | -0.3635 | No |
| 12 | PPP2R1A | PPP2R1A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), alpha isoform | 8034 | -0.051 | -0.3812 | No |
| 13 | RAD17 | RAD17 Entrez, Source | RAD17 homolog (S. pombe) | 8422 | -0.159 | -0.4061 | No |
| 14 | ATR | ATR Entrez, Source | ataxia telangiectasia and Rad3 related | 9121 | -0.388 | -0.4490 | No |
| 15 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 10238 | -0.846 | -0.5131 | No |
| 16 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 10326 | -0.885 | -0.5023 | No |
| 17 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 10419 | -0.930 | -0.4909 | No |
| 18 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 11124 | -1.284 | -0.5169 | Yes |
| 19 | FANCD2 | FANCD2 Entrez, Source | Fanconi anemia, complementation group D2 | 11482 | -1.496 | -0.5137 | Yes |
| 20 | CDC6 | CDC6 Entrez, Source | CDC6 cell division cycle 6 homolog (S. cerevisiae) | 11557 | -1.545 | -0.4891 | Yes |
| 21 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 11578 | -1.556 | -0.4604 | Yes |
| 22 | CDC25C | CDC25C Entrez, Source | cell division cycle 25C | 11628 | -1.588 | -0.4332 | Yes |
| 23 | RFC2 | RFC2 Entrez, Source | replication factor C (activator 1) 2, 40kDa | 11774 | -1.680 | -0.4111 | Yes |
| 24 | RAD51 | RAD51 Entrez, Source | RAD51 homolog (RecA homolog, E. coli) (S. cerevisiae) | 11810 | -1.703 | -0.3807 | Yes |
| 25 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 12003 | -1.841 | -0.3589 | Yes |
| 26 | RPA1 | RPA1 Entrez, Source | replication protein A1, 70kDa | 12081 | -1.900 | -0.3276 | Yes |
| 27 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 12094 | -1.910 | -0.2915 | Yes |
| 28 | TOPBP1 | TOPBP1 Entrez, Source | topoisomerase (DNA) II binding protein 1 | 12261 | -2.048 | -0.2638 | Yes |
| 29 | CLSPN | CLSPN Entrez, Source | claspin homolog (Xenopus laevis) | 12440 | -2.203 | -0.2340 | Yes |
| 30 | RFC5 | RFC5 Entrez, Source | replication factor C (activator 1) 5, 36.5kDa | 12486 | -2.247 | -0.1938 | Yes |
| 31 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 12673 | -2.441 | -0.1599 | Yes |
| 32 | RFC4 | RFC4 Entrez, Source | replication factor C (activator 1) 4, 37kDa | 13033 | -2.836 | -0.1309 | Yes |
| 33 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 13051 | -2.863 | -0.0767 | Yes |
| 34 | MCM2 | MCM2 Entrez, Source | MCM2 minichromosome maintenance deficient 2, mitotin (S. cerevisiae) | 13166 | -3.018 | -0.0265 | Yes |
| 35 | PPP2R2B | PPP2R2B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform | 13753 | -4.038 | 0.0094 | Yes |

