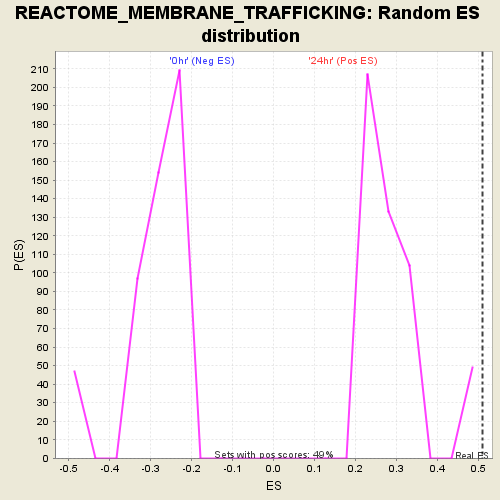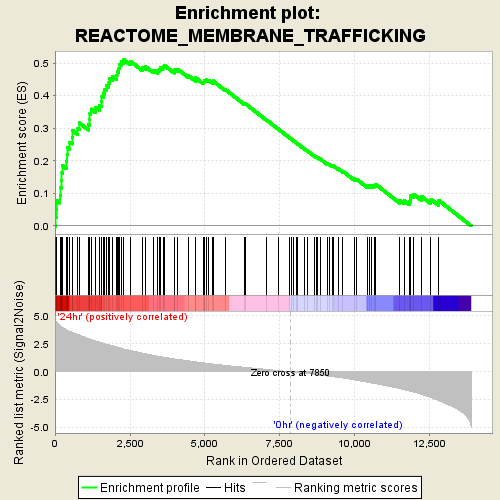
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #24hr_versus_0hr.class.cls #24hr_versus_0hr_repos |
| Phenotype | class.cls#24hr_versus_0hr_repos |
| Upregulated in class | 24hr |
| GeneSet | REACTOME_MEMBRANE_TRAFFICKING |
| Enrichment Score (ES) | 0.510523 |
| Normalized Enrichment Score (NES) | 1.7816616 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.80719745 |
| FWER p-Value | 0.491 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | AP4B1 | AP4B1 Entrez, Source | adaptor-related protein complex 4, beta 1 subunit | 15 | 4.674 | 0.0270 | Yes |
| 2 | AP1S2 | AP1S2 Entrez, Source | adaptor-related protein complex 1, sigma 2 subunit | 43 | 4.484 | 0.0520 | Yes |
| 3 | AP4E1 | AP4E1 Entrez, Source | adaptor-related protein complex 4, epsilon 1 subunit | 53 | 4.452 | 0.0782 | Yes |
| 4 | CLTB | CLTB Entrez, Source | clathrin, light chain (Lcb) | 164 | 4.096 | 0.0948 | Yes |
| 5 | VPS37A | VPS37A Entrez, Source | vacuolar protein sorting 37 homolog A (S. cerevisiae) | 194 | 4.044 | 0.1171 | Yes |
| 6 | FTL | FTL Entrez, Source | ferritin, light polypeptide | 207 | 4.017 | 0.1403 | Yes |
| 7 | HGS | HGS Entrez, Source | hepatocyte growth factor-regulated tyrosine kinase substrate | 229 | 3.991 | 0.1628 | Yes |
| 8 | YIPF6 | YIPF6 Entrez, Source | Yip1 domain family, member 6 | 249 | 3.966 | 0.1853 | Yes |
| 9 | COPA | COPA Entrez, Source | coatomer protein complex, subunit alpha | 379 | 3.752 | 0.1985 | Yes |
| 10 | AP1G1 | AP1G1 Entrez, Source | adaptor-related protein complex 1, gamma 1 subunit | 398 | 3.728 | 0.2196 | Yes |
| 11 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 413 | 3.704 | 0.2409 | Yes |
| 12 | STAM | STAM Entrez, Source | signal transducing adaptor molecule (SH3 domain and ITAM motif) 1 | 487 | 3.625 | 0.2574 | Yes |
| 13 | ARFGAP1 | ARFGAP1 Entrez, Source | ADP-ribosylation factor GTPase activating protein 1 | 570 | 3.520 | 0.2727 | Yes |
| 14 | SNX9 | SNX9 Entrez, Source | sorting nexin 9 | 582 | 3.500 | 0.2929 | Yes |
| 15 | DNM1 | DNM1 Entrez, Source | dynamin 1 | 760 | 3.344 | 0.3002 | Yes |
| 16 | SEC24D | SEC24D Entrez, Source | SEC24 related gene family, member D (S. cerevisiae) | 810 | 3.285 | 0.3164 | Yes |
| 17 | TPD52 | TPD52 Entrez, Source | tumor protein D52 | 1116 | 2.957 | 0.3121 | Yes |
| 18 | SH3D19 | SH3D19 Entrez, Source | - | 1157 | 2.925 | 0.3268 | Yes |
| 19 | GBF1 | GBF1 Entrez, Source | golgi-specific brefeldin A resistance factor 1 | 1161 | 2.921 | 0.3441 | Yes |
| 20 | PREB | PREB Entrez, Source | prolactin regulatory element binding | 1198 | 2.901 | 0.3589 | Yes |
| 21 | CHMP2B | CHMP2B Entrez, Source | chromatin modifying protein 2B | 1360 | 2.758 | 0.3639 | Yes |
| 22 | M6PR | M6PR Entrez, Source | mannose-6-phosphate receptor (cation dependent) | 1481 | 2.649 | 0.3711 | Yes |
| 23 | VPS4A | VPS4A Entrez, Source | vacuolar protein sorting 4 homolog A (S. cerevisiae) | 1546 | 2.593 | 0.3820 | Yes |
| 24 | ARCN1 | ARCN1 Entrez, Source | archain 1 | 1561 | 2.581 | 0.3966 | Yes |
| 25 | VPS37B | VPS37B Entrez, Source | vacuolar protein sorting 37 homolog B (S. cerevisiae) | 1612 | 2.547 | 0.4082 | Yes |
| 26 | NECAP1 | NECAP1 Entrez, Source | NECAP endocytosis associated 1 | 1661 | 2.502 | 0.4198 | Yes |
| 27 | CHMP5 | CHMP5 Entrez, Source | chromatin modifying protein 5 | 1707 | 2.469 | 0.4314 | Yes |
| 28 | RAB5C | RAB5C Entrez, Source | RAB5C, member RAS oncogene family | 1797 | 2.403 | 0.4394 | Yes |
| 29 | CHMP7 | CHMP7 Entrez, Source | CHMP family, member 7 | 1824 | 2.387 | 0.4519 | Yes |
| 30 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 1906 | 2.324 | 0.4600 | Yes |
| 31 | IGF2R | IGF2R Entrez, Source | insulin-like growth factor 2 receptor | 2051 | 2.223 | 0.4629 | Yes |
| 32 | ARF1 | ARF1 Entrez, Source | ADP-ribosylation factor 1 | 2068 | 2.212 | 0.4751 | Yes |
| 33 | CPD | CPD Entrez, Source | carboxypeptidase D | 2128 | 2.168 | 0.4838 | Yes |
| 34 | CHMP6 | CHMP6 Entrez, Source | chromatin modifying protein 6 | 2158 | 2.140 | 0.4946 | Yes |
| 35 | CLTC | CLTC Entrez, Source | clathrin, heavy chain (Hc) | 2202 | 2.108 | 0.5042 | Yes |
| 36 | FTH1 | FTH1 Entrez, Source | ferritin, heavy polypeptide 1 | 2285 | 2.047 | 0.5105 | Yes |
| 37 | CHMP4B | CHMP4B Entrez, Source | chromatin modifying protein 4B | 2507 | 1.908 | 0.5060 | No |
| 38 | STAM2 | STAM2 Entrez, Source | signal transducing adaptor molecule (SH3 domain and ITAM motif) 2 | 2911 | 1.669 | 0.4868 | No |
| 39 | SEC23A | SEC23A Entrez, Source | Sec23 homolog A (S. cerevisiae) | 3019 | 1.611 | 0.4887 | No |
| 40 | PIK3C2A | PIK3C2A Entrez, Source | phosphoinositide-3-kinase, class 2, alpha polypeptide | 3282 | 1.463 | 0.4785 | No |
| 41 | SEC24B | SEC24B Entrez, Source | SEC24 related gene family, member B (S. cerevisiae) | 3421 | 1.394 | 0.4769 | No |
| 42 | GJB3 | GJB3 Entrez, Source | gap junction protein, beta 3, 31kDa (connexin 31) | 3473 | 1.374 | 0.4814 | No |
| 43 | RPS27A | RPS27A Entrez, Source | ribosomal protein S27a | 3517 | 1.358 | 0.4865 | No |
| 44 | PLDN | PLDN Entrez, Source | pallidin homolog (mouse) | 3602 | 1.315 | 0.4883 | No |
| 45 | CHMP4C | CHMP4C Entrez, Source | chromatin modifying protein 4C | 3639 | 1.301 | 0.4935 | No |
| 46 | AP1M1 | AP1M1 Entrez, Source | adaptor-related protein complex 1, mu 1 subunit | 3975 | 1.152 | 0.4762 | No |
| 47 | VPS4B | VPS4B Entrez, Source | vacuolar protein sorting 4 homolog B (S. cerevisiae) | 3996 | 1.141 | 0.4816 | No |
| 48 | VPS25 | VPS25 Entrez, Source | vacuolar protein sorting 25 homolog (S. cerevisiae) | 4099 | 1.101 | 0.4808 | No |
| 49 | CTSZ | CTSZ Entrez, Source | cathepsin Z | 4456 | 0.976 | 0.4609 | No |
| 50 | NAPA | NAPA Entrez, Source | N-ethylmaleimide-sensitive factor attachment protein, alpha | 4684 | 0.890 | 0.4497 | No |
| 51 | UBA52 | UBA52 Entrez, Source | ubiquitin A-52 residue ribosomal protein fusion product 1 | 4694 | 0.886 | 0.4544 | No |
| 52 | COPZ1 | COPZ1 Entrez, Source | coatomer protein complex, subunit zeta 1 | 4938 | 0.803 | 0.4416 | No |
| 53 | CLTA | CLTA Entrez, Source | clathrin, light chain (Lca) | 4972 | 0.796 | 0.4440 | No |
| 54 | COPG | COPG Entrez, Source | coatomer protein complex, subunit gamma | 4986 | 0.792 | 0.4479 | No |
| 55 | GAK | GAK Entrez, Source | cyclin G associated kinase | 5036 | 0.776 | 0.4490 | No |
| 56 | COPB2 | COPB2 Entrez, Source | coatomer protein complex, subunit beta 2 (beta prime) | 5115 | 0.749 | 0.4478 | No |
| 57 | GJC1 | GJC1 Entrez, Source | gap junction protein, chi 1, 31.9kDa (connexin 31.9) | 5248 | 0.702 | 0.4425 | No |
| 58 | COPE | COPE Entrez, Source | coatomer protein complex, subunit epsilon | 5269 | 0.694 | 0.4452 | No |
| 59 | TXNDC5 | TXNDC5 Entrez, Source | thioredoxin domain containing 5 | 5687 | 0.572 | 0.4184 | No |
| 60 | VAMP2 | VAMP2 Entrez, Source | vesicle-associated membrane protein 2 (synaptobrevin 2) | 6334 | 0.395 | 0.3739 | No |
| 61 | SNX2 | SNX2 Entrez, Source | sorting nexin 2 | 6344 | 0.394 | 0.3756 | No |
| 62 | PUM1 | PUM1 Entrez, Source | pumilio homolog 1 (Drosophila) | 7062 | 0.209 | 0.3249 | No |
| 63 | DNM2 | DNM2 Entrez, Source | dynamin 2 | 7444 | 0.109 | 0.2979 | No |
| 64 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 7814 | 0.011 | 0.2712 | No |
| 65 | GJA4 | GJA4 Entrez, Source | gap junction protein, alpha 4, 37kDa (connexin 37) | 7884 | -0.011 | 0.2663 | No |
| 66 | GJB1 | GJB1 Entrez, Source | gap junction protein, beta 1, 32kDa (connexin 32, Charcot-Marie-Tooth neuropathy, X-linked) | 7948 | -0.028 | 0.2619 | No |
| 67 | TJP1 | TJP1 Entrez, Source | tight junction protein 1 (zona occludens 1) | 8044 | -0.055 | 0.2553 | No |
| 68 | AP3S1 | AP3S1 Entrez, Source | adaptor-related protein complex 3, sigma 1 subunit | 8103 | -0.073 | 0.2515 | No |
| 69 | TSG101 | TSG101 Entrez, Source | tumor susceptibility gene 101 | 8317 | -0.126 | 0.2369 | No |
| 70 | SNF8 | SNF8 Entrez, Source | SNF8, ESCRT-II complex subunit, homolog (S. cerevisiae) | 8427 | -0.160 | 0.2299 | No |
| 71 | OCRL | OCRL Entrez, Source | oculocerebrorenal syndrome of Lowe | 8649 | -0.227 | 0.2152 | No |
| 72 | AP2M1 | AP2M1 Entrez, Source | adaptor-related protein complex 2, mu 1 subunit | 8718 | -0.250 | 0.2118 | No |
| 73 | DTNBP1 | DTNBP1 Entrez, Source | dystrobrevin binding protein 1 | 8766 | -0.269 | 0.2100 | No |
| 74 | AP3B1 | AP3B1 Entrez, Source | adaptor-related protein complex 3, beta 1 subunit | 8847 | -0.293 | 0.2060 | No |
| 75 | SNAP23 | SNAP23 Entrez, Source | synaptosomal-associated protein, 23kDa | 9091 | -0.378 | 0.1906 | No |
| 76 | VPS37D | VPS37D Entrez, Source | vacuolar protein sorting 37 homolog D (S. cerevisiae) | 9143 | -0.396 | 0.1893 | No |
| 77 | CHMP4A | CHMP4A Entrez, Source | chromatin modifying protein 4A | 9245 | -0.426 | 0.1845 | No |
| 78 | SRC | SRC Entrez, Source | v-src sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog (avian) | 9276 | -0.435 | 0.1850 | No |
| 79 | CLTCL1 | CLTCL1 Entrez, Source | clathrin, heavy chain-like 1 | 9451 | -0.502 | 0.1754 | No |
| 80 | VPS37C | VPS37C Entrez, Source | vacuolar protein sorting 37 homolog C (S. cerevisiae) | 9604 | -0.553 | 0.1677 | No |
| 81 | HIP1R | HIP1R Entrez, Source | huntingtin interacting protein 1 related | 9982 | -0.714 | 0.1446 | No |
| 82 | CNO | CNO Entrez, Source | cappuccino homolog (mouse) | 10056 | -0.758 | 0.1439 | No |
| 83 | ARRB1 | ARRB1 Entrez, Source | arrestin, beta 1 | 10427 | -0.935 | 0.1227 | No |
| 84 | AP1M2 | AP1M2 Entrez, Source | adaptor-related protein complex 1, mu 2 subunit | 10480 | -0.961 | 0.1247 | No |
| 85 | DAB2 | DAB2 Entrez, Source | disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila) | 10564 | -1.003 | 0.1247 | No |
| 86 | SNX5 | SNX5 Entrez, Source | sorting nexin 5 | 10642 | -1.037 | 0.1254 | No |
| 87 | AP1S1 | AP1S1 Entrez, Source | adaptor-related protein complex 1, sigma 1 subunit | 10696 | -1.066 | 0.1279 | No |
| 88 | VPS28 | VPS28 Entrez, Source | vacuolar protein sorting 28 homolog (S. cerevisiae) | 11496 | -1.502 | 0.0790 | No |
| 89 | SORT1 | SORT1 Entrez, Source | sortilin 1 | 11654 | -1.607 | 0.0773 | No |
| 90 | VAMP8 | VAMP8 Entrez, Source | vesicle-associated membrane protein 8 (endobrevin) | 11826 | -1.715 | 0.0752 | No |
| 91 | DNASE2 | DNASE2 Entrez, Source | deoxyribonuclease II, lysosomal | 11857 | -1.745 | 0.0835 | No |
| 92 | MYO6 | MYO6 Entrez, Source | myosin VI | 11871 | -1.757 | 0.0932 | No |
| 93 | BLOC1S1 | BLOC1S1 Entrez, Source | biogenesis of lysosome-related organelles complex-1, subunit 1 | 11972 | -1.821 | 0.0969 | No |
| 94 | CHMP2A | CHMP2A Entrez, Source | chromatin modifying protein 2A | 12232 | -2.024 | 0.0903 | No |
| 95 | TGOLN2 | TGOLN2 Entrez, Source | trans-golgi network protein 2 | 12541 | -2.301 | 0.0818 | No |
| 96 | TPD52L1 | TPD52L1 Entrez, Source | tumor protein D52-like 1 | 12805 | -2.577 | 0.0782 | No |