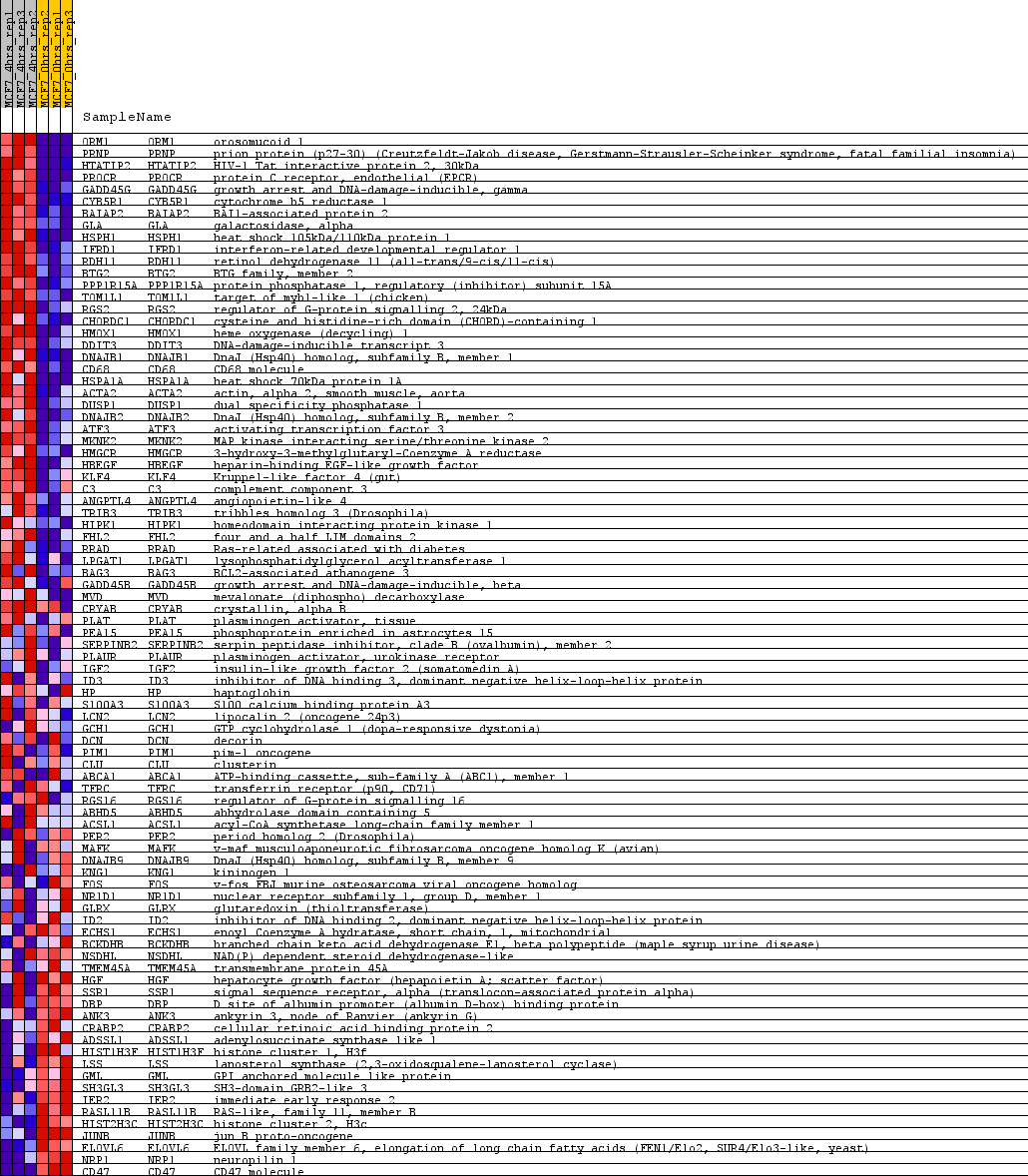
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #4hr_versus_0hr.class.cls #4hr_versus_0hr_repos |
| Phenotype | class.cls#4hr_versus_0hr_repos |
| Upregulated in class | 4hr |
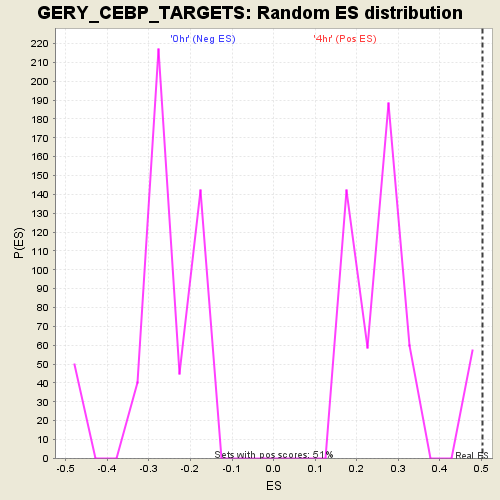
| GeneSet | GERY_CEBP_TARGETS |
| Enrichment Score (ES) | 0.5033278 |
| Normalized Enrichment Score (NES) | 1.7978662 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.5558944 |
| FWER p-Value | 0.51 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ORM1 | ORM1 Entrez, Source | orosomucoid 1 | 6 | 4.554 | 0.0400 | Yes |
| 2 | PRNP | PRNP Entrez, Source | prion protein (p27-30) (Creutzfeldt-Jakob disease, Gerstmann-Strausler-Scheinker syndrome, fatal familial insomnia) | 70 | 3.808 | 0.0692 | Yes |
| 3 | HTATIP2 | HTATIP2 Entrez, Source | HIV-1 Tat interactive protein 2, 30kDa | 121 | 3.532 | 0.0969 | Yes |
| 4 | PROCR | PROCR Entrez, Source | protein C receptor, endothelial (EPCR) | 145 | 3.414 | 0.1255 | Yes |
| 5 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 229 | 3.167 | 0.1476 | Yes |
| 6 | CYB5R1 | CYB5R1 Entrez, Source | cytochrome b5 reductase 1 | 245 | 3.129 | 0.1742 | Yes |
| 7 | BAIAP2 | BAIAP2 Entrez, Source | BAI1-associated protein 2 | 405 | 2.764 | 0.1872 | Yes |
| 8 | GLA | GLA Entrez, Source | galactosidase, alpha | 419 | 2.737 | 0.2106 | Yes |
| 9 | HSPH1 | HSPH1 Entrez, Source | heat shock 105kDa/110kDa protein 1 | 446 | 2.691 | 0.2325 | Yes |
| 10 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 506 | 2.591 | 0.2513 | Yes |
| 11 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 522 | 2.562 | 0.2729 | Yes |
| 12 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 545 | 2.520 | 0.2936 | Yes |
| 13 | PPP1R15A | PPP1R15A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 15A | 673 | 2.344 | 0.3052 | Yes |
| 14 | TOM1L1 | TOM1L1 Entrez, Source | target of myb1-like 1 (chicken) | 704 | 2.290 | 0.3234 | Yes |
| 15 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 787 | 2.197 | 0.3369 | Yes |
| 16 | CHORDC1 | CHORDC1 Entrez, Source | cysteine and histidine-rich domain (CHORD)-containing 1 | 824 | 2.146 | 0.3533 | Yes |
| 17 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 825 | 2.144 | 0.3724 | Yes |
| 18 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 911 | 2.059 | 0.3845 | Yes |
| 19 | DNAJB1 | DNAJB1 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 1 | 962 | 2.008 | 0.3987 | Yes |
| 20 | CD68 | CD68 Entrez, Source | CD68 molecule | 984 | 1.988 | 0.4148 | Yes |
| 21 | HSPA1A | HSPA1A Entrez, Source | heat shock 70kDa protein 1A | 1071 | 1.903 | 0.4254 | Yes |
| 22 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 1278 | 1.713 | 0.4257 | Yes |
| 23 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1329 | 1.670 | 0.4369 | Yes |
| 24 | DNAJB2 | DNAJB2 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 2 | 1388 | 1.620 | 0.4470 | Yes |
| 25 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 1399 | 1.610 | 0.4606 | Yes |
| 26 | MKNK2 | MKNK2 Entrez, Source | MAP kinase interacting serine/threonine kinase 2 | 1410 | 1.601 | 0.4741 | Yes |
| 27 | HMGCR | HMGCR Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A reductase | 1522 | 1.501 | 0.4793 | Yes |
| 28 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 1606 | 1.449 | 0.4862 | Yes |
| 29 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 1808 | 1.326 | 0.4834 | Yes |
| 30 | C3 | C3 Entrez, Source | complement component 3 | 1970 | 1.236 | 0.4827 | Yes |
| 31 | ANGPTL4 | ANGPTL4 Entrez, Source | angiopoietin-like 4 | 2093 | 1.171 | 0.4842 | Yes |
| 32 | TRIB3 | TRIB3 Entrez, Source | tribbles homolog 3 (Drosophila) | 2292 | 1.078 | 0.4794 | Yes |
| 33 | HIPK1 | HIPK1 Entrez, Source | homeodomain interacting protein kinase 1 | 2494 | 0.988 | 0.4736 | Yes |
| 34 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 2505 | 0.985 | 0.4816 | Yes |
| 35 | RRAD | RRAD Entrez, Source | Ras-related associated with diabetes | 2512 | 0.978 | 0.4899 | Yes |
| 36 | LPGAT1 | LPGAT1 Entrez, Source | lysophosphatidylglycerol acyltransferase 1 | 2533 | 0.969 | 0.4970 | Yes |
| 37 | BAG3 | BAG3 Entrez, Source | BCL2-associated athanogene 3 | 2564 | 0.957 | 0.5033 | Yes |
| 38 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 3191 | 0.771 | 0.4648 | No |
| 39 | MVD | MVD Entrez, Source | mevalonate (diphospho) decarboxylase | 3274 | 0.747 | 0.4655 | No |
| 40 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 3750 | 0.629 | 0.4366 | No |
| 41 | PLAT | PLAT Entrez, Source | plasminogen activator, tissue | 3937 | 0.585 | 0.4283 | No |
| 42 | PEA15 | PEA15 Entrez, Source | phosphoprotein enriched in astrocytes 15 | 4298 | 0.505 | 0.4067 | No |
| 43 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 4347 | 0.497 | 0.4076 | No |
| 44 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 4380 | 0.489 | 0.4097 | No |
| 45 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 4458 | 0.472 | 0.4083 | No |
| 46 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 4674 | 0.426 | 0.3965 | No |
| 47 | HP | HP Entrez, Source | haptoglobin | 4925 | 0.378 | 0.3817 | No |
| 48 | S100A3 | S100A3 Entrez, Source | S100 calcium binding protein A3 | 4979 | 0.369 | 0.3811 | No |
| 49 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 5056 | 0.356 | 0.3788 | No |
| 50 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 5451 | 0.284 | 0.3527 | No |
| 51 | DCN | DCN Entrez, Source | decorin | 5677 | 0.247 | 0.3386 | No |
| 52 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 5704 | 0.242 | 0.3389 | No |
| 53 | CLU | CLU Entrez, Source | clusterin | 5735 | 0.236 | 0.3388 | No |
| 54 | ABCA1 | ABCA1 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 1 | 6256 | 0.157 | 0.3025 | No |
| 55 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 6315 | 0.147 | 0.2996 | No |
| 56 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 6648 | 0.093 | 0.2764 | No |
| 57 | ABHD5 | ABHD5 Entrez, Source | abhydrolase domain containing 5 | 6752 | 0.078 | 0.2696 | No |
| 58 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 6817 | 0.068 | 0.2655 | No |
| 59 | PER2 | PER2 Entrez, Source | period homolog 2 (Drosophila) | 7181 | 0.013 | 0.2393 | No |
| 60 | MAFK | MAFK Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog K (avian) | 7492 | -0.034 | 0.2172 | No |
| 61 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 7781 | -0.081 | 0.1970 | No |
| 62 | KNG1 | KNG1 Entrez, Source | kininogen 1 | 7978 | -0.113 | 0.1838 | No |
| 63 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 8641 | -0.218 | 0.1378 | No |
| 64 | NR1D1 | NR1D1 Entrez, Source | nuclear receptor subfamily 1, group D, member 1 | 9225 | -0.315 | 0.0983 | No |
| 65 | GLRX | GLRX Entrez, Source | glutaredoxin (thioltransferase) | 9406 | -0.345 | 0.0883 | No |
| 66 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 9572 | -0.373 | 0.0797 | No |
| 67 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 9581 | -0.374 | 0.0824 | No |
| 68 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 9986 | -0.444 | 0.0571 | No |
| 69 | NSDHL | NSDHL Entrez, Source | NAD(P) dependent steroid dehydrogenase-like | 10157 | -0.474 | 0.0490 | No |
| 70 | TMEM45A | TMEM45A Entrez, Source | transmembrane protein 45A | 10275 | -0.494 | 0.0449 | No |
| 71 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 11125 | -0.678 | -0.0107 | No |
| 72 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 11329 | -0.733 | -0.0189 | No |
| 73 | DBP | DBP Entrez, Source | D site of albumin promoter (albumin D-box) binding protein | 11384 | -0.749 | -0.0162 | No |
| 74 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 11894 | -0.896 | -0.0451 | No |
| 75 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 12053 | -0.951 | -0.0481 | No |
| 76 | ADSSL1 | ADSSL1 Entrez, Source | adenylosuccinate synthase like 1 | 12214 | -1.012 | -0.0507 | No |
| 77 | HIST1H3F | HIST1H3F Entrez, Source | histone cluster 1, H3f | 12289 | -1.045 | -0.0468 | No |
| 78 | LSS | LSS Entrez, Source | lanosterol synthase (2,3-oxidosqualene-lanosterol cyclase) | 12444 | -1.109 | -0.0481 | No |
| 79 | GML | GML Entrez, Source | GPI anchored molecule like protein | 12459 | -1.117 | -0.0392 | No |
| 80 | SH3GL3 | SH3GL3 Entrez, Source | SH3-domain GRB2-like 3 | 12931 | -1.367 | -0.0613 | No |
| 81 | IER2 | IER2 Entrez, Source | immediate early response 2 | 12959 | -1.390 | -0.0509 | No |
| 82 | RASL11B | RASL11B Entrez, Source | RAS-like, family 11, member B | 13471 | -1.958 | -0.0706 | No |
| 83 | HIST2H3C | HIST2H3C Entrez, Source | histone cluster 2, H3c | 13476 | -1.964 | -0.0534 | No |
| 84 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 13487 | -1.983 | -0.0366 | No |
| 85 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 13488 | -1.984 | -0.0190 | No |
| 86 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 13646 | -2.328 | -0.0097 | No |
| 87 | CD47 | CD47 Entrez, Source | CD47 molecule | 13801 | -3.022 | 0.0059 | No |