Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | data_collapsed_to_symbols.class.cls #4hr_versus_0hr.class.cls #4hr_versus_0hr_repos |
| Phenotype | class.cls#4hr_versus_0hr_repos |
| Upregulated in class | 4hr |
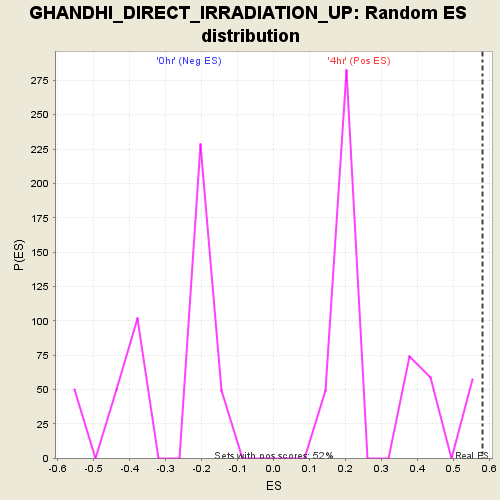
| GeneSet | GHANDHI_DIRECT_IRRADIATION_UP |
| Enrichment Score (ES) | 0.5809332 |
| Normalized Enrichment Score (NES) | 2.0278437 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.46585077 |
| FWER p-Value | 0.19 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FAS | FAS Entrez, Source | Fas (TNF receptor superfamily, member 6) | 8 | 4.549 | 0.0584 | Yes |
| 2 | BLOC1S2 | BLOC1S2 Entrez, Source | biogenesis of lysosome-related organelles complex-1, subunit 2 | 20 | 4.275 | 0.1130 | Yes |
| 3 | SESN1 | SESN1 Entrez, Source | sestrin 1 | 21 | 4.244 | 0.1680 | Yes |
| 4 | MDM2 | MDM2 Entrez, Source | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein (mouse) | 87 | 3.712 | 0.2114 | Yes |
| 5 | PLK3 | PLK3 Entrez, Source | polo-like kinase 3 (Drosophila) | 166 | 3.348 | 0.2491 | Yes |
| 6 | TNFRSF10B | TNFRSF10B Entrez, Source | tumor necrosis factor receptor superfamily, member 10b | 177 | 3.315 | 0.2913 | Yes |
| 7 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 251 | 3.115 | 0.3264 | Yes |
| 8 | TNFRSF10C | TNFRSF10C Entrez, Source | tumor necrosis factor receptor superfamily, member 10c, decoy without an intracellular domain | 311 | 2.976 | 0.3607 | Yes |
| 9 | POLH | POLH Entrez, Source | polymerase (DNA directed), eta | 334 | 2.925 | 0.3970 | Yes |
| 10 | ASCC3 | ASCC3 Entrez, Source | activating signal cointegrator 1 complex subunit 3 | 432 | 2.718 | 0.4252 | Yes |
| 11 | SLC7A11 | SLC7A11 Entrez, Source | solute carrier family 7, (cationic amino acid transporter, y+ system) member 11 | 487 | 2.616 | 0.4552 | Yes |
| 12 | BBC3 | BBC3 Entrez, Source | BCL2 binding component 3 | 641 | 2.387 | 0.4751 | Yes |
| 13 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 867 | 2.106 | 0.4861 | Yes |
| 14 | GADD45A | GADD45A Entrez, Source | growth arrest and DNA-damage-inducible, alpha | 960 | 2.009 | 0.5055 | Yes |
| 15 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 1121 | 1.856 | 0.5179 | Yes |
| 16 | LRP8 | LRP8 Entrez, Source | low density lipoprotein receptor-related protein 8, apolipoprotein e receptor | 1239 | 1.751 | 0.5321 | Yes |
| 17 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 1248 | 1.743 | 0.5542 | Yes |
| 18 | HSPA4L | HSPA4L Entrez, Source | heat shock 70kDa protein 4-like | 1446 | 1.567 | 0.5602 | Yes |
| 19 | CXCL3 | CXCL3 Entrez, Source | chemokine (C-X-C motif) ligand 3 | 1562 | 1.475 | 0.5710 | Yes |
| 20 | BTG3 | BTG3 Entrez, Source | BTG family, member 3 | 1676 | 1.398 | 0.5809 | Yes |
| 21 | DDIT4L | DDIT4L Entrez, Source | DNA-damage-inducible transcript 4-like | 2238 | 1.106 | 0.5547 | No |
| 22 | THSD1 | THSD1 Entrez, Source | thrombospondin, type I, domain containing 1 | 2745 | 0.899 | 0.5297 | No |
| 23 | PTPRE | PTPRE Entrez, Source | protein tyrosine phosphatase, receptor type, E | 3070 | 0.805 | 0.5167 | No |
| 24 | EYA3 | EYA3 Entrez, Source | eyes absent homolog 3 (Drosophila) | 3673 | 0.650 | 0.4816 | No |
| 25 | RRM2B | RRM2B Entrez, Source | ribonucleotide reductase M2 B (TP53 inducible) | 3932 | 0.586 | 0.4705 | No |
| 26 | LOC284454 | LOC284454 Entrez, Source | - | 4159 | 0.536 | 0.4611 | No |
| 27 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 4336 | 0.499 | 0.4548 | No |
| 28 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 4347 | 0.497 | 0.4605 | No |
| 29 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 4440 | 0.476 | 0.4600 | No |
| 30 | RELB | RELB Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog B, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3 (avian) | 4498 | 0.464 | 0.4619 | No |
| 31 | BMP2 | BMP2 Entrez, Source | bone morphogenetic protein 2 | 4569 | 0.449 | 0.4627 | No |
| 32 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 4771 | 0.406 | 0.4534 | No |
| 33 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 5049 | 0.357 | 0.4380 | No |
| 34 | IL11 | IL11 Entrez, Source | interleukin 11 | 5064 | 0.355 | 0.4416 | No |
| 35 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 5451 | 0.284 | 0.4173 | No |
| 36 | GDNF | GDNF Entrez, Source | glial cell derived neurotrophic factor | 5871 | 0.213 | 0.3898 | No |
| 37 | LAMB3 | LAMB3 Entrez, Source | laminin, beta 3 | 5900 | 0.209 | 0.3904 | No |
| 38 | E2F7 | E2F7 Entrez, Source | E2F transcription factor 7 | 5932 | 0.206 | 0.3909 | No |
| 39 | POU5F1 | POU5F1 Entrez, Source | POU domain, class 5, transcription factor 1 | 6158 | 0.171 | 0.3768 | No |
| 40 | MT1A | MT1A Entrez, Source | metallothionein 1A (functional) | 6502 | 0.119 | 0.3535 | No |
| 41 | ZC3H12C | ZC3H12C Entrez, Source | zinc finger CCCH-type containing 12C | 7106 | 0.023 | 0.3102 | No |
| 42 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 7146 | 0.018 | 0.3076 | No |
| 43 | CLDN1 | CLDN1 Entrez, Source | claudin 1 | 7317 | -0.007 | 0.2954 | No |
| 44 | CCK | CCK Entrez, Source | cholecystokinin | 7343 | -0.011 | 0.2937 | No |
| 45 | NFKBIZ | NFKBIZ Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta | 7745 | -0.075 | 0.2657 | No |
| 46 | TNFAIP3 | TNFAIP3 Entrez, Source | tumor necrosis factor, alpha-induced protein 3 | 7870 | -0.096 | 0.2579 | No |
| 47 | FAM20A | FAM20A Entrez, Source | family with sequence similarity 20, member A | 8364 | -0.175 | 0.2245 | No |
| 48 | MT1L | MT1L Entrez, Source | metallothionein 1L (pseudogene) | 9033 | -0.285 | 0.1799 | No |
| 49 | MT1E | MT1E Entrez, Source | metallothionein 1E (functional) | 9274 | -0.324 | 0.1667 | No |
| 50 | MYBL1 | MYBL1 Entrez, Source | v-myb myeloblastosis viral oncogene homolog (avian)-like 1 | 9336 | -0.334 | 0.1666 | No |
| 51 | SLC30A1 | SLC30A1 Entrez, Source | solute carrier family 30 (zinc transporter), member 1 | 9475 | -0.357 | 0.1613 | No |
| 52 | MT1B | MT1B Entrez, Source | metallothionein 1B (functional) | 9737 | -0.398 | 0.1476 | No |
| 53 | MT1X | MT1X Entrez, Source | metallothionein 1X | 9837 | -0.420 | 0.1458 | No |
| 54 | GPR68 | GPR68 Entrez, Source | G protein-coupled receptor 68 | 9924 | -0.434 | 0.1452 | No |
| 55 | MT1G | MT1G Entrez, Source | metallothionein 1G | 10044 | -0.455 | 0.1425 | No |
| 56 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 10138 | -0.471 | 0.1419 | No |
| 57 | MT2A | MT2A Entrez, Source | metallothionein 2A | 10638 | -0.561 | 0.1131 | No |
| 58 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 11391 | -0.751 | 0.0684 | No |
| 59 | CYP26B1 | CYP26B1 Entrez, Source | cytochrome P450, family 26, subfamily B, polypeptide 1 | 11853 | -0.884 | 0.0465 | No |
| 60 | SLC16A6 | SLC16A6 Entrez, Source | solute carrier family 16, member 6 (monocarboxylic acid transporter 7) | 12209 | -1.009 | 0.0339 | No |
| 61 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 12297 | -1.048 | 0.0411 | No |
| 62 | DKK1 | DKK1 Entrez, Source | dickkopf homolog 1 (Xenopus laevis) | 12779 | -1.270 | 0.0228 | No |
| 63 | FRMD4A | FRMD4A Entrez, Source | FERM domain containing 4A | 13564 | -2.140 | -0.0062 | No |
| 64 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 13613 | -2.255 | 0.0195 | No |