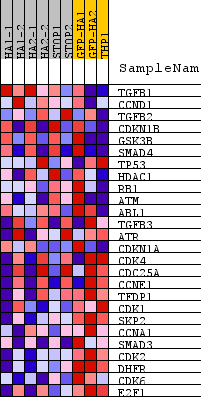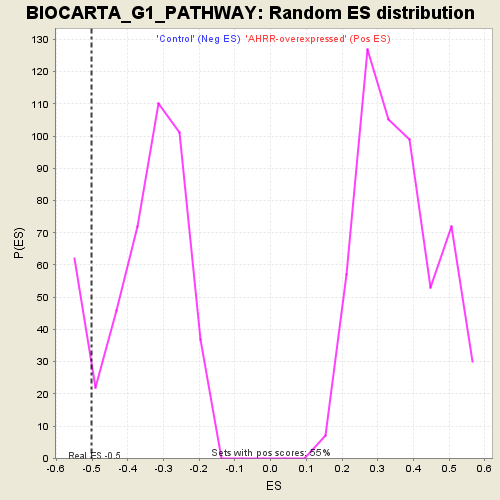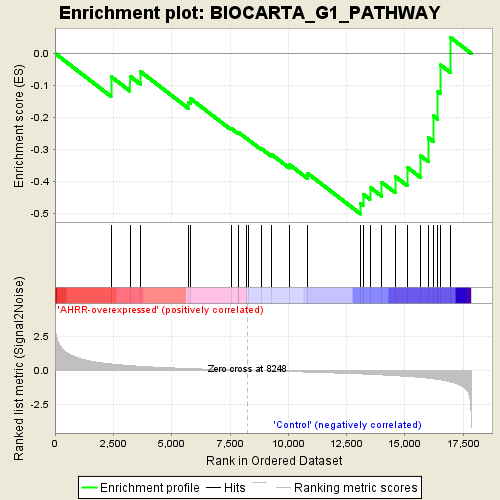
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA.class.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | BIOCARTA_G1_PATHWAY |
| Enrichment Score (ES) | -0.50244874 |
| Normalized Enrichment Score (NES) | -1.4082075 |
| Nominal p-value | 0.13777778 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.991 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TGFB1 | na | 2399 | 0.481 | -0.0722 | No | ||
| 2 | CCND1 | na | 3212 | 0.359 | -0.0712 | No | ||
| 3 | TGFB2 | na | 3660 | 0.306 | -0.0566 | No | ||
| 4 | CDKN1B | na | 5709 | 0.140 | -0.1534 | No | ||
| 5 | GSK3B | na | 5796 | 0.135 | -0.1406 | No | ||
| 6 | SMAD4 | na | 7554 | 0.037 | -0.2345 | No | ||
| 7 | TP53 | na | 7860 | 0.019 | -0.2492 | No | ||
| 8 | HDAC1 | na | 7874 | 0.019 | -0.2475 | No | ||
| 9 | RB1 | na | 8213 | 0.002 | -0.2662 | No | ||
| 10 | ATM | na | 8283 | -0.001 | -0.2699 | No | ||
| 11 | ABL1 | na | 8829 | -0.027 | -0.2969 | No | ||
| 12 | TGFB3 | na | 9261 | -0.048 | -0.3149 | No | ||
| 13 | ATR | na | 10040 | -0.086 | -0.3474 | No | ||
| 14 | CDKN1A | na | 10829 | -0.127 | -0.3751 | No | ||
| 15 | CDK4 | na | 13098 | -0.259 | -0.4688 | Yes | ||
| 16 | CDC25A | na | 13205 | -0.268 | -0.4399 | Yes | ||
| 17 | CCNE1 | na | 13504 | -0.293 | -0.4186 | Yes | ||
| 18 | TFDP1 | na | 14006 | -0.335 | -0.4033 | Yes | ||
| 19 | CDK1 | na | 14598 | -0.390 | -0.3859 | Yes | ||
| 20 | SKP2 | na | 15108 | -0.445 | -0.3567 | Yes | ||
| 21 | CCNA1 | na | 15669 | -0.523 | -0.3203 | Yes | ||
| 22 | SMAD3 | na | 16014 | -0.581 | -0.2641 | Yes | ||
| 23 | CDK2 | na | 16222 | -0.618 | -0.1955 | Yes | ||
| 24 | DHFR | na | 16410 | -0.663 | -0.1200 | Yes | ||
| 25 | CDK6 | na | 16528 | -0.697 | -0.0361 | Yes | ||
| 26 | E2F1 | na | 16952 | -0.845 | 0.0499 | Yes |