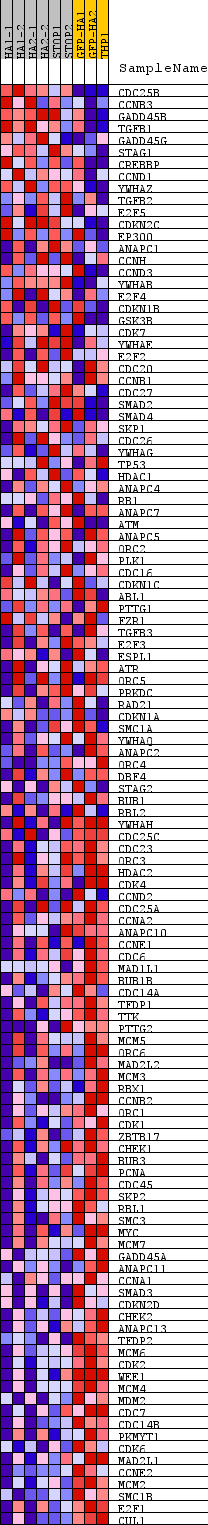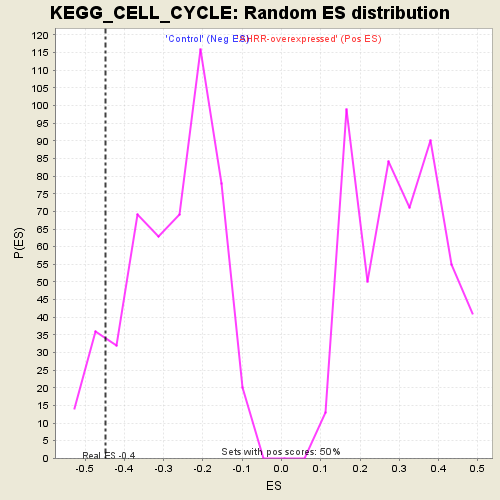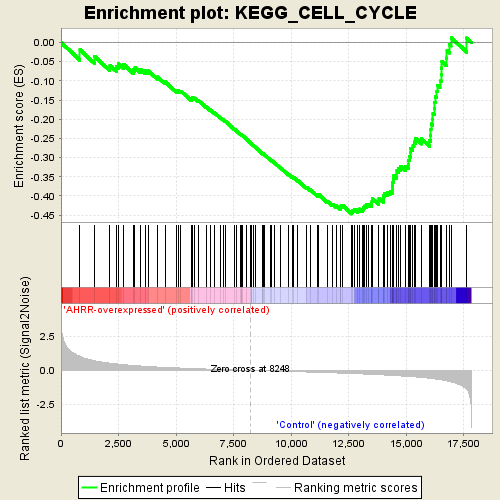
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA.class.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | KEGG_CELL_CYCLE |
| Enrichment Score (ES) | -0.44695765 |
| Normalized Enrichment Score (NES) | -1.6145761 |
| Nominal p-value | 0.078470826 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.679 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDC25B | na | 821 | 1.036 | -0.0190 | No | ||
| 2 | CCNB3 | na | 1457 | 0.713 | -0.0361 | No | ||
| 3 | GADD45B | na | 2125 | 0.535 | -0.0596 | No | ||
| 4 | TGFB1 | na | 2399 | 0.481 | -0.0624 | No | ||
| 5 | GADD45G | na | 2497 | 0.462 | -0.0557 | No | ||
| 6 | STAG1 | na | 2710 | 0.426 | -0.0564 | No | ||
| 7 | CREBBP | na | 3142 | 0.368 | -0.0710 | No | ||
| 8 | CCND1 | na | 3212 | 0.359 | -0.0655 | No | ||
| 9 | YWHAZ | na | 3450 | 0.329 | -0.0702 | No | ||
| 10 | TGFB2 | na | 3660 | 0.306 | -0.0739 | No | ||
| 11 | E2F5 | na | 3792 | 0.292 | -0.0736 | No | ||
| 12 | CDKN2C | na | 4196 | 0.250 | -0.0898 | No | ||
| 13 | EP300 | na | 4523 | 0.225 | -0.1023 | No | ||
| 14 | ANAPC1 | na | 5038 | 0.185 | -0.1264 | No | ||
| 15 | CCNH | na | 5111 | 0.181 | -0.1257 | No | ||
| 16 | CCND3 | na | 5209 | 0.174 | -0.1266 | No | ||
| 17 | YWHAB | na | 5668 | 0.144 | -0.1486 | No | ||
| 18 | E2F4 | na | 5689 | 0.142 | -0.1460 | No | ||
| 19 | CDKN1B | na | 5709 | 0.140 | -0.1434 | No | ||
| 20 | GSK3B | na | 5796 | 0.135 | -0.1447 | No | ||
| 21 | CDK7 | na | 5952 | 0.125 | -0.1501 | No | ||
| 22 | YWHAE | na | 6312 | 0.104 | -0.1676 | No | ||
| 23 | E2F2 | na | 6489 | 0.093 | -0.1751 | No | ||
| 24 | CDC20 | na | 6648 | 0.084 | -0.1818 | No | ||
| 25 | CCNB1 | na | 6949 | 0.068 | -0.1970 | No | ||
| 26 | CDC27 | na | 7056 | 0.062 | -0.2013 | No | ||
| 27 | SMAD2 | na | 7159 | 0.056 | -0.2056 | No | ||
| 28 | SMAD4 | na | 7554 | 0.037 | -0.2269 | No | ||
| 29 | SKP1 | na | 7644 | 0.032 | -0.2311 | No | ||
| 30 | CDC26 | na | 7802 | 0.022 | -0.2393 | No | ||
| 31 | YWHAG | na | 7848 | 0.020 | -0.2413 | No | ||
| 32 | TP53 | na | 7860 | 0.019 | -0.2415 | No | ||
| 33 | HDAC1 | na | 7874 | 0.019 | -0.2417 | No | ||
| 34 | ANAPC4 | na | 8048 | 0.010 | -0.2512 | No | ||
| 35 | RB1 | na | 8213 | 0.002 | -0.2604 | No | ||
| 36 | ANAPC7 | na | 8280 | -0.001 | -0.2641 | No | ||
| 37 | ATM | na | 8283 | -0.001 | -0.2642 | No | ||
| 38 | ANAPC5 | na | 8366 | -0.005 | -0.2687 | No | ||
| 39 | ORC2 | na | 8456 | -0.010 | -0.2734 | No | ||
| 40 | PLK1 | na | 8766 | -0.025 | -0.2902 | No | ||
| 41 | CDC16 | na | 8769 | -0.025 | -0.2897 | No | ||
| 42 | CDKN1C | na | 8781 | -0.025 | -0.2896 | No | ||
| 43 | ABL1 | na | 8829 | -0.027 | -0.2915 | No | ||
| 44 | PTTG1 | na | 9083 | -0.040 | -0.3048 | No | ||
| 45 | FZR1 | na | 9161 | -0.043 | -0.3080 | No | ||
| 46 | TGFB3 | na | 9261 | -0.048 | -0.3123 | No | ||
| 47 | E2F3 | na | 9560 | -0.062 | -0.3275 | No | ||
| 48 | ESPL1 | na | 9879 | -0.078 | -0.3434 | No | ||
| 49 | ATR | na | 10040 | -0.086 | -0.3501 | No | ||
| 50 | ORC5 | na | 10124 | -0.091 | -0.3524 | No | ||
| 51 | PRKDC | na | 10283 | -0.098 | -0.3587 | No | ||
| 52 | RAD21 | na | 10672 | -0.118 | -0.3775 | No | ||
| 53 | CDKN1A | na | 10829 | -0.127 | -0.3830 | No | ||
| 54 | SMC1A | na | 11142 | -0.143 | -0.3968 | No | ||
| 55 | YWHAQ | na | 11177 | -0.145 | -0.3949 | No | ||
| 56 | ANAPC2 | na | 11579 | -0.169 | -0.4131 | No | ||
| 57 | ORC4 | na | 11802 | -0.181 | -0.4209 | No | ||
| 58 | DBF4 | na | 11961 | -0.189 | -0.4248 | No | ||
| 59 | STAG2 | na | 12165 | -0.201 | -0.4309 | No | ||
| 60 | BUB1 | na | 12168 | -0.202 | -0.4257 | No | ||
| 61 | RBL2 | na | 12239 | -0.206 | -0.4243 | No | ||
| 62 | YWHAH | na | 12642 | -0.228 | -0.4409 | Yes | ||
| 63 | CDC25C | na | 12684 | -0.232 | -0.4372 | Yes | ||
| 64 | CDC23 | na | 12757 | -0.237 | -0.4350 | Yes | ||
| 65 | ORC3 | na | 12898 | -0.246 | -0.4364 | Yes | ||
| 66 | HDAC2 | na | 12970 | -0.251 | -0.4338 | Yes | ||
| 67 | CDK4 | na | 13098 | -0.259 | -0.4341 | Yes | ||
| 68 | CCND2 | na | 13142 | -0.263 | -0.4296 | Yes | ||
| 69 | CDC25A | na | 13205 | -0.268 | -0.4261 | Yes | ||
| 70 | CCNA2 | na | 13271 | -0.273 | -0.4225 | Yes | ||
| 71 | ANAPC10 | na | 13375 | -0.282 | -0.4209 | Yes | ||
| 72 | CCNE1 | na | 13504 | -0.293 | -0.4204 | Yes | ||
| 73 | CDC6 | na | 13515 | -0.293 | -0.4133 | Yes | ||
| 74 | MAD1L1 | na | 13537 | -0.294 | -0.4067 | Yes | ||
| 75 | BUB1B | na | 13804 | -0.317 | -0.4134 | Yes | ||
| 76 | CDC14A | na | 13819 | -0.318 | -0.4058 | Yes | ||
| 77 | TFDP1 | na | 14006 | -0.335 | -0.4074 | Yes | ||
| 78 | TTK | na | 14028 | -0.336 | -0.3998 | Yes | ||
| 79 | PTTG2 | na | 14074 | -0.340 | -0.3934 | Yes | ||
| 80 | MCM5 | na | 14184 | -0.351 | -0.3903 | Yes | ||
| 81 | ORC6 | na | 14302 | -0.361 | -0.3874 | Yes | ||
| 82 | MAD2L2 | na | 14398 | -0.370 | -0.3830 | Yes | ||
| 83 | MCM3 | na | 14407 | -0.371 | -0.3737 | Yes | ||
| 84 | RBX1 | na | 14419 | -0.372 | -0.3645 | Yes | ||
| 85 | CCNB2 | na | 14435 | -0.374 | -0.3555 | Yes | ||
| 86 | ORC1 | na | 14468 | -0.377 | -0.3473 | Yes | ||
| 87 | CDK1 | na | 14598 | -0.390 | -0.3444 | Yes | ||
| 88 | ZBTB17 | na | 14601 | -0.390 | -0.3342 | Yes | ||
| 89 | CHEK1 | na | 14674 | -0.399 | -0.3278 | Yes | ||
| 90 | BUB3 | na | 14763 | -0.408 | -0.3220 | Yes | ||
| 91 | PCNA | na | 14965 | -0.429 | -0.3220 | Yes | ||
| 92 | CDC45 | na | 15097 | -0.444 | -0.3177 | Yes | ||
| 93 | SKP2 | na | 15108 | -0.445 | -0.3066 | Yes | ||
| 94 | RBL1 | na | 15135 | -0.448 | -0.2962 | Yes | ||
| 95 | SMC3 | na | 15178 | -0.452 | -0.2867 | Yes | ||
| 96 | MYC | na | 15185 | -0.453 | -0.2751 | Yes | ||
| 97 | MCM7 | na | 15298 | -0.469 | -0.2691 | Yes | ||
| 98 | GADD45A | na | 15344 | -0.474 | -0.2591 | Yes | ||
| 99 | ANAPC11 | na | 15408 | -0.483 | -0.2500 | Yes | ||
| 100 | CCNA1 | na | 15669 | -0.523 | -0.2509 | Yes | ||
| 101 | SMAD3 | na | 16014 | -0.581 | -0.2550 | Yes | ||
| 102 | CDKN2D | na | 16057 | -0.588 | -0.2418 | Yes | ||
| 103 | CHEK2 | na | 16077 | -0.594 | -0.2273 | Yes | ||
| 104 | ANAPC13 | na | 16085 | -0.595 | -0.2120 | Yes | ||
| 105 | TFDP2 | na | 16160 | -0.604 | -0.2002 | Yes | ||
| 106 | MCM6 | na | 16167 | -0.607 | -0.1846 | Yes | ||
| 107 | CDK2 | na | 16222 | -0.618 | -0.1714 | Yes | ||
| 108 | WEE1 | na | 16242 | -0.624 | -0.1560 | Yes | ||
| 109 | MCM4 | na | 16263 | -0.629 | -0.1405 | Yes | ||
| 110 | MDM2 | na | 16337 | -0.647 | -0.1276 | Yes | ||
| 111 | CDC7 | na | 16354 | -0.651 | -0.1114 | Yes | ||
| 112 | CDC14B | na | 16475 | -0.679 | -0.1002 | Yes | ||
| 113 | PKMYT1 | na | 16521 | -0.694 | -0.0845 | Yes | ||
| 114 | CDK6 | na | 16528 | -0.697 | -0.0665 | Yes | ||
| 115 | MAD2L1 | na | 16556 | -0.705 | -0.0495 | Yes | ||
| 116 | CCNE2 | na | 16756 | -0.769 | -0.0404 | Yes | ||
| 117 | MCM2 | na | 16776 | -0.776 | -0.0211 | Yes | ||
| 118 | SMC1B | na | 16893 | -0.819 | -0.0060 | Yes | ||
| 119 | E2F1 | na | 16952 | -0.845 | 0.0130 | Yes | ||
| 120 | CUL1 | na | 17633 | -1.410 | 0.0117 | Yes |