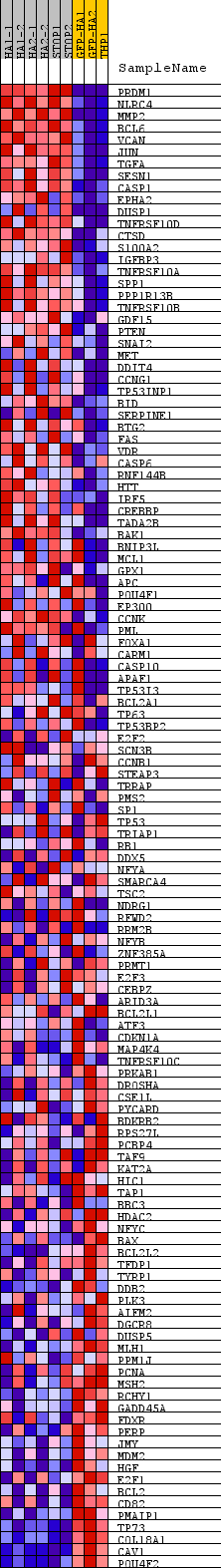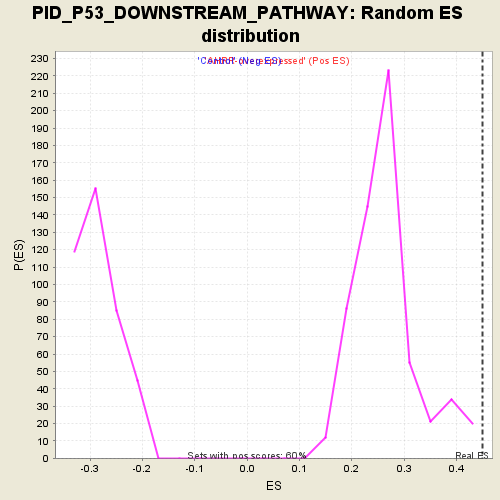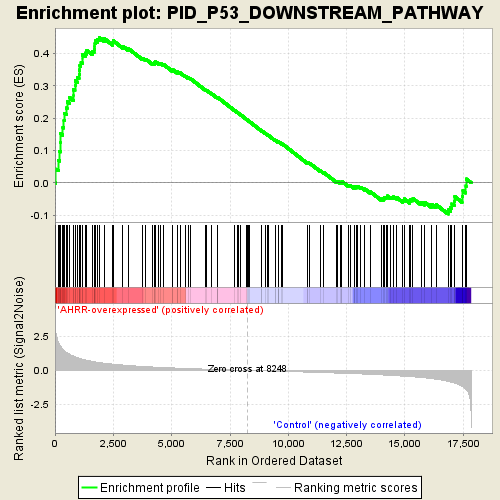
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA.class.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | AHRR-overexpressed |
| GeneSet | PID_P53_DOWNSTREAM_PATHWAY |
| Enrichment Score (ES) | 0.45015523 |
| Normalized Enrichment Score (NES) | 1.6960099 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.21744376 |
| FWER p-Value | 0.454 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PRDM1 | na | 35 | 2.825 | 0.0430 | Yes | ||
| 2 | NLRC4 | na | 144 | 2.079 | 0.0700 | Yes | ||
| 3 | MMP2 | na | 193 | 1.904 | 0.0976 | Yes | ||
| 4 | BCL6 | na | 231 | 1.829 | 0.1247 | Yes | ||
| 5 | VCAN | na | 239 | 1.778 | 0.1526 | Yes | ||
| 6 | JUN | na | 334 | 1.569 | 0.1723 | Yes | ||
| 7 | TGFA | na | 380 | 1.491 | 0.1935 | Yes | ||
| 8 | SESN1 | na | 396 | 1.460 | 0.2159 | Yes | ||
| 9 | CASP1 | na | 486 | 1.332 | 0.2321 | Yes | ||
| 10 | EPHA2 | na | 517 | 1.294 | 0.2510 | Yes | ||
| 11 | DUSP1 | na | 626 | 1.188 | 0.2638 | Yes | ||
| 12 | TNFRSF10D | na | 782 | 1.063 | 0.2720 | Yes | ||
| 13 | CTSD | na | 786 | 1.061 | 0.2887 | Yes | ||
| 14 | S100A2 | na | 855 | 1.010 | 0.3010 | Yes | ||
| 15 | IGFBP3 | na | 864 | 1.001 | 0.3165 | Yes | ||
| 16 | TNFRSF10A | na | 967 | 0.934 | 0.3256 | Yes | ||
| 17 | SPP1 | na | 1028 | 0.892 | 0.3364 | Yes | ||
| 18 | PPP1R13B | na | 1044 | 0.887 | 0.3497 | Yes | ||
| 19 | TNFRSF10B | na | 1062 | 0.880 | 0.3627 | Yes | ||
| 20 | GDF15 | na | 1110 | 0.859 | 0.3737 | Yes | ||
| 21 | PTEN | na | 1170 | 0.825 | 0.3835 | Yes | ||
| 22 | SNAI2 | na | 1171 | 0.823 | 0.3967 | Yes | ||
| 23 | MET | na | 1294 | 0.778 | 0.4022 | Yes | ||
| 24 | DDIT4 | na | 1359 | 0.751 | 0.4105 | Yes | ||
| 25 | CCNG1 | na | 1594 | 0.664 | 0.4079 | Yes | ||
| 26 | TP53INP1 | na | 1673 | 0.639 | 0.4136 | Yes | ||
| 27 | BID | na | 1705 | 0.631 | 0.4220 | Yes | ||
| 28 | SERPINE1 | na | 1710 | 0.630 | 0.4318 | Yes | ||
| 29 | BTG2 | na | 1743 | 0.622 | 0.4399 | Yes | ||
| 30 | FAS | na | 1824 | 0.600 | 0.4449 | Yes | ||
| 31 | VDR | na | 1896 | 0.582 | 0.4502 | Yes | ||
| 32 | CASP6 | na | 2115 | 0.537 | 0.4464 | No | ||
| 33 | RNF144B | na | 2476 | 0.466 | 0.4335 | No | ||
| 34 | HTT | na | 2486 | 0.465 | 0.4404 | No | ||
| 35 | IRF5 | na | 2911 | 0.398 | 0.4228 | No | ||
| 36 | CREBBP | na | 3142 | 0.368 | 0.4157 | No | ||
| 37 | TADA2B | na | 3751 | 0.297 | 0.3861 | No | ||
| 38 | BAK1 | na | 3897 | 0.281 | 0.3824 | No | ||
| 39 | BNIP3L | na | 4194 | 0.250 | 0.3697 | No | ||
| 40 | MCL1 | na | 4251 | 0.246 | 0.3704 | No | ||
| 41 | GPX1 | na | 4265 | 0.245 | 0.3736 | No | ||
| 42 | APC | na | 4307 | 0.241 | 0.3751 | No | ||
| 43 | POU4F1 | na | 4450 | 0.230 | 0.3708 | No | ||
| 44 | EP300 | na | 4523 | 0.225 | 0.3703 | No | ||
| 45 | CCNK | na | 4643 | 0.215 | 0.3670 | No | ||
| 46 | PML | na | 5033 | 0.186 | 0.3480 | No | ||
| 47 | FOXA1 | na | 5052 | 0.184 | 0.3499 | No | ||
| 48 | CARM1 | na | 5232 | 0.173 | 0.3426 | No | ||
| 49 | CASP10 | na | 5242 | 0.173 | 0.3448 | No | ||
| 50 | APAF1 | na | 5363 | 0.164 | 0.3406 | No | ||
| 51 | TP53I3 | na | 5587 | 0.149 | 0.3304 | No | ||
| 52 | BCL2A1 | na | 5708 | 0.140 | 0.3259 | No | ||
| 53 | TP63 | na | 5786 | 0.136 | 0.3237 | No | ||
| 54 | TP53BP2 | na | 6451 | 0.095 | 0.2878 | No | ||
| 55 | E2F2 | na | 6489 | 0.093 | 0.2872 | No | ||
| 56 | SCN3B | na | 6705 | 0.082 | 0.2763 | No | ||
| 57 | CCNB1 | na | 6949 | 0.068 | 0.2637 | No | ||
| 58 | STEAP3 | na | 6968 | 0.067 | 0.2637 | No | ||
| 59 | TRRAP | na | 6973 | 0.066 | 0.2646 | No | ||
| 60 | PMS2 | na | 7695 | 0.029 | 0.2243 | No | ||
| 61 | SP1 | na | 7818 | 0.022 | 0.2178 | No | ||
| 62 | TP53 | na | 7860 | 0.019 | 0.2158 | No | ||
| 63 | TRIAP1 | na | 7938 | 0.015 | 0.2117 | No | ||
| 64 | RB1 | na | 8213 | 0.002 | 0.1962 | No | ||
| 65 | DDX5 | na | 8260 | -0.001 | 0.1937 | No | ||
| 66 | NFYA | na | 8285 | -0.001 | 0.1923 | No | ||
| 67 | SMARCA4 | na | 8334 | -0.004 | 0.1897 | No | ||
| 68 | TSC2 | na | 8870 | -0.030 | 0.1600 | No | ||
| 69 | NDRG1 | na | 8871 | -0.030 | 0.1604 | No | ||
| 70 | RFWD2 | na | 9014 | -0.037 | 0.1530 | No | ||
| 71 | RRM2B | na | 9114 | -0.041 | 0.1481 | No | ||
| 72 | NFYB | na | 9143 | -0.043 | 0.1472 | No | ||
| 73 | ZNF385A | na | 9437 | -0.057 | 0.1315 | No | ||
| 74 | PRMT1 | na | 9471 | -0.058 | 0.1306 | No | ||
| 75 | E2F3 | na | 9560 | -0.062 | 0.1266 | No | ||
| 76 | CEBPZ | na | 9575 | -0.063 | 0.1268 | No | ||
| 77 | ARID3A | na | 9584 | -0.063 | 0.1274 | No | ||
| 78 | BCL2L1 | na | 9719 | -0.069 | 0.1209 | No | ||
| 79 | ATF3 | na | 9747 | -0.071 | 0.1205 | No | ||
| 80 | CDKN1A | na | 10829 | -0.127 | 0.0615 | No | ||
| 81 | MAP4K4 | na | 10836 | -0.127 | 0.0632 | No | ||
| 82 | TNFRSF10C | na | 10921 | -0.132 | 0.0606 | No | ||
| 83 | PRKAB1 | na | 11377 | -0.157 | 0.0374 | No | ||
| 84 | DROSHA | na | 11519 | -0.166 | 0.0321 | No | ||
| 85 | CSE1L | na | 12084 | -0.197 | 0.0034 | No | ||
| 86 | PYCARD | na | 12103 | -0.198 | 0.0055 | No | ||
| 87 | BDKRB2 | na | 12229 | -0.205 | 0.0018 | No | ||
| 88 | RPS27L | na | 12290 | -0.209 | 0.0017 | No | ||
| 89 | PCBP4 | na | 12297 | -0.209 | 0.0047 | No | ||
| 90 | TAF9 | na | 12597 | -0.226 | -0.0086 | No | ||
| 91 | KAT2A | na | 12666 | -0.230 | -0.0088 | No | ||
| 92 | HIC1 | na | 12829 | -0.242 | -0.0141 | No | ||
| 93 | TAP1 | na | 12836 | -0.242 | -0.0105 | No | ||
| 94 | BBC3 | na | 12933 | -0.248 | -0.0120 | No | ||
| 95 | HDAC2 | na | 12970 | -0.251 | -0.0100 | No | ||
| 96 | NFYC | na | 13111 | -0.261 | -0.0138 | No | ||
| 97 | BAX | na | 13274 | -0.273 | -0.0186 | No | ||
| 98 | BCL2L2 | na | 13518 | -0.293 | -0.0276 | No | ||
| 99 | TFDP1 | na | 14006 | -0.335 | -0.0498 | No | ||
| 100 | TYRP1 | na | 14072 | -0.340 | -0.0480 | No | ||
| 101 | DDB2 | na | 14109 | -0.344 | -0.0446 | No | ||
| 102 | PLK3 | na | 14196 | -0.352 | -0.0438 | No | ||
| 103 | AIFM2 | na | 14235 | -0.355 | -0.0403 | No | ||
| 104 | DGCR8 | na | 14402 | -0.370 | -0.0438 | No | ||
| 105 | DUSP5 | na | 14499 | -0.380 | -0.0432 | No | ||
| 106 | MLH1 | na | 14623 | -0.392 | -0.0438 | No | ||
| 107 | PPM1J | na | 14907 | -0.424 | -0.0531 | No | ||
| 108 | PCNA | na | 14965 | -0.429 | -0.0495 | No | ||
| 109 | MSH2 | na | 15206 | -0.455 | -0.0558 | No | ||
| 110 | RCHY1 | na | 15233 | -0.459 | -0.0499 | No | ||
| 111 | GADD45A | na | 15344 | -0.474 | -0.0486 | No | ||
| 112 | FDXR | na | 15695 | -0.527 | -0.0599 | No | ||
| 113 | PERP | na | 15852 | -0.550 | -0.0600 | No | ||
| 114 | JMY | na | 16127 | -0.600 | -0.0659 | No | ||
| 115 | MDM2 | na | 16337 | -0.647 | -0.0674 | No | ||
| 116 | HGF | na | 16852 | -0.804 | -0.0836 | No | ||
| 117 | E2F1 | na | 16952 | -0.845 | -0.0757 | No | ||
| 118 | BCL2 | na | 17015 | -0.875 | -0.0653 | No | ||
| 119 | CD82 | na | 17114 | -0.921 | -0.0561 | No | ||
| 120 | PMAIP1 | na | 17141 | -0.940 | -0.0426 | No | ||
| 121 | TP73 | na | 17450 | -1.173 | -0.0413 | No | ||
| 122 | COL18A1 | na | 17487 | -1.203 | -0.0242 | No | ||
| 123 | CAV1 | na | 17617 | -1.372 | -0.0096 | No | ||
| 124 | POU4F2 | na | 17626 | -1.395 | 0.0121 | No |