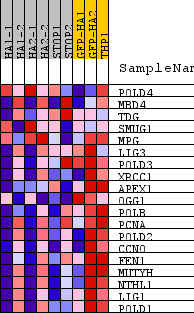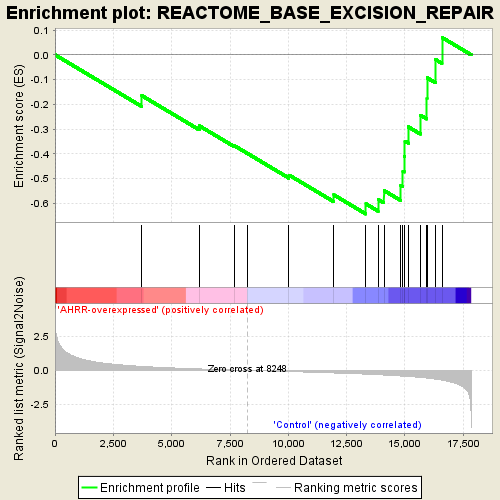
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | normalized_counts_per_kilobase_for_GSEA.class.cls #AHRR-overexpressed_versus_Control |
| Phenotype | class.cls#AHRR-overexpressed_versus_Control |
| Upregulated in class | Control |
| GeneSet | REACTOME_BASE_EXCISION_REPAIR |
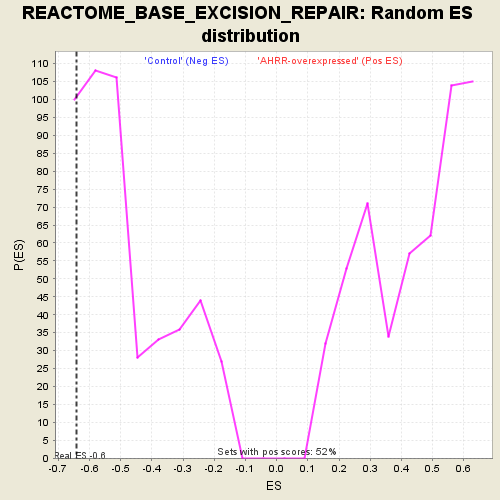
| Enrichment Score (ES) | -0.6428084 |
| Normalized Enrichment Score (NES) | -1.3452711 |
| Nominal p-value | 0.1307054 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | POLD4 | na | 3720 | 0.300 | -0.1647 | No | ||
| 2 | MBD4 | na | 6187 | 0.111 | -0.2868 | No | ||
| 3 | TDG | na | 7701 | 0.029 | -0.3675 | No | ||
| 4 | SMUG1 | na | 8240 | 0.000 | -0.3977 | No | ||
| 5 | MPG | na | 10029 | -0.086 | -0.4854 | No | ||
| 6 | LIG3 | na | 11955 | -0.189 | -0.5657 | No | ||
| 7 | POLD3 | na | 13330 | -0.279 | -0.6019 | Yes | ||
| 8 | XRCC1 | na | 13850 | -0.321 | -0.5840 | Yes | ||
| 9 | APEX1 | na | 14103 | -0.343 | -0.5479 | Yes | ||
| 10 | OGG1 | na | 14825 | -0.414 | -0.5276 | Yes | ||
| 11 | POLB | na | 14915 | -0.424 | -0.4704 | Yes | ||
| 12 | PCNA | na | 14965 | -0.429 | -0.4103 | Yes | ||
| 13 | POLD2 | na | 15002 | -0.434 | -0.3487 | Yes | ||
| 14 | CCNO | na | 15144 | -0.449 | -0.2908 | Yes | ||
| 15 | FEN1 | na | 15663 | -0.522 | -0.2433 | Yes | ||
| 16 | MUTYH | na | 15946 | -0.568 | -0.1759 | Yes | ||
| 17 | NTHL1 | na | 15949 | -0.569 | -0.0927 | Yes | ||
| 18 | LIG1 | na | 16316 | -0.642 | -0.0192 | Yes | ||
| 19 | POLD1 | na | 16591 | -0.714 | 0.0701 | Yes |